- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Volume 64(2); 2026 > Article
-
Review
Armored RNA technology as a clinical diagnostics tool for future pandemic preparedness - Jin Hao Tan1, Prashant Mainali1, Wei Zhang2, Dave Siak-Wei Ow1,*
-
Journal of Microbiology 2026;64(2):e2510016.
DOI: https://doi.org/10.71150/jm.2510016
Published online: February 28, 2026
1Microbial Cell Bioprocessing, Bioprocessing Technology Institute (BTI), Agency for Science, Technology and Research (A*STAR), Singapore 138668, Republic of Singapore
2Downstream Processing, Bioprocessing Technology Institute (BTI), Agency for Science, Technology and Research (A*STAR), Singapore 138668, Republic of Singapore
- *Correspondence Dave Siak-Wei Ow dave_ow@a-star.edu.sg
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
- 1,051 Views
- 68 Download
ABSTRACT
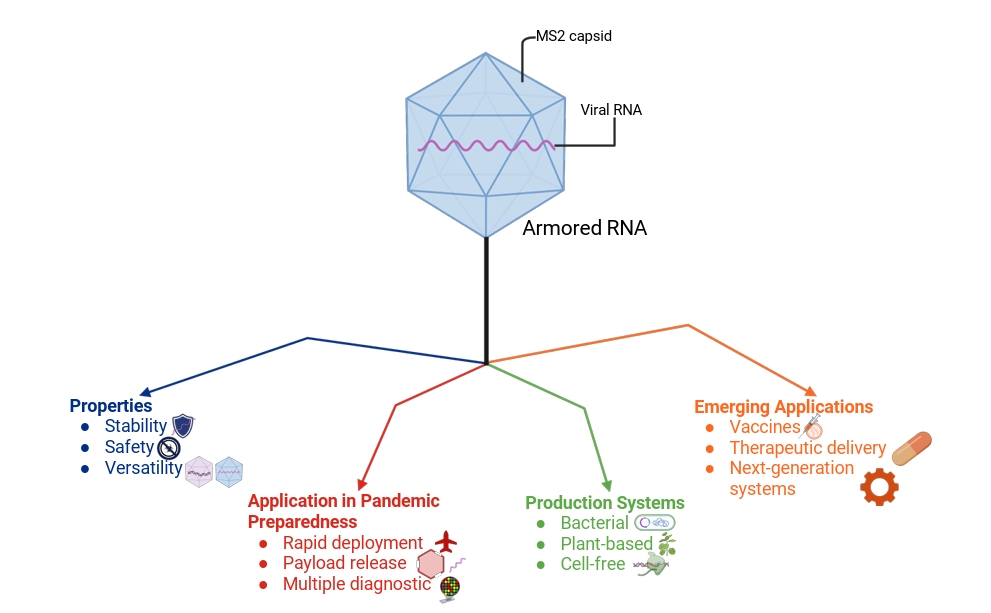
- The COVID-19 pandemic highlighted the critical role of reliable molecular diagnostics in outbreak response and the vulnerabilities of existing systems to delays and reagent instability. Armored RNA technology, which packages RNA within bacteriophage-derived capsids, offers a robust solution by combining nuclease resistance, safety, and versatility into a single platform. Armored RNA has become a trusted internal and external control for RT-qPCR and RT-LAMP, enabling accurate detection across a wide range of viral pathogens. Also, recent advances in alternative expression systems, such as plant-based and cell-free platforms, as well as the use of more stable scaffolds from bacteriophage Qβ, are enhancing yield, stability, and accessibility of armored RNA. Engineering innovations, including capsid polymorphism and optimized downstream purification, further improve efficiency and broaden possible applications. Looking ahead, armored RNA holds promise not only as a diagnostic standard but also as a delivery vehicle for vaccines and therapeutics. Encapsulation of self-amplifying RNA, small interfering RNA, or microRNA could open new pathways for rapid-response vaccines and targeted therapies, aligning this technology with the future of precision medicine. By uniting stability, scalability, and adaptability, armored RNA represents a critical component of global health preparedness, with the potential to strengthen diagnostic resilience and accelerate biomedical countermeasures in future pandemics.
Introduction
The Armored RNA (aRNA) Technology
Application of aRNA in Pandemic Preparedness
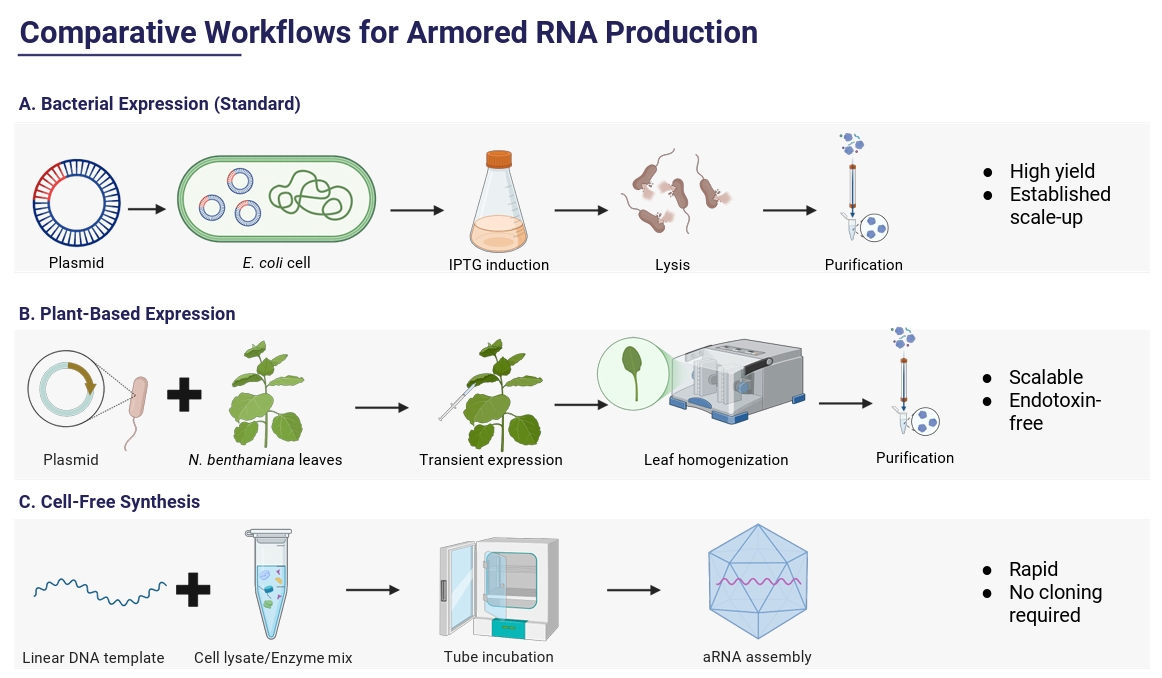
Alternative Production Systems
Limitations and Considerations
Future Directions and Emerging Applications
Closing Summary
Acknowledgments
This research was supported by the Agency for Science, Technology and Research (A∗STAR), Singapore, and the authors gratefully acknowledge the financial support of the MTC – Young Investigator Research Grant (YIRG grant) M24N8c0107. All figures were created with BioRender.com.
Conflict of Interest
The authors declare that they have no competing interests.
Ethics Statement
Not applicable.
| Publication year | Viral target | Production system | RT-PCR limit of detection (LOD) | Reference |
|---|---|---|---|---|
| 1998 | HIV-1 | E. coli DH5α | Not tested | Pasloske et al. (1998) |
| 1999 | Hepatitis C | E. coli DH5α | Not tested | WalkerPeach et al. (1999) |
| 2005 | Enterovirus | Not specified | Not specified | Donia et al. (2005) |
| 2006 | Classical swine fever virus (CSFV), foot-and-mouth disease virus (FMDV), vesicular stomatitis virus (VSV) | Cell-free production | 102 to 103 copies for CSFV, and 103 to 104 copies for FMDV, 10 to 102 target copies for VSV | Hietala and Crossley (2006) |
| 2007 | Rebulla | E. coli | Not specified | Zhao et al. (2007) |
| 2007 | Influenza A, influenza B, SARS | E. coli BL21 (DE3) | 101 copies/μl of AR-2 | Yu et al. (2008) |
| 2008 | SARS-CoV | E. coli DH5α | 25 copies/reaction | Stevenson et al. (2008) |
| 2009 | HIV-1 | E. coli BL21 (DE3) | 50 copies/ml | Zhan et al. (2009) |
| 2010 | Influenza A, influenza B, RSV, H1N1 | E. coli DH5α | Not reported | Hymas et al. (2010) |
| 2011 | EV71, CA16, pan-EV | E. coli DH5α | Not reported | Song et al. (2011) |
| 2013 | Influenza A (H1N1, H5N1, H9N2) | E. coli DH5α | 100–101 copies/ml | Chen et al. (2013) |
| 2013 | Influenza A (H7N9) | E. coli BL21 (DE3) | Not reported | Sun et al. (2013) |
| 2015 | Foot-and-mouth disease | Nicotiana benthamiana | Not reported | Madi et al. (2015) |
| 2015 | Measles | E. coli BL21 (DE3) | Not reported | Zhang et al. (2015b) |
| 2015 | HBV, HPV | E. coli BL21 (DE3) | Not tested | Zhang et al. (2015a) |
| 2016 | MERS-CoV | E. coli BL21 (DE3) | Not reported | Zhang et al. (2016a) |
| 2020 | SARS-CoV-2 | E. coli BL21 (DE3) | 103 copies/ml | Goncharova et al. (2021) |
| 2021 | Zika | E. coli BL21 (DE3) | Not reported | Lin et al. (2017) |
| 2022 | SARS-CoV-2 | Nicotiana benthamiana | Not reported | Peyret et al. (2022) |
| 2024 | nOPV2 poliovirus | E. coli BL21 (DE3) | 103 copies/ml | Dolgova et al. (2024) |
| 2025 | Measles | E. coli BL21 (DE3) | 103 copies/ml | Chayeb et al. (2025) |
- Asor R, Schlicksup CJ, Zhao Z, Zlotnick A, Raviv U. 2020. Rapidly forming early intermediate structures dictate the pathway of capsid assembly. J Am Chem Soc. 142: 7868–7882. ArticlePubMedPMCLink
- Beld M, Minnaar R, Weel J, Sol C, Damen M, et al. 2004. Highly sensitive assay for detection of enterovirus in clinical specimens by reverse transcription-PCR with an armored RNA internal control. J Clin Microbiol. 42: 3059–3064. ArticlePubMedPMCLink
- Biddlecome A, Habte HH, McGrath KM, Sambanthamoorthy S, Wurm M, et al. 2019. Delivery of self-amplifying RNA vaccines in in vitro reconstituted virus-like particles. PLoS One. 14: e0215031. ArticlePubMedPMC
- Biela AP, Naskalska A, Fatehi F, Twarock R, Heddle JG. 2022. Programmable polymorphism of a virus-like particle. Commun Mater. 3: 7.ArticlePubMedPMCPDF
- Boom R, Sol C, Beld M, Weel J, Goudsmit J, et al. 1999. Improved silica-guanidiniumthiocyanate DNA isolation procedure based on selective binding of bovine alpha-casein to silica particles. J Clin Microbiol. 37: 615–619. ArticlePubMedPMCLink
- Boom R, Sol CJ, Salimans MM, Jansen CL, Wertheim-van Dillen PM, et al. 1990. Rapid and simple method for purification of nucleic acids. J Clin Microbiol. 28: 495–503. ArticlePubMedPMCLink
- Chan SK, Du P, Ignacio C, Mehta S, Newton IG, et al. 2021. Virus-like particles as positive controls for COVID-19 RT-LAMP diagnostic assays. Biomacromolecules. 22: 1231–1243. ArticlePubMed
- Chayeb VA, Dolgova AS, Popova MR, Zheleznova NV, Shirobokova SA, et al. 2025. Development and evaluation of a new measles detection assay using real-time RT-PCR. Int J Mol Sci. 26: 1801.ArticlePubMedPMC
- Chen Y, Liu T, Cai L, Du H, Li M. 2013. A one-step RT-PCR array for detection and differentiation of zoonotic influenza viruses H5N1, H9N2, and H1N1. J Clin Lab Anal. 27: 450–460. ArticlePubMedPMC
- Chung YH, Volckaert BA, Steinmetz NF. 2023. Development of a modular NTA:His tag viral vaccine for co-delivery of antigen and adjuvant. Bioconjug Chem. 34: 269–278. ArticlePubMedPMCLink
- Costa VG, Gomes AJ, Bittar C, Geraldini DB, Previdelli da Conceição PJ, et al. 2023. Burden of influenza and respiratory syncytial viruses in suspected COVID-19 patients: a cross-sectional and meta-analysis study. Viruses. 15: 665.ArticlePubMedPMC
- Crone MA, Freemont PS. 2022. Simple low-cost production of DNA MS2 virus-like particles as molecular diagnostic controls. GEN Biotechnol. 1: 496–503. ArticlePubMedPMC
- Crone MA, Priestman M, Ciechonska M, Jensen K, Sharp DJ, et al. 2020. A role for biofoundries in rapid development and validation of automated SARS-CoV-2 clinical diagnostics. Nat Commun. 11: 4464.ArticlePubMedPMCPDF
- de Maddalena LL, Niederholtmeyer H, Turtola M, Swank ZN, Belogurov GA, et al. 2016. GreA and GreB enhance expression of Escherichia coli RNA polymerase promoters in a reconstituted transcription-translation system. ACS Synth Biol. 5: 929–935. ArticlePubMed
- Dolgova AS, Kanaeva OI, Antonov SA, Shabalina AV, Klyuchnikova EO, et al. 2024. Qualitative real-time RT-PCR assay for nOPV2 poliovirus detection. J Virol Methods. 329: 114984.ArticlePubMed
- Donia D, Divizia M, Pana’ A. 2005. Use of armored RNA as a standard to construct a calibration curve for real-time RT-PCR. J Virol Methods. 126: 157–163. ArticlePubMed
- Galaway FA, Stockley PG. 2013. MS2 virus-like particles: a robust, semisynthetic targeted drug delivery platform. Mol Pharm. 10: 59–68. ArticlePubMed
- Gholami M, Ravanshad M, Baesi K, Samiee SM, Hosseini Rozbahani N, et al. 2018. Preparation and evaluation of ribonuclease-resistant viral HIV RNA standards based on armored RNA technology. Iran Biomed J. 22: 394–400. ArticlePubMedPMCPDF
- Goncharova EA, Dedkov VG, Dolgova AS, Kassirov IS, Safonova MV, et al. 2021. One-step quantitative RT-PCR assay with armored RNA controls for detection of SARS-CoV-2. J Med Virol. 93: 1694–1701. ArticlePubMedPMCLink
- Guo W, Alarcon E, Sanchez JE, Xiao C, Li L. 2024. Modeling viral capsid assembly: a review of computational strategies and applications. Cells. 13: 2088.ArticlePubMedPMC
- Hashemi K, Ghahramani Seno MM, Ahmadian MR, Malaekeh-Nikouei B, Bassami MR, et al. 2021. Optimizing the synthesis and purification of MS2 virus-like particles. Sci Rep. 11: 19851.ArticlePubMedPMCPDF
- Hemmati F, Hemmati-Dinarvand M, Karimzade M, Rutkowska D, Eskandari MH, et al. 2022. Plant-derived VLP: a worthy platform to produce vaccine against SARS-CoV-2. Biotechnol Lett. 44: 45–57. ArticlePubMedPMCPDF
- Hietala SK, Crossley BM. 2006. Armored RNA as virus surrogate in a real-time reverse transcriptase PCR assay proficiency panel. J Clin Microbiol. 44: 67–70. ArticlePubMedPMCLink
- Hsu LM, Vo NV, Chamberlin MJ. 1995. Escherichia coli transcript cleavage factors GreA and GreB stimulate promoter escape and gene expression in vivo and in vitro. Proc Natl Acad Sci USA. 92: 11588–11592. ArticlePubMedPMC
- Huang Q, Cheng Y, Guo Q, Li Q. 2006. Preparation of a chimeric armored RNA as a versatile calibrator for multiple virus assays. Clin Chem. 52: 1446–1448. ArticlePubMedPMCPDF
- Huang X, Wang X, Zhang J, Xia N, Zhao Q. 2017. Escherichia coli-derived virus-like particles in vaccine development. NPJ Vaccines. 2: 3.ArticlePubMedPMCPDF
- Hunt AC, Rasor BJ, Seki K, Ekas HM, Warfel KF, et al. 2025. Cell-free gene expression: methods and applications. Chem Rev. 125: 91–149. ArticlePubMedLink
- Hymas WC, Mills A, Ferguson S, Langer J, She RC, et al. 2010. Development of a multiplex real-time RT-PCR assay for detection of influenza A, influenza B, RSV and typing of the 2009-H1N1 influenza virus. J Virol Methods. 167: 113–118. ArticlePubMed
- Inbanathan FY, Wijesinghe PR, Naidoo D, Buddha N, Salvador EC, et al. 2024. Assessment of public health laboratory preparedness and response in WHO South-East Asia region during the COVID-19 pandemic: lessons learned and future directions. Lancet Reg Health Southeast Asia. 31: 100496.ArticlePubMedPMC
- Jester BJ, Uyeki TM, Patel A, Koonin L, Jernigan DB. 2018. 100 years of medical countermeasures and pandemic influenza preparedness. Am J Public Health. 108: 1469–1472. ArticlePubMedPMC
- Johnson RA, White RC, Disbrow GL. 2022. Advancing development of medical countermeasures: incorporating COVID-19 lessons learned into future pandemic preparedness planning. Hum Vaccin Immunother. 18: 2129930.ArticlePubMedPMC
- Kumar P, Singh RK, Shahgholian A. 2024. Learnings from COVID-19 for managing humanitarian supply chains: systematic literature review and future research directions. Ann Oper Res. 335: 899–935. ArticlePubMedPMCPDF
- Lee JM, Jansen R, Sanderson KE, Guerra F, Keller-Olaman S, et al. 2023. Public health emergency preparedness for infectious disease emergencies: a scoping review of recent evidence. BMC Public Health. 23: 420.ArticlePubMedPMCPDF
- Liang M, Li Y, Yang J, Liu C, Lin H, et al. 2025. Construction and application of MS2 bacteriophage virus-like particles for SARS-CoV-2 detection using a single-plasmid system. J Microbiol Methods. 240: 107344.ArticlePubMed
- Lin G, Zhang K, Zhang D, Han Y, Xie J, et al. 2017. Fast preparation of a long chimeric armored RNA as controls for external quality assessment for molecular detection of Zika virus. Clin Chim Acta. 466: 138–144. ArticlePubMed
- Liu X, Chang X, Rothen D, Derveni M, Krenger P, et al. 2021. AP205 VLPs based on dimerized capsid proteins accommodate RBM domain of SARS-CoV-2 and serve as an attractive vaccine candidate. Vaccines. 9: 403.Article
- Liu L, Qiao B, Rong J, Wang E, Yu K. 2024. The removal of endotoxins in the actual production process. Int J Public Health Med Res. 2: 165–172. ArticleLink
- Madi M, Mioulet V, King DP, Lomonossoff GP, Montague NP. 2015. Development of a non-infectious encapsidated positive control RNA for molecular assays to detect foot-and-mouth disease virus. J Virol Methods. 220: 27–34. ArticlePubMedPMC
- Mikel P, Vasickova P, Kralik P. 2015. Methods for preparation of MS2 phage-like particles and their utilization as process control viruses in RT-PCR and qRT-PCR detection of RNA viruses from food matrices and clinical specimens. Food Environ. Virol. 7: 96–111. ArticlePubMedPMCPDF
- Pabbaraju K, Wong AA, Douesnard M, Ma R, Gill K, et al. 2020. A public health laboratory response to the pandemic. J Clin Microbiol. 58: e01110-20.ArticlePubMedPMCLink
- Pan Y, Zhang Y, Jia T, Zhang K, Li J, et al. 2012. Development of a microRNA delivery system based on bacteriophage MS2 virus-like particles. FEBS J. 279: 1198–1208. ArticlePubMed
- Pasloske BL, Walkerpeach CR, Obermoeller RD, Winkler M, DuBois DB. 1998. Armored RNA technology for production of ribonuclease-resistant viral RNA controls and standards. J Clin Microbiol. 36: 3590–3594. ArticlePubMedPMCLink
- Peabody DS. 1993. The RNA binding site of bacteriophage MS2 coat protein. EMBO J. 12: 595–600. ArticlePubMedPMCLink
- Peabody DS, Peabody J, Bradfute SB, Chackerian B. 2021. RNA phage VLP-based vaccine platforms. Pharmaceuticals. 14: 764.ArticlePubMedPMC
- Peyret H, Groppelli E, Clark D, Eckersley N, Planche T, et al. 2022. Production and use of encapsidated RNA mimics as positive control reagents for SARS-CoV-2 RT-qPCR diagnostics. J Virol Methods. 300: 114372.ArticlePubMedPMC
- Rozhkov M, Ivanov D, Blackhurst J, Nair A. 2022. Adapting supply chain operations in anticipation of and during the COVID-19 pandemic. Omega. 110: 102635.ArticlePubMedPMC
- Silva-Pilipich N, Beloki U, Salaberry L, Smerdou C. 2024. Self-amplifying RNA: a second revolution of mRNA vaccines against COVID-19. Vaccines. 12: 318.ArticlePubMedPMC
- Song L, Sun S, Li B, Pan Y, Li W, et al. 2011. External quality assessment for enterovirus 71 and coxsackievirus A16 detection by reverse transcription-PCR using armored RNA as a virus surrogate. J Clin Microbiol. 49: 3591–3595. ArticlePubMedPMCLink
- Stevenson J, Hymas W, Hillyard D. 2008. The use of armored RNA as a multi-purpose internal control for RT-PCR. J Virol Methods. 150: 73–76. ArticlePubMedPMC
- Sun Y, Jia T, Sun Y, Han Y, Wang L, et al. 2013. External quality assessment for avian influenza A (H7N9) virus detection using armored RNA. J Clin Microbiol. 51: 4055–4059. ArticlePubMedPMCLink
- Uddin MN, Roni MA. 2021. Challenges of storage and stability of mRNA-based COVID-19 vaccines. Vaccines. 9: 1033.ArticlePubMedPMC
- Vallet T, Vignuzzi M. 2025. Self-amplifying RNA: advantages and challenges of a versatile platform for vaccine development. Viruses. 17: 566.ArticlePubMedPMC
- WalkerPeach CR, Winkler M, DuBois DB, Pasloske BL. 1999. Ribonuclease-resistant RNA controls (armored RNA) for reverse transcription-PCR, branched DNA, and genotyping assays for hepatitis C virus. Clin Chem. 45: 2079–2085. ArticlePubMedPDF
- Wei B, Wei Y, Zhang K, Yang C, Wang J, et al. 2008a. Construction of armored RNA containing long-size chimeric RNA by increasing the number and affinity of the pac site in exogenous RNA and sequence coding coat protein of the MS2 bacteriophage. Intervirology. 51: 144–150. ArticlePubMedPMCLink
- Wei Y, Yang C, Wei B, Huang J, Wang L, et al. 2008b. RNase-resistant virus-like particles containing long chimeric RNA sequences produced by two-plasmid coexpression system. J Clin Microbiol. 46: 1734–1740. ArticlePubMedPMCLink
- WHO, World Health Organization. 2020. WHO Director-General’s opening remarks at the media briefing on COVID-19 - 11 March 2020. World Health Organization; Available from https://www.who.int/news-room/speeches/item/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19---11-march-2020.
- Wright AM, Snowdon A, Saunders M, Trampas D. 2024. The necessity of healthcare supply chain resilience for crisis preparedness. Healthc Manage Forum. 37: 95–100. ArticlePubMedPMCLink
- Xie T, Dai GY, Chen WF, Yang CP, Huang YJ, et al. 2025. Pandemic triggered emergency supply chain management innovations: a scientometric analysis based on bibliometrics and dynamic topic models. Disaster Med Public Health Prep. 19: e88.ArticlePubMed
- Yao L, Li F, Qu M, Guo Y, Jiang Y, et al. 2019. Development and evaluation of a novel armored RNA technology using bacteriophage Qβ. Food Environ Virol. 11: 383–392. ArticlePubMedPDF
- Yu XF, Pan JC, Ye R, Xiang HQ, Kou Y, et al. 2008. Preparation of armored RNA as a control for multiplex real-time reverse transcription-PCR detection of influenza virus and severe acute respiratory syndrome coronavirus. J Clin Microbiol. 46: 837–841. ArticlePubMedPMCLink
- Zhan S, Li J, Xu R, Wang L, Zhang K, et al. 2009. Armored long RNA controls or standards for branched DNA assay for detection of human immunodeficiency virus type 1. J Clin Microbiol. 47: 2571–2576. ArticlePubMedPMCLink
- Zhang L, Hao M, Zhang K, Zhang R, Lin G, et al. 2016a. External quality assessment for the molecular detection of MERS-CoV in China. J Clin Virol. 75: 5–9. ArticlePubMedPMC
- Zhang L, Sun Y, Chang L, Jia T, Wang G, et al. 2015a. A novel method to produce armored double-stranded DNA by encapsulation of MS2 viral capsids. Appl Microbiol Biotechnol. 99: 7047–7057. ArticlePubMedPMCPDF
- Zhang D, Sun Y, Jia T, Zhang L, Wang G, et al. 2015b. External quality assessment for the detection of measles virus by reverse transcription-PCR using armored RNA. PLoS One. 10: e0134681. ArticlePubMedPMC
- Zhang M, Wang N, Jing H, Deng J, Wu S, et al. 2016b. Method of preparing armored RNA with non-cellular expression system. CN105734046A. Available from https://patents.google.com/patent/CN105734046A/en.
- Zhang M, Wang N, Jing H, Deng J, Wu S. 2019. Method for preparing armored RNA by cell-free expression system. CN Patent CN105734046B. Available from https://patents.google.com/patent/CN105734046B/en.
- Zhao L, Ma Y, Zhao S, Yang N. 2007. Armored RNA as positive control and standard for quantitative reverse transcription-polymerase chain reaction assay for rubella virus. Arch Virol. 152: 219–224. ArticlePubMedPDF
- Zilberzwige-Tal S, Alon DM, Gazit D, Zachariah S, Hollander A, et al. 2021. Genetically encoding ultrastable virus-like particles encapsulating functional DNA nanostructures in living bacteria. ACS Synth Biol. 10: 1798–1807. ArticlePubMed
References
Figure & Data
References
Citations

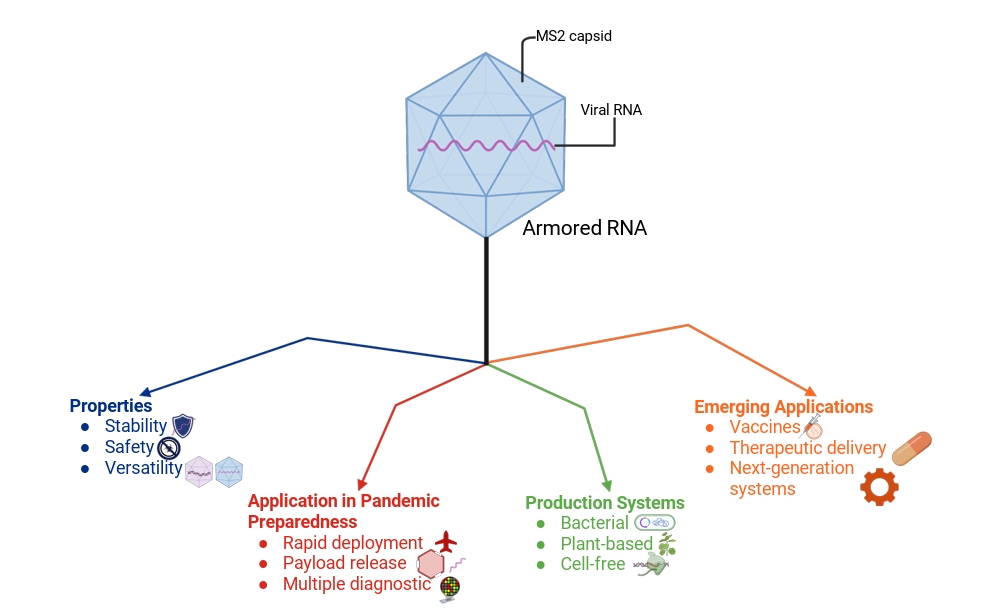
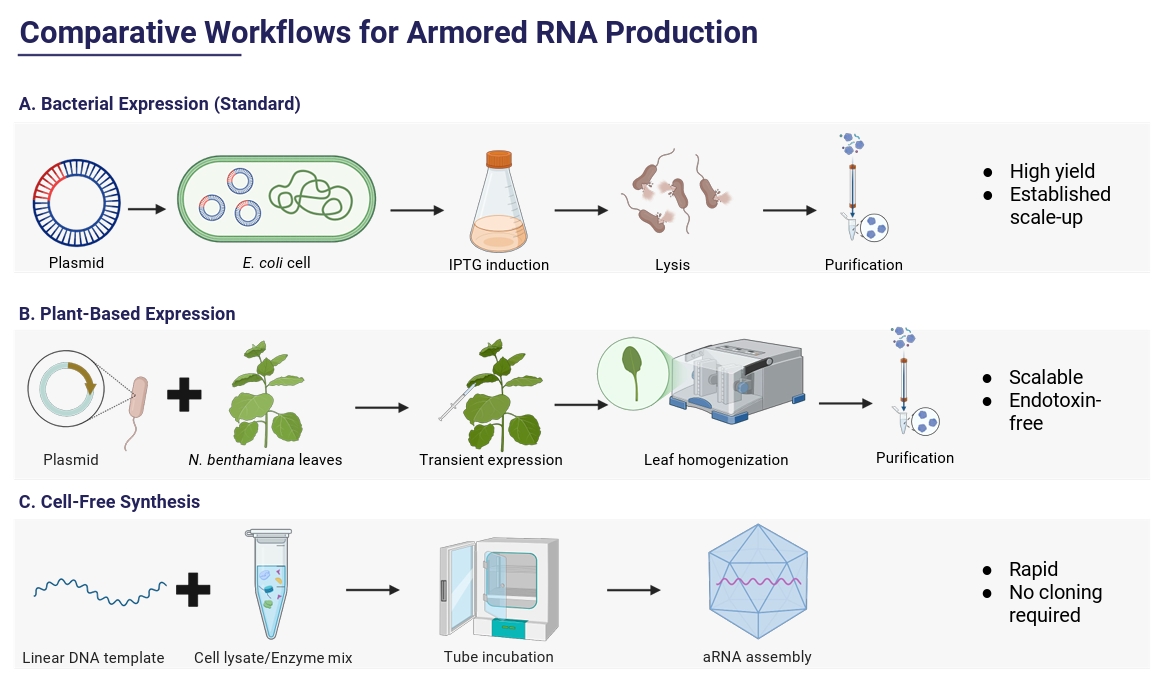
Fig. 1.
Fig. 2.
| Publication year | Viral target | Production system | RT-PCR limit of detection (LOD) | Reference |
|---|---|---|---|---|
| 1998 | HIV-1 | E. coli DH5α | Not tested | |
| 1999 | Hepatitis C | E. coli DH5α | Not tested | |
| 2005 | Enterovirus | Not specified | Not specified | |
| 2006 | Classical swine fever virus (CSFV), foot-and-mouth disease virus (FMDV), vesicular stomatitis virus (VSV) | Cell-free production | 102 to 103 copies for CSFV, and 103 to 104 copies for FMDV, 10 to 102 target copies for VSV | |
| 2007 | Rebulla | E. coli | Not specified | |
| 2007 | Influenza A, influenza B, SARS | E. coli BL21 (DE3) | 101 copies/μl of AR-2 | |
| 2008 | SARS-CoV | E. coli DH5α | 25 copies/reaction | |
| 2009 | HIV-1 | E. coli BL21 (DE3) | 50 copies/ml | |
| 2010 | Influenza A, influenza B, RSV, H1N1 | E. coli DH5α | Not reported | |
| 2011 | EV71, CA16, pan-EV | E. coli DH5α | Not reported | |
| 2013 | Influenza A (H1N1, H5N1, H9N2) | E. coli DH5α | 100–101 copies/ml | |
| 2013 | Influenza A (H7N9) | E. coli BL21 (DE3) | Not reported | |
| 2015 | Foot-and-mouth disease | Nicotiana benthamiana | Not reported | |
| 2015 | Measles | E. coli BL21 (DE3) | Not reported | |
| 2015 | HBV, HPV | E. coli BL21 (DE3) | Not tested | |
| 2016 | MERS-CoV | E. coli BL21 (DE3) | Not reported | |
| 2020 | SARS-CoV-2 | E. coli BL21 (DE3) | 103 copies/ml | |
| 2021 | Zika | E. coli BL21 (DE3) | Not reported | |
| 2022 | SARS-CoV-2 | Nicotiana benthamiana | Not reported | |
| 2024 | nOPV2 poliovirus | E. coli BL21 (DE3) | 103 copies/ml | |
| 2025 | Measles | E. coli BL21 (DE3) | 103 copies/ml |
| Characteristic | Bacterial systems | Plant-based systems | Cell-free production |
|---|---|---|---|
| Production speed | Fast (hours–1 day) | Slower (days for transient expression) | Very fast (hours, no growth) |
| Yield | High, scalable in fermenters | Moderate; variable per biomass | Lower but adequate for small batches |
| Cost | Low | Moderate–high | High per unit; economical only at small scale |
| Assembly complexity | Well-established but may need optimization | Handles complex VLPs; more variable | Highly controlled but technically demanding |
| Product quality & Stability | Good if purified; risk of endotoxin | Batch variability; plant contaminants possible | High control; must guard against RNase |
| Biosafety | Non-infectious, low risk | Low pathogen risk after purification | Extremely low risk; no living host |
| Scalability | Easy to scale to bioreactors | More space and resources needed; feasible | Limited by reagent cost and supply |
| Regulatory/Contaminants | Well understood; bacterial nucleic acids must be removed | Less established; possible plant compounds | Few biological contaminants; reagent purity critical |
| Storage & Shipping | Stable when frozen/lyophilized | Similar to bacterial system after purification | Most flexible; can be lyophilized, potentially room-temperature stable |
Table 1.
Table 2.
TOP
 MSK
MSK
 Cite this Article
Cite this Article



