- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Volume 64(4); 2026 > Article
-
Full article
Functional characterization of spike RBD mutations in SARS-CoV-2 Omicron-derived subvariants KP.3.1.1, LP.8.1, and NB.1.8.1. - Yeong Jun Kim1,*, Seon Jae Jeong1, Hye-Ra Lee1,2,*
-
Journal of Microbiology 2026;64(4):e2511014.
DOI: https://doi.org/10.71150/jm.2511014
Published online: April 6, 2026
1Department of Biotechnology and Bioinformatics, College of Science and Technology, Korea University, Sejong 30019, Republic of Korea
2Department of Lab Medicine, College of Medicine, Korea University, Seoul 08308, Republic of Korea
- *Correspondence. Yeong Jun Kim kyj1994@korea.ac.kr Hye-Ra Lee leehr@korea.ac.kr
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
- 929 Views
- 23 Download
ABSTRACT
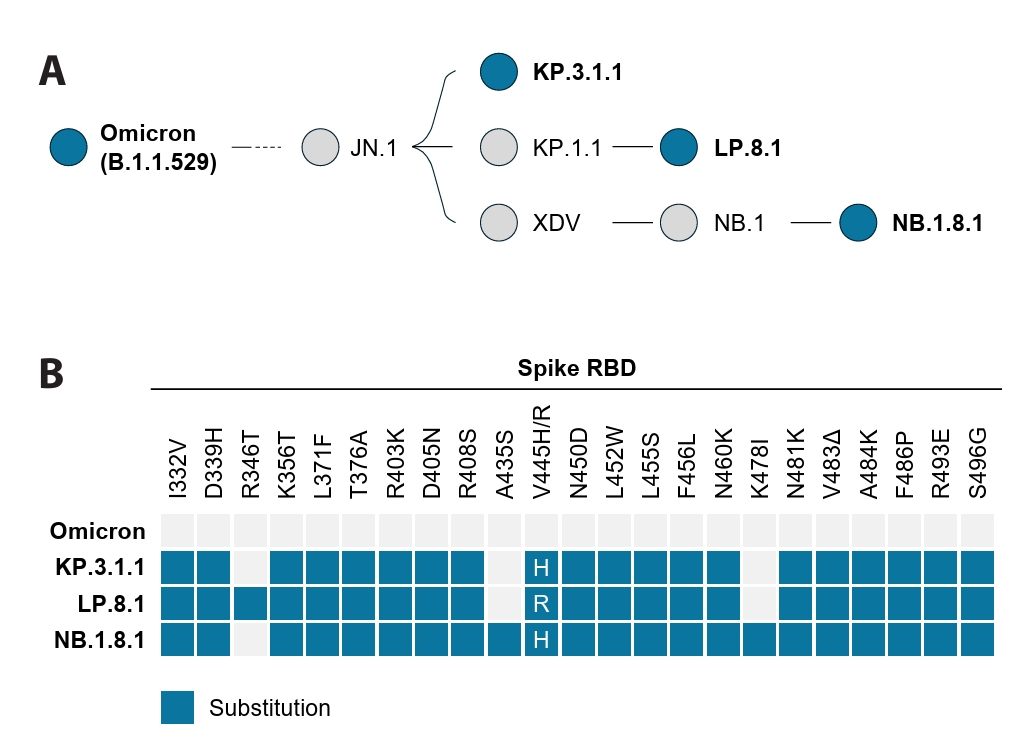
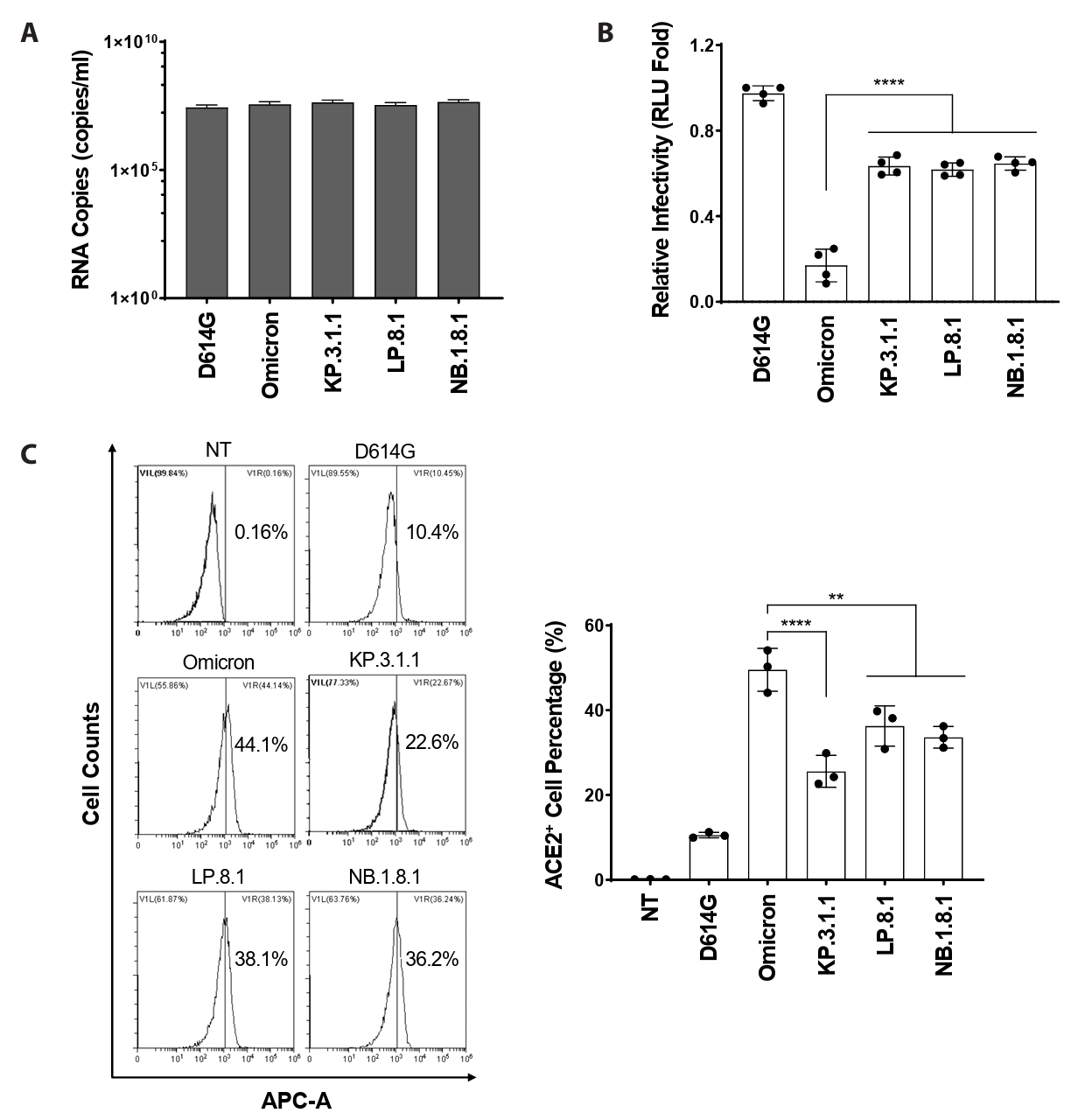
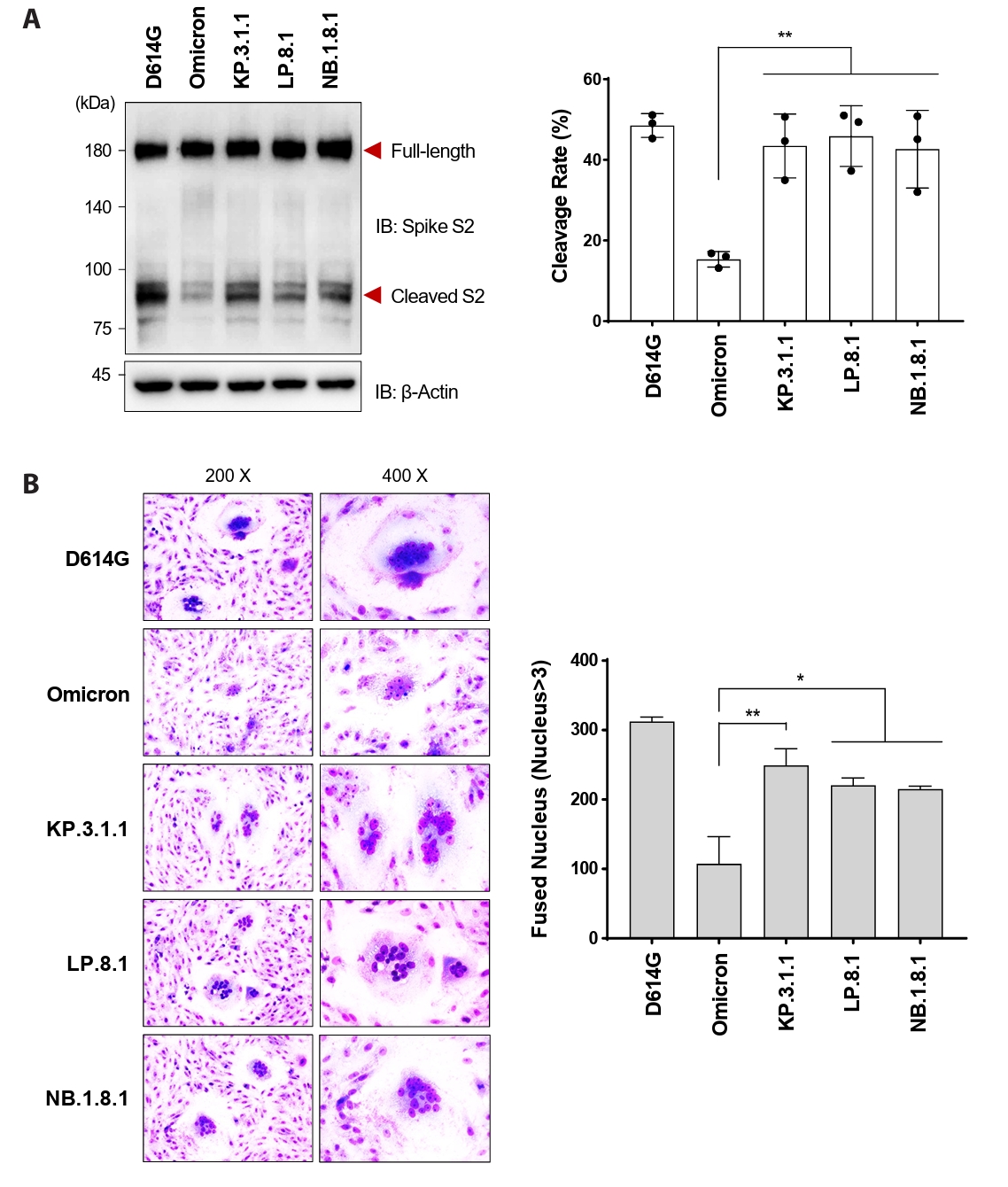
- Following the global spread of SARS-CoV-2 Omicron (B.1.1.529), its subvariants KP.3.1.1, LP.8.1, and NB.1.8.1 disseminated worldwide. By April 2025, the epidemiological landscape of these subvariants had become distinct, with LP.8.1 emerging as the predominant variant, KP.3.1.1 persisting as a co-circulating variant under monitoring (VUM), and NB.1.8.1 exhibiting a significant increase in prevalence. Despite their epidemiological prominence, the functional consequences of spike mutations defining these emerging subvariants remain poorly understood. Here, we systematically dissected the entry properties conferred by their receptor-binding domain (RBD) mutations using a pseudovirus system. Our results demonstrate that all three subvariants exhibited substantially higher infectivity than ancestral Omicron. Unexpectedly, this enhanced infectivity occurred despite reduced ACE2 binding affinity. Rather, increased viral entry consistently correlated with elevated spike cleavage efficiency and fusogenicity, suggesting a compensatory evolutionary strategy in which enhanced spike processing and fusion contribute to enhanced entry despite reduced receptor engagement. These findings provide a virological explanation for the accelerated global spread of these subvariants and highlight the importance of monitoring functional shifts in spike-mediated entry that may influence SARS-CoV-2 transmission dynamics.
Introduction
Materials and Methods
Results
Discussion
Acknowledgments
This work was supported by Grant of the National Research Foundation (NRF) funded by the Korean government (MSIT, Korea), RS-2021-NR056550, H.R.L., Grant of the Korea Health Technology R&D project through the Korean Health Industry Development Institute (KHIDI), Ministry of Health & Welfare, Republic of Korea, RS-2023-KH136036, H.R.L., and the Korea-US Collaborative Research Project funded by the Ministry of Science, ICT & Future Planning and the Ministry of Health and Welfare of Korea, RS-2024-00467046, H.R.L.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
- Cao Y, Jian F, Wang J, Yu Y, Song W, et al. 2023. Imprinted SARS-CoV-2 humoral immunity induces convergent Omicron RBD evolution. Nature. 614: 521–529. ArticlePubMedPMCPDF
- Cui Z, Liu P, Wang N, Wang L, Fan K, et al. 2022. Structural and functional characterizations of infectivity and immune evasion of SARS-CoV-2 Omicron. Cell. 185: 860–871. ArticlePubMedPMC
- Eckert DM, Kim PS. 2001. Mechanisms of viral membrane fusion and its inhibition. Annu Rev Biochem. 70: 777–810. ArticlePubMed
- Harvey WT, Carabelli AM, Jackson B, Gupta RK, Thomson EC, et al. 2021. SARS-CoV-2 variants, spike mutations and immune escape. Nat Rev Microbiol. 19: 409–424. ArticlePubMedPMCPDF
- Hoffmann M, Kleine-Weber H, Schroeder S, Kruger N, Herrler T, et al. 2020. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell. 181: 271–280. ArticlePubMedPMC
- Hu B, Chan JF, Liu H, Liu Y, Chai Y, et al. 2022. Spike mutations contributing to the altered entry preference of SARS-CoV-2 Omicron BA.1 and BA.2. Emerg Microbes Infect. 11: 2275–2287. ArticlePubMedPMC
- Kim YJ, Jang US, Soh SM, Lee JY, Lee HR, et al. 2021. The impact on infectivity and neutralization efficiency of SARS-CoV-2 lineage B.1.351 pseudovirus. Viruses. 13: 633.ArticlePubMedPMC
- Kimura I, Yamasoba D, Nasser H, Zahradnik J, Kosugi Y, et al. 2022. The SARS-CoV-2 spike S375F mutation characterizes the Omicron BA.1 variant. iScience. 25: 105720.ArticlePubMedPMC
- Mannar D, Saville JW, Zhu X, Srivastava SS, Berezuk AM, et al. 2022. SARS-CoV-2 Omicron variant: antibody evasion and cryo-EM structure of spike protein-ACE2 complex. Science. 375: 760–764. ArticlePubMedPMC
- Millet JK, Whittaker GR. 2016. Murine leukemia virus (MLV)-based coronavirus spike-pseudotyped particle production and infection. Bio Protoc. 6: e2035. ArticlePubMedPMC
- Park YJ, Pinto D, Walls AC, Liu Z, De Marco A, et al. 2022. Imprinted antibody responses against SARS-CoV-2 Omicron sublineages. Science. 378: 619–627. ArticlePubMed
- Planas D, Veyer D, Baidaliuk A, Staropoli I, Guivel-Benhassine F, et al. 2021. Reduced sensitivity of SARS-CoV-2 variant Delta to antibody neutralization. Nature. 596: 276–280. ArticlePubMedPDF
- Reeves JD, Gallo SA, Ahmad N, Miamidian JL, Harvey PE, et al. 2002. Sensitivity of HIV-1 to entry inhibitors correlates with envelope/coreceptor affinity, receptor density, and fusion kinetics. Proc Natl Acad Sci USA. 99: 16249–16254. ArticlePubMedPMC
- Tao K, Tzou PL, Nouhin J, Gupta RK, de Oliveira T, et al. 2021. The biological and clinical significance of emerging SARS-CoV-2 variants. Nat Rev Genet. 22: 757–773. ArticlePubMedPMCPDF
- Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, et al. 2020. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell. 181: 281–292. ArticlePubMedPMC
- Wang L, Zhao J, Nguyen LNT, Adkins JL, Schank M, et al. 2021. Blockade of SARS-CoV-2 spike protein-mediated cell-cell fusion using COVID-19 convalescent plasma. Sci Rep. 11: 5558.ArticlePubMedPMCPDF
- Willett BJ, Grove J, MacLean OA, Wilkie C, De Lorenzo G, et al. 2022. SARS-CoV-2 Omicron is an immune escape variant with an altered cell entry pathway. Nat Microbiol. 7: 1161–1179. ArticlePubMedPMC
- Zhou P, Yang XL, Wang XG, Hu B, Zhang L, et al. 2020. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 579: 270–273. ArticlePubMedPMC
References
Supplementary Information
References
Citations

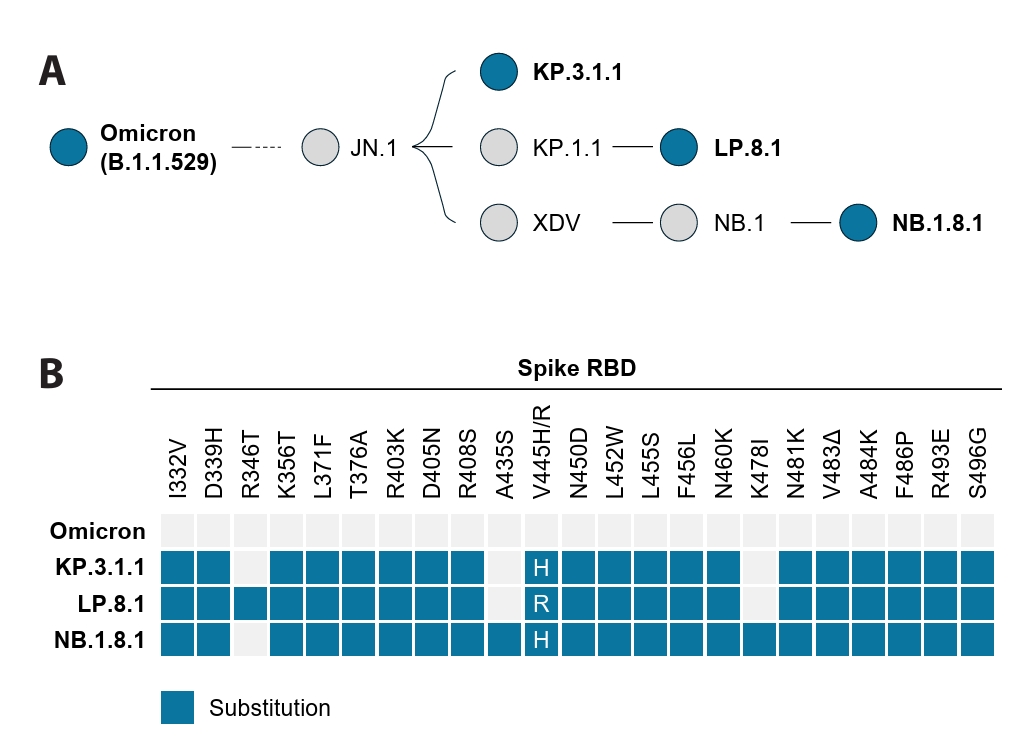
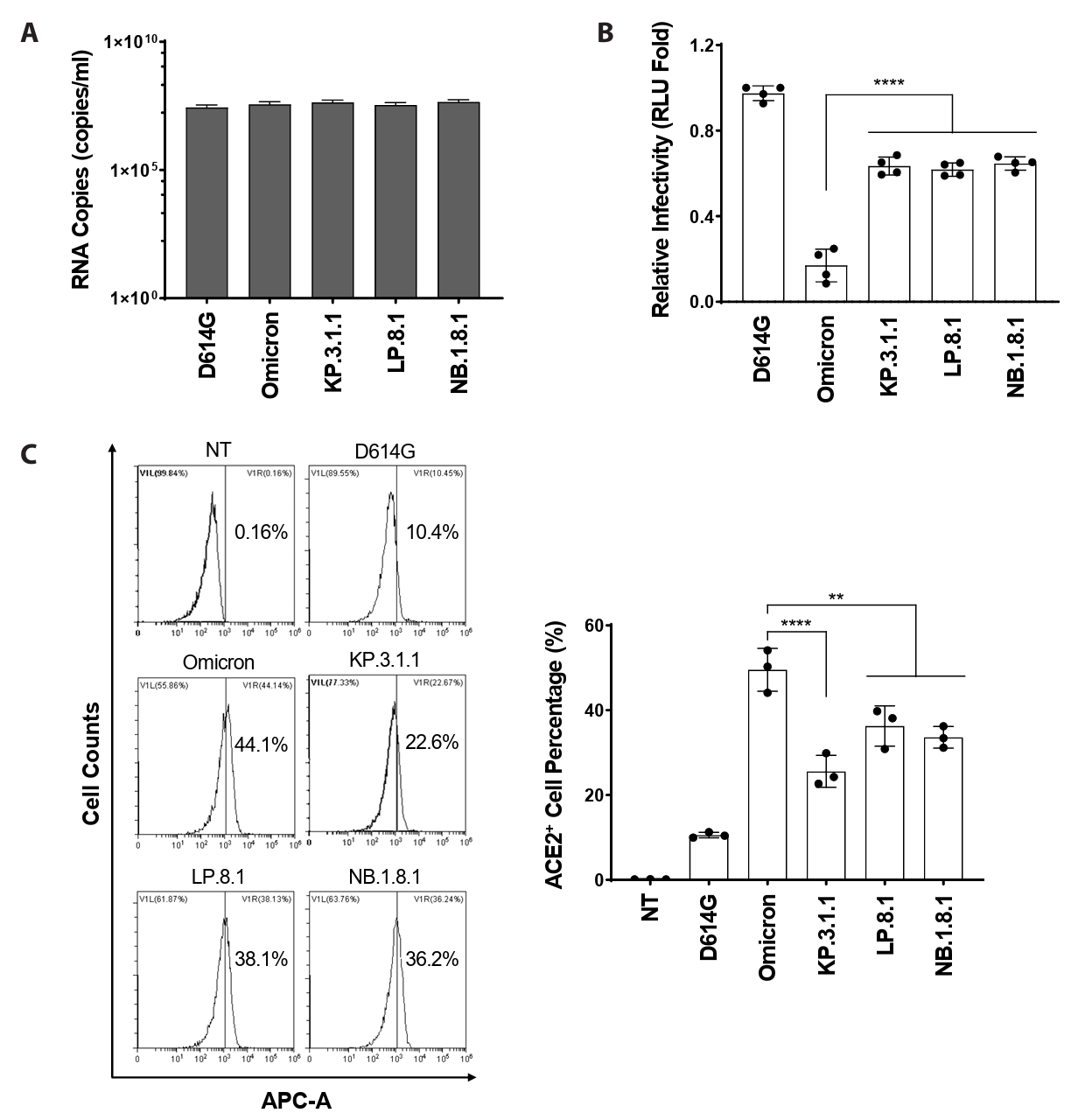
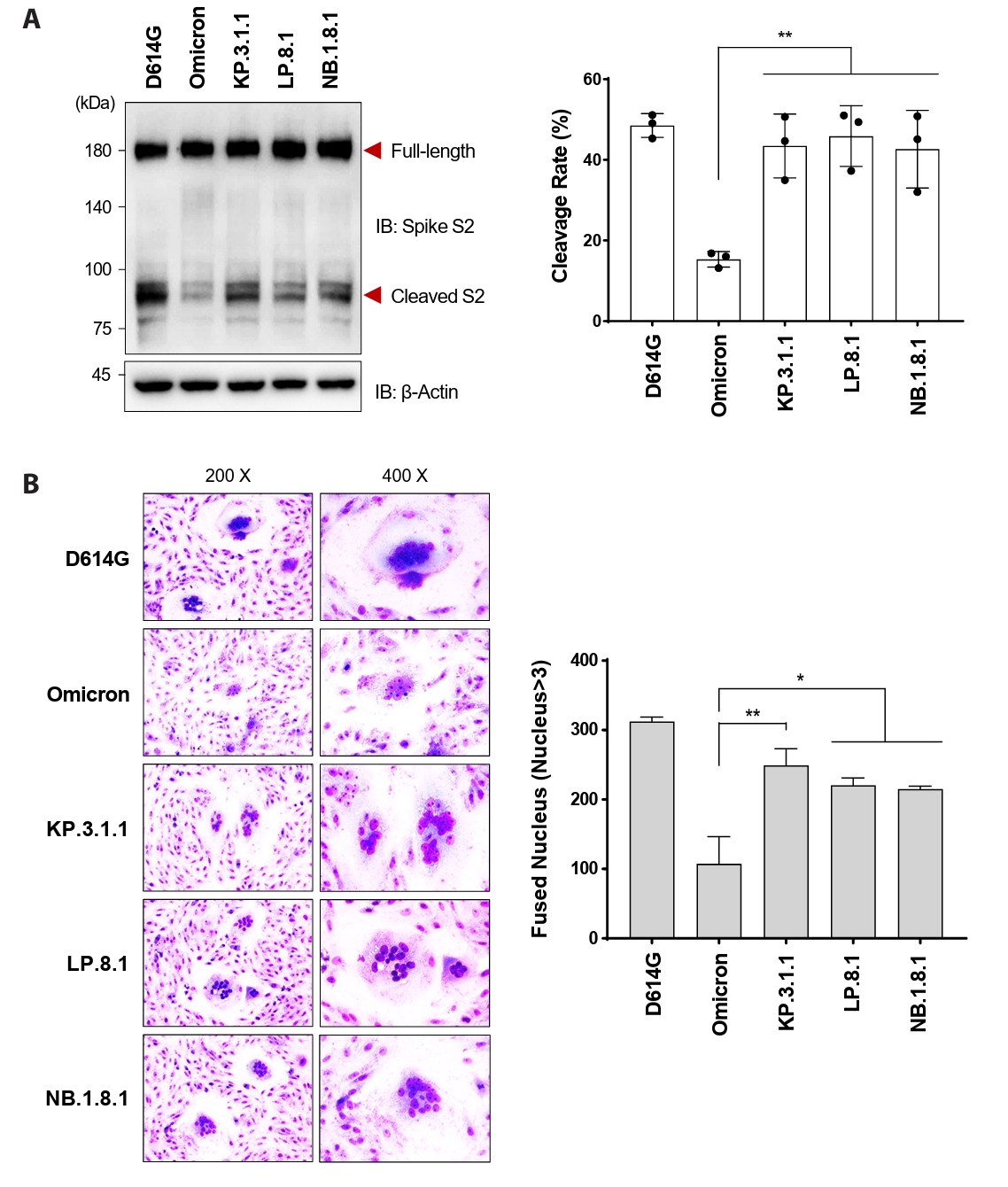
Fig. 1.
Fig. 2.
Fig. 3.
| Group | Primer | Sequence |
|---|---|---|
| Omicron | G339D F | 5'-CTGTGTCCATTTGATGAGGTGTTCAATG-3' |
| G339D R | 5'-CATTGAACACCTCATCAAATGGACACAG-3' | |
| K417N F | 5'-GACAAACAGGCAACATTGCTGACTACA-3' | |
| K417N R | 5'-TGTAGTCAGCAATGTTGCCTGTTTGTC-3' | |
| S371L F | 5'-TCTGTGCTCTACAACCTTGCCTCCTTCAGCAC-3' | |
| S371L R | 5'-GTGCTGAAGGAGGCAAGGTTGTAGAGCACAGA-3' | |
| S373P/S375F F | 5'-GCTCTACAACCTTGCCCCCTTCTTCACCTTCAAGTGTTAT-3' | |
| S373P/S375F R | 5'-ATAACACTTGAAGGTGAAGAAGGGGGCAAGGTTGTAGAGC-3' | |
| N440K F | 5'-GGAACAGCAACAAACTGGACAGCAAGG-3' | |
| N440K R | 5'-CCTTGCTGTCCAGTTTGTTGCTGTTCC-3' | |
| G446S F | 5'-GGACAGCAAGGTGAGCGGCAACTACAACT-3' | |
| G446S R | 5'-AGTTGTAGTTGCCGCTCACCTTGCTGTCC-3' | |
| S477N/T478K F | 5'-TACCAGGCTGGCAACAAACCATGTAATGGA-3' | |
| S477N/T478K R | 5'-TCCATTACATGGTTTGTTGCCAGCCTGGTA-3' | |
| E484A F | 5'-TGTAATGGAGTGGCGGGCTTCAACTGT-3' | |
| E484A R | 5'-ACAGTTGAAGCCCGCCACTCCATTACA-3' | |
| Q493R/G496S/Q498R F | 5'-TTACTTTCCACTCAAATCCTATAGCTTCCGACCAACCAATGGA-3' | |
| Q493R/G496S/Q498R R | 5'-TCCATTGGTTGGTCGGAAGCTATAGGATTTGAGTGGAAAGTAA-3' | |
| N501Y/Y505H F | 5'-CTTCCAACCAACCTATGGAGTGGGCCACCAACCATACAG-3' | |
| N501Y/Y505H R | 5'-CTGTATGGTTGGTGGCCCACTCCATAGGTTGGTTGGAAG-3' | |
| KP.3.1.1, LP.8.1, NB.1.8.1 | I332V F | 5′-GAGGTTTCCAAACGTCACCAACCTGTG-3′ |
| I332V R | 5'-CACAGGTTGGTGACGTTTGGAAACCTC-3' | |
| D339H F | 5'-CCTGTGTCCATTTCATGAGGTGTTCAA-3' | |
| D339H R | 5'-TTGAACACCTCATGAAATGGACACAGG-3' | |
| K356T F | 5'-ATGCCTGGAACAGGACGAGGATTAGCAACTG-3' | |
| K356T R | 5'-CAGTTGCTAATCCTCGTCCTGTTCCAGGCAT-3' | |
| L371F F | 5'-TGTGCTCTACAACTTTGCCCCCTTCTT-3' | |
| L371F R | 5'-AAGAAGGGGGCAAAGTTGTAGAGCACA-3' | |
| T376A F | 5'-TGCCCCCTTCTTCGCCTTCAAGTGTTA-3' | |
| T376A R | 5'-TAACACTTGAAGGCGAAGAAGGGGGCA-3' | |
| R403K-D405N F | 5'-TCCTTTGTGATTAAGGGAAATGAGGTGAGACA-3' | |
| R403K-D405N R | 5'-TGTCTCACCTCATTTCCCTTAATCACAAAGGA-3' | |
| R408S F | 5'-GAAATGAGGTGAGTCAGATTGCCCCTG-3' | |
| R408S R | 5'-CAGGGGCAATCTGACTCACCTCATTTC-3' | |
| N450D-L452W F | 5'-CAGCGGCAACTACGACTACTGGTACAGACTGTTCA-3' | |
| N450D-L452W R | 5'-TGAACAGTCTGTACCAGTAGTCGTAGTTGCCGCTG-3' | |
| L455S-F456L F | 5'-CTACTGGTACAGATCGTTGAGGAAGAGCAACC-3' | |
| L455S-F456L R | 5'-GGTTGCTCTTCCTCAACGATCTGTACCAGTAG-3' | |
| N460K F | 5'-TGAGGAAGAGCAAGCTGAAACCATTTG-3' | |
| N460K R | 5'-CAAATGGTTTCAGCTTGCTCTTCCTCA-3' | |
| N481K-V483Δ F | 5'-ACAAACCATGTAAGGGAAAGGCGGGCTTCAAC-3' | |
| N481K-V483Δ R | 5'-GTTGAAGCCCGCCTTTCCCTTACATGGTTTGT-3' | |
| A484K-F486P F | 5'-AACCATGTAAGGGAAAGGGCCCCAACTGTTACTTTC-3' | |
| A484K-F486P R | 5'-GAAAGTAACAGTTGGGGCCCTTTCCCTTACATGGTT-3' | |
| R493E F | 5'-TTACTTTCCACTCGAATCCTATGGCTT-3' | |
| R493E R | 5'-AAGCCATAGGATTCGAGTGGAAAGTAA-3' | |
| S496G F | 5'-ACTCCAATCCTATGGCTTCCGACCAAC-3' | |
| S496G R | 5'-GTTGGTCGGAAGCCATAGGATTGGAGT-3' | |
| KP.3.1.1, NB.1.8.1 | V445H F | 5'-ACTGGACAGCAAGCACAGCGGCAACTACA-3' |
| V445H R | 5'-TGTAGTTGCCGCTGTGCTTGCTGTCCAGT-3' | |
| LP.8.1 | R346T F | 5'-TTCAATGCCACCACGTTTGCCTCTGTC-3' |
| R346T R | 5'-GACAGAGGCAAACGTGGTGGCATTGAA-3' | |
| V445R F | 5'-ACTGGACAGCAAGCGCAGCGGCAACTACG-3' | |
| V445R R | 5'-CGTAGTTGCCGCTGCGCTTGCTGTCCAGT-3' | |
| NB.1.8.1 | A435S F | 5'-AGGCTGTGTGATTTCCTGGAACAGCAA-3' |
| A435S R | 5'-TTGCTGTTCCAGGAAATCACACAGCCT-3' | |
| K478I F | 5'-CAGGCTGGCAACATACCATGTAAGGGA-3' | |
| K478I R | 5'-TCCCTTACATGGTATGTTGCCAGCCTG-3' |
Table 1.
TOP
 MSK
MSK
 ePub Link
ePub Link Cite this Article
Cite this Article




