- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Volume 64(3); 2026 > Review
-
Minireview
Recent trends in dual-acting hybrid antibiotics and combination therapies against Gram-negative pathogens - Ji Eun Son, Umji Choi, Gyubin Han, Jeongho Lee, Chang-Ro Lee*
-
Journal of Microbiology 2026;64(3):e2601004.
DOI: https://doi.org/10.71150/jm.2601004
Published online: March 31, 2026
Department of Biological Sciences, Myongji University, Yongin 17058, Republic of Korea
- *Correspondence Chang-Ro Lee crlee@mju.ac.kr
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
- 1,095 Views
- 56 Download
- 1 Crossref
ABSTRACT
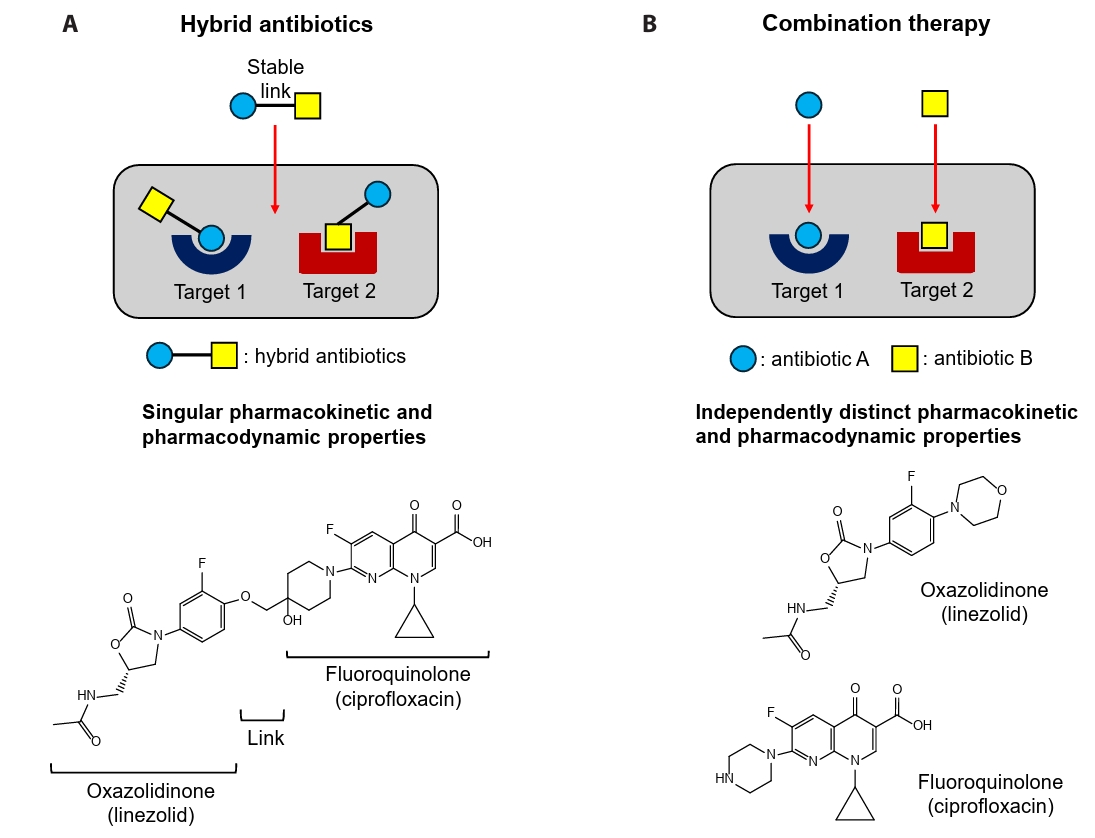
- Antibiotic resistance poses a serious challenge to public health worldwide; however, the development of new antibiotic classes for combating bacterial infections, especially those caused by Gram-negative pathogens, has slowed in recent years. Dual-acting hybrid antibiotics with a metabolically non-cleavable covalent bond represent an emerging strategy for developing novel antibiotic classes to overcome antibiotic resistance. The covalent connection between two antibiotics results in a fixed pharmacokinetic profile of a single molecule and can impede bacterial efflux. However, as most antibiotics do not have membrane-destabilizing activity, the resulting increase in molecular weight by connection of two antibiotics could limit their activity against Gram-negative bacteria, whose outer membrane forms a strong barrier blocking the penetration of high-molecular weight antibiotics. Here, we review recent developments in dual-acting hybrid antibiotics targeting Gram-negative bacteria, with a focus on their antibacterial efficacy. We also discuss combination therapy strategies in which the underlying molecular mechanisms of synergy have been characterized. Finally, we outline future directions for the rational design of hybrid antibiotics against Gram-negative pathogens.
Introduction
Quinolone-Containing Antibiotic Hybrids
Other Antibiotic Hybrids
Mechanisms Underlying Synergistic Effects of Combination Therapy
Conclusion
Acknowledgments
This work was supported by a research grant from Basic Science Research Program through the National Research Foundation of Korea funded by the Ministry of Education (NRF-RS-2023-00246684) and the Regional Innovation System & Education (RISE) program through the Gyeonggi RISE Center, funded by the Ministry of Education and the Gyeonggi-do (2025-RISE-09-A15).
| Hybrid antibiotics | Target pathogens | MIC (mg/ml) | MIC in the resistant mutant strains | Development stage | Comments | Reference |
|---|---|---|---|---|---|---|
| Ciprofloxacin–cephalosporin | E. coli | 0.18 | NDM-expressing E. coli: 0.05 | In vitro bacteria assay | Evans et al. (2019) | |
| Ciprofloxacin–neomycin | E. coli | 0.75–6 | Neomycin-resistant E. coli: 6 | In vitro bacteria assay | Pokrovskaya et al. (2009) | |
| Ciprofloxacin–kanamycin | E. coli | 0.28–12 | Kanamycin-resistant E. coli: 0.1–3.0 | In vitro bacteria assay | The hybrids had significantly lower resistance frequencies in E. coli than each drug alone or their combination mixture. | Shavit et al. (2017) |
| Ciprofloxacin–tobramycin | P. aeruginosa | 4 | ND | In vitro bacteria assay | The compound showed strong synergy with fluoroquinolone against multidrug-resistant P. aeruginosa, A. baumannii, and K. pneumoniae. | Gorityala et al. (2016) |
| Ciprofloxacin–tobramycin | P. aeruginosa | 64 | ND | In vitro bacteria assay | The compound showed strong synergy with ciprofloxacin, levofloxacin, and moxifloxacin against multidrug-resistant P. aeruginosa. | Dhiman et al. (2023) |
| Ciprofloxacin–erythromycin | H. influenzae | 4 | ND | In vitro bacteria assay | Hutinec et al. (2010) | |
| Ciprofloxacin–erythromycin | H. influenzae, M. catarrhalis | H. influenzae: 2 | ND | In vitro bacteria assay | Ma et al. (2019) | |
| M. catarrhalis: 0.5 | ||||||
| Quinolone–telithromycin | H. influenzae, E. coli, K. pneumoniae | H. influenzae: 2 | ND | In vitro bacteria assay | Fan et al. (2020) | |
| E. coli: 8 | ||||||
| K. pneumoniae: 16 | ||||||
| Quinolone–telithromycin | H. influenzae, M. pneumoniae | H. influenzae: 4 M. pneumoniae: 0.008 | Erythromycin-resistant M. pneumoniae: 8 | In vitro bacteria assay | Liu et al. (2022) | |
| Ciprofloxacin–oxazolidinone | H. influenzae, E. coli | H. influenzae: 0.06 | ND | In vitro bacteria assay | Hubschwerlen et al. (2003a) | |
| E. coli: 1 | ||||||
| Ciprofloxacin–oxazolidinone | H. influenzae, E. coli | H. influenzae: 0.03 | ND | In vitro bacteria assay | Hubschwerlen et al. (2003b) | |
| E. coli: 0.5 | ||||||
| Fluoroquinolone–oxazolidinone | H. influenzae, E. coli, P. aeruginosa | H. influenzae: 0.016 | ND | In vitro bacteria assay | Liu et al. (2019) | |
| E. coli: 1 | ||||||
| P. aeruginosa: 16 | ||||||
| Fluoroquinolone–oxazolidinone (cadazolid) | E. coli | E. coli: 8 | ND | Discontinued in 2018 after two Phase III trials | Scaiola et al. (2019) | |
| Fluoroquinolone–rifampicin | H. influenzae, E. coli, N. gonorrhoeae, H. pylori | H. influenzae: 0.012 | ND | Phase II for acute bacterial skin and skin structure infection | Yuan et al. (2020) | |
| E. coli: 0.25 | ||||||
| N. gonorrhoeae: 0.06 | ||||||
| H. pylori: 0.12 | ||||||
| Fluoroquinolone–kanglemycin | E. coli | 4 | ND | In vitro bacteria assay | Peek et al. (2022) | |
| Ciprofloxacin–sulfonamide | E. coli | 0.013 | ND | In vitro bacteria assay | Ibrahim et al. (2022) | |
| Quinolone–trimethoprim | P. aeruginosa, K. pneumoniae | P. aeruginosa: 8–16 | ND | In vitro bacteria assay | Karoli et al. (2012) | |
| K. pneumoniae: 8 | ||||||
| Quinolone–metronidazole | P. aeruginosa, E. coli | P. aeruginosa: 2 | ND | In vitro bacteria assay | Several compounds showed appropriate ranges to pharmacokinetic behaviors and no obvious toxicity to human hepatocyte cells. | Cui et al. (2014) |
| E. coli: 0.5 | ||||||
| Quinolone–metronidazole | P. aeruginosa, E. coli | P. aeruginosa: 0.25 | ND | In vitro bacteria assay | Zhang et al. (2015) | |
| E. coli: 0.25 | ||||||
| Quinolone–fluoroquinolone | P. aeruginosa | 312.5 | ND | In vitro bacteria assay | Panda et al. (2015) | |
| Ciprofloxacin dimer | E. coli, P. aeruginosa | P. aeruginosa: 0.125 | ND | In vitro bacteria assay | Ross et al. (2015) | |
| E. coli: 0.03 | ||||||
| Polymyxin B3–tobramycin- | E. coli, K. pneumoniae, A. baumannii, P. aeruginosa | P. aeruginosa: 2 | Carbapenem-resistant MDR/XDR P. aeruginosa: 2–16 | In vitro bacteria assay | The compound showed strong synergy with minocycline, rifampicin, and vancomycin against multidrug-resistant P. aeruginosa. | Domalaon et al. (2017) |
| A. baumannii: 16 | Colistin-resistant P. aeruginosa: 4–32 | |||||
| E. coli: 8 | ||||||
| K. pneumoniae: 128 | ||||||
| Polymyxin E–vancomycin (vancomyxin) | K. pneumoniae, A. baumannii, P. aeruginosa | K. pneumoniae: 8 | ND | In vitro bacteria assay | van Groesen et al. (2021) | |
| P. aeruginosa: 16 | ||||||
| A. baumannii: 16 | ||||||
| Vancomycin–nisin | K. pneumoniae, M. catarrhalis | K. pneumoniae: 16 | ND | In vitro bacteria assay | Arnusch et al. (2008) | |
| M. catarrhalis: 16 | ||||||
| Rifamycin–nitroimidazole (TNP-2198) | H. pylori | H. pylori: 0.004 | Rifamycin-resistant H. pylori: 0.5 | Phase II for H. pylori, vaginosis and CDAD ongoing since 2021/2022 | Ma et al. (2022) | |
| Rifamycin- and metronidazole-resistant H. pylori: 0.5 | ||||||
| Neomycin B–triclosan | E. coli, K. pneumoniae, A. baumannii, P. aeruginosa | E. coli: 0.25 | ND | In vitro bacteria assay | Findlay et al. (2012) | |
| K. pneumoniae: 1 | ||||||
| P. aeruginosa: 64 | ||||||
| A. baumannii: 8 | ||||||
| Sisomicin–gentamicin | E. coli, K. pneumoniae, P. aeruginosa | E. coli: 0.5 | ND | In vitro bacteria assay | Hanessian et al. (2011) | |
| K. pneumoniae: 0.25 | ||||||
| P. aeruginosa: 0.5 | ||||||
| Linezolid–sparsomycin | H. influenzae | 4 | ND | In vitro bacteria assay | Zhou et al. (2008) | |
| Oxazolidinone–cephalosporin–siderophore | E. coli, A. baumannii, P. aeruginosa | E. coli: 0.025 | ADC-1-expressing A. baumannii: 6 | In vitro bacteria assay | Liu et al. (2018) | |
| A. baumannii: 0.4 | ||||||
| P. aeruginosa: 0.4 | ||||||
| Mupirocin–holomycin | E. coli | 64 | ND | In vitro bacteria assay | Johnson et al. (2024) |
| Combination antibiotics | Target pathogens | FICI | Synergy mechanisms | Clinical uses | Comments | Reference |
|---|---|---|---|---|---|---|
| Trimethoprim and sulfamethoxazole | E. coli | 0.31 | Mutual potentiation (Sulfamethoxazole potentiates trimethoprim by limiting de novo dihydrofolate production and trimethoprim potentiates sulfamethoxazole activity through inhibition of dihydropterin pyrophosphate synthesis) | Clinical use | Minato et al. (2018) | |
| Amikacin and nitrofurantoin | E. coli, K. pneumoniae | E. coli: 0.375 | Amikacin induces bacterial envelope stress by introducing mistranslated proteins, thereby constitutively activating the CpxAR two‐component system. The activation of Cpx signaling stimulates the expression of bacterial major nitroreductases (NfsAB). Nitroreductases overexpression generates considerable quantity of lethal reactive intermediates via nitroreduction and promotes the prodrug activation of nitrofurantoin. | No | Ren et al. (2023) | |
| K. pneumoniae: 0.5 | ||||||
| Nalidixic acid and tetracycline | Multidrug-resistant A. baumannii and E. coli | Multidrug-resistant A. baumannii: 0.1875–0.5 | Enhanced uptake and reduced efflux of tetracycline by nalidixic acid explain the basis of synergy between nalidixic acid and tetracycline. | No | Nalidixic acid and tetracycline combination did not display synergy against susceptible A. baumannii and E. coli isolates. | Gaurav et al. (2021) |
| Flavomycin and colistin | E. coli, mcr-1-positive E. coli | E. coli: 0.19–0.49 | The synergy is manifested as an augmented penetration of the E. coli OM by colistin, leading to increased intracellular accumulation of flavomycin and enhanced cell killing thereafter. | No | Huang et al. (2024) | |
| Novobiocin and colistin | Colistin-resistant A. baumannii and K. pneumoniae | Colistin-resistant A. baumannii: 0.129 | Novobiocin binds and activates the ATPase LptB that powers lipopolysaccharide transport, which enhances polymyxin activity. | No | May et al. (2017); Mandler et al. (2018); Mattingly et al. (2020) | |
| Colistin-resistant K. pneumoniae: 0.012 | ||||||
| Hygromycin A and macrolides | B. burgdorferi | 0.5 | Hygromycin A cooperatively binds ribosomes with nascent peptide exit tunnel-targeting macrolides and slows down their dissociation, which potentiates macrolide’s efficacy in bacterial growth inhibition and early killing. | No | Chen et al. (2023) |
- Antimicrobial Resistance Collaborators. 2022. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet. 399: 629–655. ArticlePubMedPMC
- Arnusch CJ, Bonvin AM, Verel AM, Jansen WT, Liskamp RM, et al. 2008. The vancomycin-nisin(1–12) hybrid restores activity against vancomycin resistant Enterococci. Biochemistry. 47: 12661–12663. ArticlePubMed
- Bozdogan B, Appelbaum PC. 2004. Oxazolidinones: activity, mode of action, and mechanism of resistance. Int J Antimicrob Agents. 23: 113–119. ArticlePubMed
- Brogden RN, Carmine AA, Heel RC, Speight TM, Avery GS. 1982. Trimethoprim: a review of its antibacterial activity, pharmacokinetics and therapeutic use in urinary tract infections. Drugs. 23: 405–430. ArticlePubMed
- Bulitta JB, Ly NS, Landersdorfer CB, Wanigaratne NA, Velkov T, et al. 2015. Two mechanisms of killing of Pseudomonas aeruginosa by tobramycin assessed at multiple inocula via mechanism-based modeling. Antimicrob Agents Chemother. 59: 2315–2327. ArticlePubMedPMCLink
- Butler MS, Blaskovich MA, Cooper MA. 2017. Antibiotics in the clinical pipeline at the end of 2015. J Antibiot (Tokyo). 70: 3–24. ArticlePubMedPDF
- Calvori C, Frontali L, Leoni L, Tecce G. 1965. Effect of rifamycin on protein synthesis. Nature. 207: 417–418. ArticlePubMedPDF
- Chen CW, Leimer N, Syroegin EA, Dunand C, Bulman ZP, et al. 2023. Structural insights into the mechanism of overcoming Erm-mediated resistance by macrolides acting together with hygromycin-A. Nat Commun. 14: 4196.ArticlePubMedPMCPDF
- Cui SF, Peng LP, Zhang HZ, Rasheed S, Vijaya Kumar K, et al. 2014. Novel hybrids of metronidazole and quinolones: synthesis, bioactive evaluation, cytotoxicity, preliminary antimicrobial mechanism and effect of metal ions on their transportation by human serum albumin. Eur J Med Chem. 86: 318–334. ArticlePubMed
- Dhiman S, Ramirez D, Li Y, Kumar A, Arthur G, et al. 2023. Chimeric tobramycin-based adjuvant TOB-TOB-CIP potentiates fluoroquinolone and β-lactam antibiotics against multidrug-resistant Pseudomonas aeruginosa. ACS Infect Dis. 9: 864–885. ArticlePubMedLink
- Domalaon R, Yang X, Lyu Y, Zhanel GG, Schweizer F. 2017. Polymyxin B3-tobramycin hybrids with Pseudomonas aeruginosa-selective antibacterial activity and strong potentiation of rifampicin, minocycline, and vancomycin. ACS Infect Dis. 3: 941–954. ArticlePubMed
- Evans LE, Krishna A, Ma Y, Webb TE, Marshall DC, et al. 2019. Exploitation of antibiotic resistance as a novel drug target: development of a β-lactamase-activated antibacterial prodrug. J Med Chem. 62: 4411–4425. ArticlePubMedPMC
- Fan BZ, Hiasa H, Lv W, Brody S, Yang ZY, et al. 2020. Design, synthesis and structure-activity relationships of novel 15-membered macrolides: Quinolone/quinoline-containing sidechains tethered to the C-6 position of azithromycin acylides. Eur J Med Chem. 193: 112222.ArticlePubMed
- Findlay B, Zhanel GG, Schweizer F. 2012. Neomycin-phenolic conjugates: polycationic amphiphiles with broad-spectrum antibacterial activity, low hemolytic activity and weak serum protein binding. Bioorg Med Chem Lett. 22: 1499–1503. ArticlePubMed
- Gaurav A, Gupta V, Shrivastava SK, Pathania R. 2021. Mechanistic insights into synergy between nalidixic acid and tetracycline against clinical isolates of Acinetobacter baumannii and Escherichia coli. Commun Biol. 4: 542.ArticlePubMedPMCPDF
- Gorityala BK, Guchhait G, Fernando DM, Deo S, McKenna SA, et al. 2016. Adjuvants based on hybrid antibiotics overcome resistance in Pseudomonas aeruginosa and enhance fluoroquinolone efficacy. Angew Chem Int Ed Engl. 55: 555–559. ArticlePubMedLink
- Hanessian S, Maianti JP, Matias RD, Feeney LA, Armstrong ES. 2011. Hybrid aminoglycoside antibiotics via Tsuji palladium-catalyzed allylic deoxygenation. Org Lett. 13: 6476–6479. ArticlePubMed
- Herzog IM, Green KD, Berkov-Zrihen Y, Feldman M, Vidavski RR, et al. 2012. 6''-Thioether tobramycin analogues: towards selective targeting of bacterial membranes. Angew Chem Int Ed Engl. 51: 5652–5656. ArticlePubMedPMC
- Huang Y, Zhu Y, Yue HY, Liu YY, Deng LM, et al. 2024. Flavomycin restores colistin susceptibility in multidrug-resistant Gram-negative bacteria. mSystems. 9: e00109-24.ArticlePubMedPMCLink
- Hubschwerlen C, Specklin JL, Baeschlin DK, Borer Y, Haefeli S, et al. 2003a. Structure-activity relationship in the oxazolidinone-quinolone hybrid series: influence of the central spacer on the antibacterial activity and the mode of action. Bioorg Med Chem Lett. 13: 4229–4233. Article
- Hubschwerlen C, Specklin JL, Sigwalt C, Schroeder S, Locher HH. 2003b. Design, synthesis and biological evaluation of oxazolidinone-quinolone hybrids. Bioorg Med Chem. 11: 2313–2319. Article
- Hutinec A, Derek M, Lazarevski G, Sunjic V, Paljetak HC, et al. 2010. Novel 8a-aza-8a-homoerythromycin—4''-(3-substituted-amino)propionates with broad spectrum antibacterial activity. Bioorg Med Chem Lett. 20: 3244–3249. ArticlePubMed
- Ibrahim NM, Fahim SH, Hassan M, Farag AE, Georgey HH. 2022. Design and synthesis of ciprofloxacin-sulfonamide hybrids to manipulate ciprofloxacin pharmacological qualities: Potency and side effects. Eur J Med Chem. 228: 114021.ArticlePubMed
- Johnson RM, Li K, Chen X, Morgan GL, Aube J, et al. 2024. The hybrid antibiotic thiomarinol A overcomes intrinsic resistance in Escherichia coli using a privileged dithiolopyrrolone moiety. ACS Infect Dis. 10: 582–593. ArticlePubMedPMCLink
- Karoli T, Mamidyala SK, Zuegg J, Fry SR, Tee EH, et al. 2012. Structure aided design of chimeric antibiotics. Bioorg Med Chem Lett. 22: 2428–2433. ArticlePubMed
- Khoshnood S, Heidary M, Asadi A, Soleimani S, Motahar M, et al. 2019. A review on mechanism of action, resistance, synergism, and clinical implications of mupirocin against Staphylococcus aureus. Biomed Pharmacother. 109: 1809–1818. ArticlePubMed
- Koh AJJ, Thombare V, Hussein M, Rao GG, Li J, et al. 2023. Bifunctional antibiotic hybrids: A review of clinical candidates. Front Pharmacol. 14: 1158152.ArticlePubMedPMC
- Kratky M, Vinsova J, Volkova M, Buchta V, Trejtnar F, et al. 2012. Antimicrobial activity of sulfonamides containing 5-chloro-2-hydroxybenzaldehyde and 5-chloro-2-hydroxybenzoic acid scaffold. Eur J Med Chem. 50: 433–440. ArticlePubMed
- Lawson DM, Stevenson CE. 2012. Structural and functional dissection of aminocoumarin antibiotic biosynthesis: a review. J Struct Funct Genomics. 13: 125–133. ArticlePubMedPDF
- Lee CR, Cho IH, Jeong BC, Lee SH. 2013. Strategies to minimize antibiotic resistance. Int J Environ Res Public Health. 10: 4274–4305. ArticlePubMedPMC
- Leimer N, Wu X, Imai Y, Morrissette M, Pitt N, et al. 2021. A selective antibiotic for Lyme disease. Cell. 184: 5405–5418.e16. ArticlePubMedPMC
- Leitsch D. 2019. A review on metronidazole: an old warhorse in antimicrobial chemotherapy. Parasitology. 146: 1167–1178. ArticlePubMed
- Li J, Nation RL, Turnidge JD, Milne RW, Coulthard K, et al. 2006. Colistin: the re-emerging antibiotic for multidrug-resistant Gram-negative bacterial infections. Lancet Infect Dis. 6: 589–601. ArticlePubMed
- Liras P. 2014. Holomycin, a dithiolopyrrolone compound produced by Streptomyces clavuligerus. Appl Microbiol Biotechnol. 98: 1023–1030. ArticlePubMedPDF
- Liu XP, Lv W, Zhao F, Ding J, Zhang JR, et al. 2022. Design and synthesis of novel macrolones bridged with linkers from 11,12-positions of macrolides. Bioorg Med Chem Lett. 68: 128761.ArticlePubMed
- Liu R, Miller PA, Vakulenko SB, Stewart NK, Boggess WC, et al. 2018. A synthetic dual drug sideromycin induces Gram-negative bacteria to commit suicide with a Gram-positive antibiotic. J Med Chem. 61: 3845–3854. ArticlePubMed
- Liu L, Shao L, Li J, Cui H, Li B, et al. 2019. Synthesis, antibacterial activities, mode of action and acute toxicity studies of new oxazolidinone-fluoroquinolone hybrids. Molecules. 24: 1641.ArticlePubMedPMC
- Ma Z, He S, Yuan Y, Zhuang Z, Liu Y, et al. 2022. Design, synthesis, and characterization of TNP-2198, a dual-targeted rifamycin-nitroimidazole conjugate with potent activity against microaerophilic and anaerobic bacterial pathogens. J Med Chem. 65: 4481–4495. ArticlePubMedPMC
- Ma CX, Lv W, Li YX, Fan BZ, Han X, et al. 2019. Design, synthesis and structure-activity relationships of novel macrolones: hybrids of 2-fluoro 9-oxime ketolides and carbamoyl quinolones with highly improved activity against resistant pathogens. Eur J Med Chem. 169: 1–20. ArticlePubMed
- Mahdizade Ari M, Dashtbin S, Ghasemi F, Shahroodian S, Kiani P, et al. 2023. Nitrofurantoin: properties and potential in treatment of urinary tract infection: a narrative review. Front Cell Infect Microbiol. 13: 1148603.ArticlePubMedPMC
- Mandler MD, Baidin V, Lee J, Pahil KS, Owens TW, et al. 2018. Novobiocin enhances polymyxin activity by stimulating lipopolysaccharide transport. J Am Chem Soc. 140: 6749–6753. ArticlePubMedPMC
- Mattingly AE, Cox KE, Smith R, Melander RJ, Ernst RK, et al. 2020. Screening an established natural product library identifies secondary metabolites that potentiate conventional antibiotics. ACS Infect Dis. 6: 2629–2640. ArticlePubMedPMC
- May JM, Owens TW, Mandler MD, Simpson BW, Lazarus MB, et al. 2017. The antibiotic novobiocin binds and activates the ATPase that powers lipopolysaccharide transport. J Am Chem Soc. 139: 17221–17224. ArticlePubMedPMC
- Minato Y, Dawadi S, Kordus SL, Sivanandam A, Aldrich CC, et al. 2018. Mutual potentiation drives synergy between trimethoprim and sulfamethoxazole. Nat Commun. 9: 1003.ArticlePubMedPMCPDF
- Mingeot-Leclercq MP, Glupczynski Y, Tulkens PM. 1999. Aminoglycosides: activity and resistance. Antimicrob Agents Chemother. 43: 727–737. ArticlePubMedPMCLink
- Odds FC. 2003. Synergy, antagonism, and what the chequerboard puts between them. J Antimicrob Chemother. 52: 1.ArticlePubMed
- O'Shea R, Moser HE. 2008. Physicochemical properties of antibacterial compounds: implications for drug discovery. J Med Chem. 51: 2871–2878. ArticlePubMed
- Ostash B, Doud E, Fedorenko V. 2010. The molecular biology of moenomycins: towards novel antibiotics based on inhibition of bacterial peptidoglycan glycosyltransferases. Biol Chem. 391: 499–504. ArticlePubMed
- Ottenheijm HC, van den Broek LA, Ballesta JP, Zylicz Z. 1986. Chemical and biological aspects of sparsomycin, an antibiotic from Streptomyces. Prog Med Chem. 23: 219–268. ArticlePubMed
- Panda SS, Liaqat S, Girgis AS, Samir A, Hall CD, et al. 2015. Novel antibacterial active quinolone-fluoroquinolone conjugates and 2D-QSAR studies. Bioorg Med Chem Lett. 25: 3816–3821. ArticlePubMed
- Peek J, Koirala B, Brady SF. 2022. Synthesis and evaluation of dual-action kanglemycin-fluoroquinolone hybrid antibiotics. Bioorg Med Chem Lett. 57: 128484.ArticlePubMed
- Pokrovskaya V, Baasov T. 2010. Dual-acting hybrid antibiotics: a promising strategy to combat bacterial resistance. Expert Opin Drug Discov. 5: 883–902. ArticlePubMed
- Pokrovskaya V, Belakhov V, Hainrichson M, Yaron S, Baasov T. 2009. Design, synthesis, and evaluation of novel fluoroquinolone-aminoglycoside hybrid antibiotics. J Med Chem. 52: 2243–2254. ArticlePubMed
- Ren H, Zhong Z, Zhou S, Wei Y, Liang Y, et al. 2023. CpxA/R-controlled nitroreductase expression as target for combinatorial therapy against uropathogens by promoting reactive oxygen species generation. Adv Sci (Weinh). 10: 2300938. ArticlePubMedPMC
- Ross AG, Benton BM, Chin D, De Pascale G, Fuller J, et al. 2015. Synthesis of ciprofloxacin dimers for evaluation of bacterial permeability in atypical chemical space. Bioorg Med Chem Lett. 25: 3468–3475. ArticlePubMed
- Scaiola A, Leibundgut M, Boehringer D, Caspers P, Bur D, et al. 2019. Structural basis of translation inhibition by cadazolid, a novel quinoxolidinone antibiotic. Sci Rep. 9: 5634.ArticlePubMedPMCPDF
- Schalk IJ. 2025. Bacterial siderophores: diversity, uptake pathways and applications. Nat Rev Microbiol. 23: 24–40. ArticlePubMedPDF
- Schumacher A, Trittler R, Bohnert JA, Kummerer K, Pages JM, et al. 2007. Intracellular accumulation of linezolid in Escherichia coli, Citrobacter freundii and Enterobacter aerogenes: role of enhanced efflux pump activity and inactivation. J Antimicrob Chemother. 59: 1261–1264. ArticlePubMed
- Shavit M, Pokrovskaya V, Belakhov V, Baasov T. 2017. Covalently linked kanamycin-ciprofloxacin hybrid antibiotics as a tool to fight bacterial resistance. Bioorg Med Chem. 25: 2917–2925. ArticlePubMed
- Smilack JD. 1999. Trimethoprim-sulfamethoxazole. Mayo Clin Proc. 74: 730–734. ArticlePubMed
- Theuretzbacher U. 2020. Dual-mechanism antibiotics. Nat Microbiol. 5: 984–985. ArticlePubMedPDF
- Thomas CM, Nielsen KM. 2005. Mechanisms of, and barriers to, horizontal gene transfer between bacteria. Nat Rev Microbiol. 3: 711–721. ArticlePubMedPDF
- van Groesen E, Slingerland CJ, Innocenti P, Mihajlovic M, Masereeuw R, et al. 2021. Vancomyxins: vancomycin-polymyxin nonapeptide conjugates that retain anti-Gram-positive activity with enhanced potency against Gram-negative strains. ACS Infect Dis. 7: 2746–2754. ArticlePubMedPMCLink
- Vazquez-Laslop N, Mankin AS. 2018. How macrolide antibiotics work. Trends Biochem Sci. 43: 668–684. ArticlePubMedPMC
- Velkov T, Roberts KD, Nation RL, Thompson PE, Li J. 2013. Pharmacology of polymyxins: new insights into an 'old' class of antibiotics. Future Microbiol. 8: 711–724. ArticlePubMedPMC
- Yuan Y, Wang X, Xu X, Liu Y, Li C, et al. 2020. Evaluation of a dual-acting antibacterial agent, TNP-2092, on gut microbiota and potential application in the treatment of gastrointestinal and liver disorders. ACS Infect Dis. 6: 820–831. ArticlePubMed
- Zhanel GG, Dueck M, Hoban DJ, Vercaigne LM, Embil JM, et al. 2001. Review of macrolides and ketolides: focus on respiratory tract infections. Drugs. 61: 443–498. ArticlePubMed
- Zhang L, Kumar KV, Geng RX, Zhou CH. 2015. Design and biological evaluation of novel quinolone-based metronidazole derivatives as potent Cu2+ mediated DNA-targeting antibacterial agents. Bioorg Med Chem Lett. 25: 3699–3705. ArticlePubMed
- Zhou J, Bhattacharjee A, Chen S, Chen Y, Duffy E, et al. 2008. Design at the atomic level: design of biaryloxazolidinones as potent orally active antibiotics. Bioorg Med Chem Lett. 18: 6175–6178. ArticlePubMed
References
Supplementary Information
References
Citations

- Pioneering strategies for overcoming bacterial drug resistance
Byoung Sik Kim
Journal of Microbiology.2026; 64(3): e2603100. CrossRef
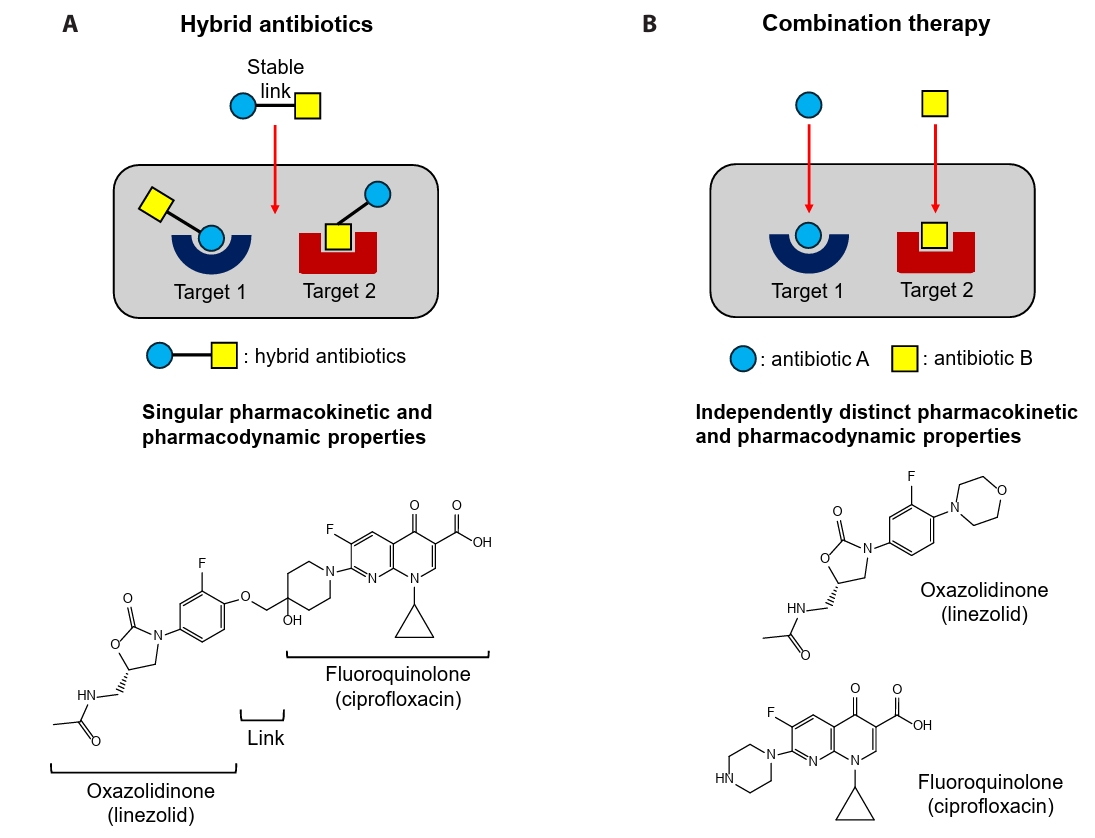
Fig. 1.
| Hybrid antibiotics | Target pathogens | MIC (mg/ml) | MIC in the resistant mutant strains | Development stage | Comments | Reference |
|---|---|---|---|---|---|---|
| Ciprofloxacin–cephalosporin | E. coli | 0.18 | NDM-expressing E. coli: 0.05 | In vitro bacteria assay | ||
| Ciprofloxacin–neomycin | E. coli | 0.75–6 | Neomycin-resistant E. coli: 6 | In vitro bacteria assay | ||
| Ciprofloxacin–kanamycin | E. coli | 0.28–12 | Kanamycin-resistant E. coli: 0.1–3.0 | In vitro bacteria assay | The hybrids had significantly lower resistance frequencies in E. coli than each drug alone or their combination mixture. | |
| Ciprofloxacin–tobramycin | P. aeruginosa | 4 | ND | In vitro bacteria assay | The compound showed strong synergy with fluoroquinolone against multidrug-resistant P. aeruginosa, A. baumannii, and K. pneumoniae. | |
| Ciprofloxacin–tobramycin | P. aeruginosa | 64 | ND | In vitro bacteria assay | The compound showed strong synergy with ciprofloxacin, levofloxacin, and moxifloxacin against multidrug-resistant P. aeruginosa. | |
| Ciprofloxacin–erythromycin | H. influenzae | 4 | ND | In vitro bacteria assay | ||
| Ciprofloxacin–erythromycin | H. influenzae, M. catarrhalis | H. influenzae: 2 | ND | In vitro bacteria assay | ||
| M. catarrhalis: 0.5 | ||||||
| Quinolone–telithromycin | H. influenzae, E. coli, K. pneumoniae | H. influenzae: 2 | ND | In vitro bacteria assay | ||
| E. coli: 8 | ||||||
| K. pneumoniae: 16 | ||||||
| Quinolone–telithromycin | H. influenzae, M. pneumoniae | H. influenzae: 4 M. pneumoniae: 0.008 | Erythromycin-resistant M. pneumoniae: 8 | In vitro bacteria assay | ||
| Ciprofloxacin–oxazolidinone | H. influenzae, E. coli | H. influenzae: 0.06 | ND | In vitro bacteria assay | ||
| E. coli: 1 | ||||||
| Ciprofloxacin–oxazolidinone | H. influenzae, E. coli | H. influenzae: 0.03 | ND | In vitro bacteria assay | ||
| E. coli: 0.5 | ||||||
| Fluoroquinolone–oxazolidinone | H. influenzae, E. coli, P. aeruginosa | H. influenzae: 0.016 | ND | In vitro bacteria assay | ||
| E. coli: 1 | ||||||
| P. aeruginosa: 16 | ||||||
| Fluoroquinolone–oxazolidinone (cadazolid) | E. coli | E. coli: 8 | ND | Discontinued in 2018 after two Phase III trials | ||
| Fluoroquinolone–rifampicin | H. influenzae, E. coli, N. gonorrhoeae, H. pylori | H. influenzae: 0.012 | ND | Phase II for acute bacterial skin and skin structure infection | ||
| E. coli: 0.25 | ||||||
| N. gonorrhoeae: 0.06 | ||||||
| H. pylori: 0.12 | ||||||
| Fluoroquinolone–kanglemycin | E. coli | 4 | ND | In vitro bacteria assay | ||
| Ciprofloxacin–sulfonamide | E. coli | 0.013 | ND | In vitro bacteria assay | ||
| Quinolone–trimethoprim | P. aeruginosa, K. pneumoniae | P. aeruginosa: 8–16 | ND | In vitro bacteria assay | ||
| K. pneumoniae: 8 | ||||||
| Quinolone–metronidazole | P. aeruginosa, E. coli | P. aeruginosa: 2 | ND | In vitro bacteria assay | Several compounds showed appropriate ranges to pharmacokinetic behaviors and no obvious toxicity to human hepatocyte cells. | |
| E. coli: 0.5 | ||||||
| Quinolone–metronidazole | P. aeruginosa, E. coli | P. aeruginosa: 0.25 | ND | In vitro bacteria assay | ||
| E. coli: 0.25 | ||||||
| Quinolone–fluoroquinolone | P. aeruginosa | 312.5 | ND | In vitro bacteria assay | ||
| Ciprofloxacin dimer | E. coli, P. aeruginosa | P. aeruginosa: 0.125 | ND | In vitro bacteria assay | ||
| E. coli: 0.03 | ||||||
| Polymyxin B3–tobramycin- | E. coli, K. pneumoniae, A. baumannii, P. aeruginosa | P. aeruginosa: 2 | Carbapenem-resistant MDR/XDR P. aeruginosa: 2–16 | In vitro bacteria assay | The compound showed strong synergy with minocycline, rifampicin, and vancomycin against multidrug-resistant P. aeruginosa. | |
| A. baumannii: 16 | Colistin-resistant P. aeruginosa: 4–32 | |||||
| E. coli: 8 | ||||||
| K. pneumoniae: 128 | ||||||
| Polymyxin E–vancomycin (vancomyxin) | K. pneumoniae, A. baumannii, P. aeruginosa | K. pneumoniae: 8 | ND | In vitro bacteria assay | ||
| P. aeruginosa: 16 | ||||||
| A. baumannii: 16 | ||||||
| Vancomycin–nisin | K. pneumoniae, M. catarrhalis | K. pneumoniae: 16 | ND | In vitro bacteria assay | ||
| M. catarrhalis: 16 | ||||||
| Rifamycin–nitroimidazole (TNP-2198) | H. pylori | H. pylori: 0.004 | Rifamycin-resistant H. pylori: 0.5 | Phase II for H. pylori, vaginosis and CDAD ongoing since 2021/2022 | ||
| Rifamycin- and metronidazole-resistant H. pylori: 0.5 | ||||||
| Neomycin B–triclosan | E. coli, K. pneumoniae, A. baumannii, P. aeruginosa | E. coli: 0.25 | ND | In vitro bacteria assay | ||
| K. pneumoniae: 1 | ||||||
| P. aeruginosa: 64 | ||||||
| A. baumannii: 8 | ||||||
| Sisomicin–gentamicin | E. coli, K. pneumoniae, P. aeruginosa | E. coli: 0.5 | ND | In vitro bacteria assay | ||
| K. pneumoniae: 0.25 | ||||||
| P. aeruginosa: 0.5 | ||||||
| Linezolid–sparsomycin | H. influenzae | 4 | ND | In vitro bacteria assay | ||
| Oxazolidinone–cephalosporin–siderophore | E. coli, A. baumannii, P. aeruginosa | E. coli: 0.025 | ADC-1-expressing A. baumannii: 6 | In vitro bacteria assay | ||
| A. baumannii: 0.4 | ||||||
| P. aeruginosa: 0.4 | ||||||
| Mupirocin–holomycin | E. coli | 64 | ND | In vitro bacteria assay |
| Combination antibiotics | Target pathogens | FICI | Synergy mechanisms | Clinical uses | Comments | Reference |
|---|---|---|---|---|---|---|
| Trimethoprim and sulfamethoxazole | E. coli | 0.31 | Mutual potentiation (Sulfamethoxazole potentiates trimethoprim by limiting de novo dihydrofolate production and trimethoprim potentiates sulfamethoxazole activity through inhibition of dihydropterin pyrophosphate synthesis) | Clinical use | ||
| Amikacin and nitrofurantoin | E. coli, K. pneumoniae | E. coli: 0.375 | Amikacin induces bacterial envelope stress by introducing mistranslated proteins, thereby constitutively activating the CpxAR two‐component system. The activation of Cpx signaling stimulates the expression of bacterial major nitroreductases (NfsAB). Nitroreductases overexpression generates considerable quantity of lethal reactive intermediates via nitroreduction and promotes the prodrug activation of nitrofurantoin. | No | ||
| K. pneumoniae: 0.5 | ||||||
| Nalidixic acid and tetracycline | Multidrug-resistant A. baumannii and E. coli | Multidrug-resistant A. baumannii: 0.1875–0.5 | Enhanced uptake and reduced efflux of tetracycline by nalidixic acid explain the basis of synergy between nalidixic acid and tetracycline. | No | Nalidixic acid and tetracycline combination did not display synergy against susceptible A. baumannii and E. coli isolates. | |
| Flavomycin and colistin | E. coli, mcr-1-positive E. coli | E. coli: 0.19–0.49 | The synergy is manifested as an augmented penetration of the E. coli OM by colistin, leading to increased intracellular accumulation of flavomycin and enhanced cell killing thereafter. | No | ||
| Novobiocin and colistin | Colistin-resistant A. baumannii and K. pneumoniae | Colistin-resistant A. baumannii: 0.129 | Novobiocin binds and activates the ATPase LptB that powers lipopolysaccharide transport, which enhances polymyxin activity. | No | ||
| Colistin-resistant K. pneumoniae: 0.012 | ||||||
| Hygromycin A and macrolides | B. burgdorferi | 0.5 | Hygromycin A cooperatively binds ribosomes with nascent peptide exit tunnel-targeting macrolides and slows down their dissociation, which potentiates macrolide’s efficacy in bacterial growth inhibition and early killing. | No |
Not determined.
Not determined.
Table 1.
Table 2.
TOP
 MSK
MSK
 ePub Link
ePub Link Cite this Article
Cite this Article


