- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Volume 63(3); 2025 > Article
-
Review
Harnessing organelle engineering to facilitate biofuels and biochemicals production in yeast - Phuong Hoang Nguyen Tran1,2, Taek Soon Lee1,2,*
-
Journal of Microbiology 2025;63(3):e2501006.
DOI: https://doi.org/10.71150/jm.2501006
Published online: March 28, 2025
1Joint BioEnergy Institute, Emeryville 94608, CA, USA
2Biological Systems & Engineering Division, Lawrence Berkeley National Laboratory, Berkeley 94720, CA, USA
- *Correspondence Taek Soon Lee tslee@lbl.gov
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
ABSTRACT
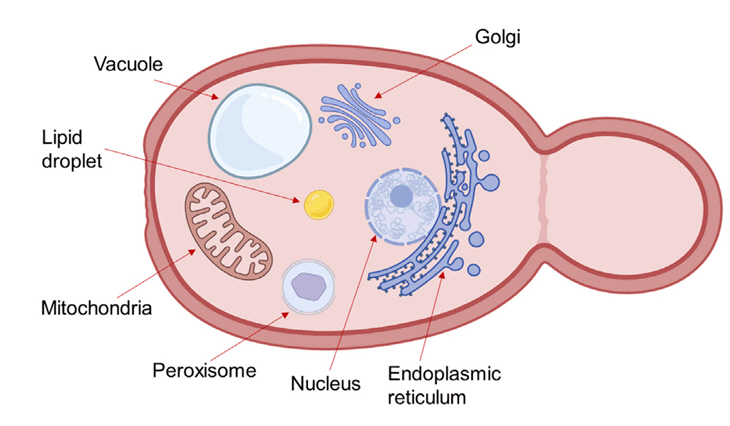
- Microbial biosynthesis using yeast species offers numerous advantages to produce industrially relevant biofuels and biochemicals. Conventional metabolic engineering approaches in yeast focus on biosynthetic pathways in the cytoplasm, but these approaches are disturbed by various undesired factors including metabolic crosstalk, competing pathways and insufficient precursors. Given that eukaryotic cells contain subcellular organelles with distinct physicochemical properties, an emerging strategy to overcome cytosolic pathway engineering bottlenecks is through repurposing these organelles as specialized microbial cell factories for enhanced production of valuable chemicals. Here, we review recent progress and significant outcomes of harnessing organelle engineering for biofuels and biochemicals production in both conventional and non-conventional yeasts. We highlight key engineering strategies for the compartmentalization of biosynthetic pathways within specific organelles such as mitochondria, peroxisomes, and endoplasmic reticulum; involved in engineering of signal peptide, cofactor and energy enhancement, organelle biogenesis and dual subcellular engineering. Finally, we discuss the potential and challenges of organelle engineering for future studies and propose an automated pipeline to fully exploit this approach.
Introduction
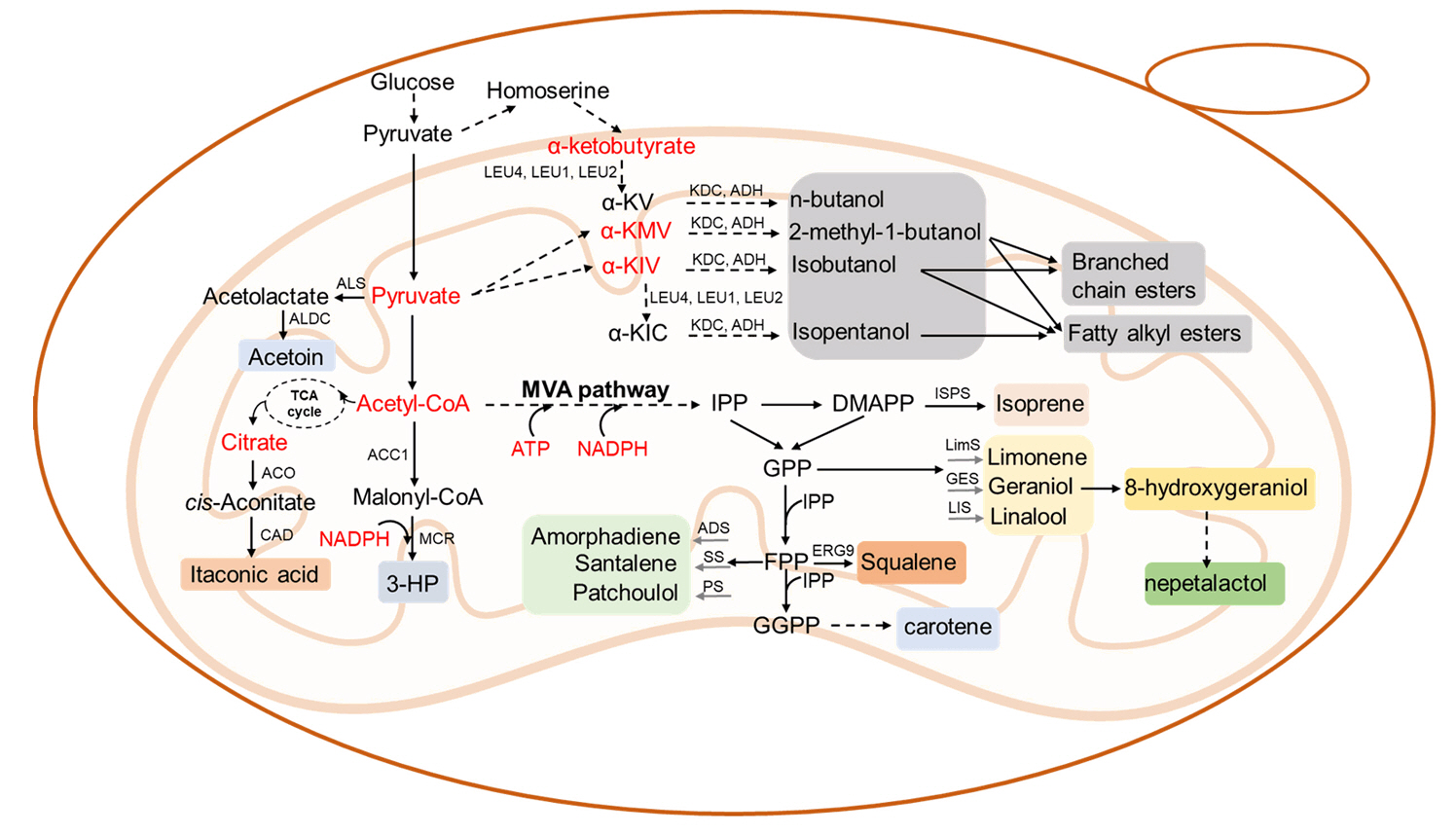
Mitochondrial Engineering
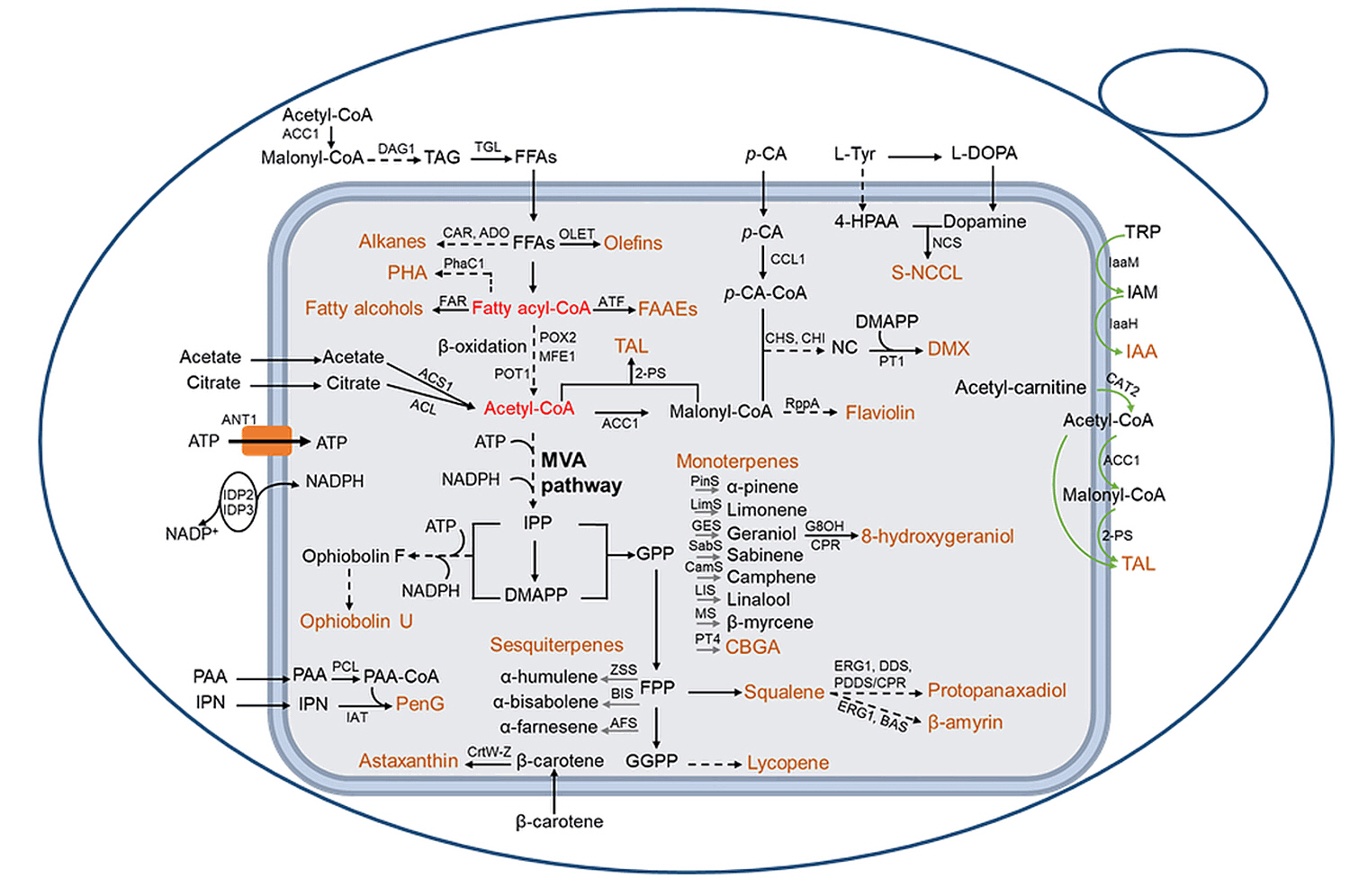
Peroxisomal Engineering
Endoplasmic Reticulum (ER) Engineering
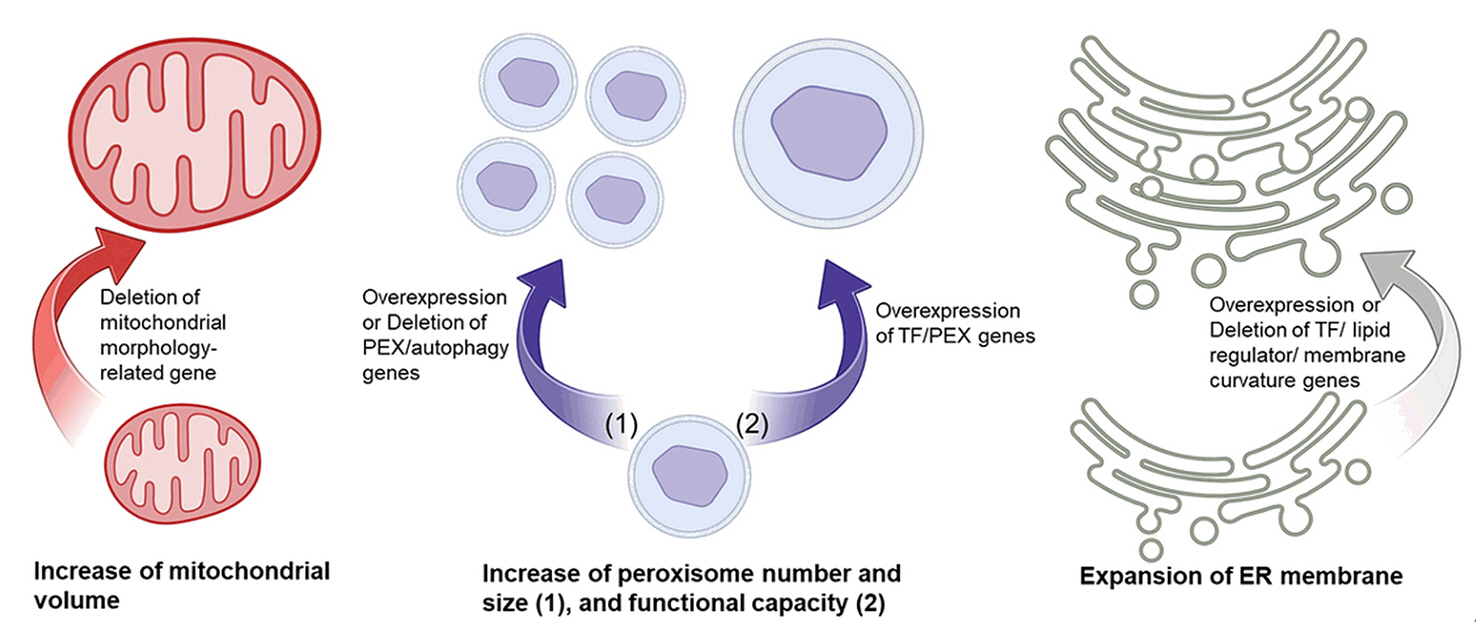
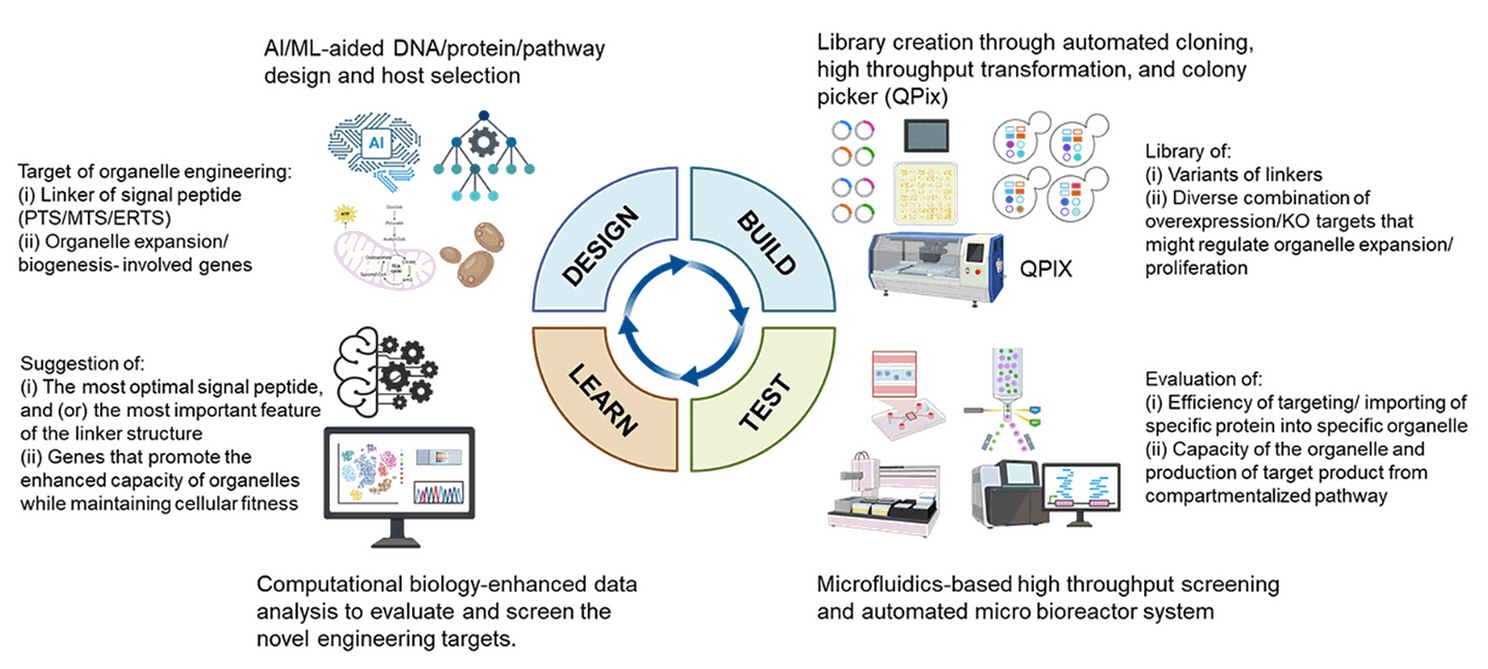
Potentials and Challenges of the Yeast Organelle Engineering
Acknowledgments
We thank Dr. David Carruthers at Lawrence Berkeley National Laboratory for helpful comments on the manuscript. This work was supported by the US Department of Defense (DOD)’s Tri-Service Biotechnology for a Resilient Supply Chain (T-BRSC) program through award AWD00007196 to Lawrence Berkeley National Laboratory.
Conflict of Interest
The authors declare that they have no conflict of interest.
| Product | Yeast species | MTS type | Single/Dual pathway* | Engineering description | Titer | Scale | References | |
|---|---|---|---|---|---|---|---|---|
| Biochemical | Isoprene | S. cerevisiae | COX4 | Dual | · Complete MVA pathway in mitochondria (two copies of tHMG1) | 2.53 g/L | Bioreactor | Lv et al. (20141, 2016) |
| · Diploid strain formation by mating the mitochondrial engineered strain with YXM10 strain1 | ||||||||
| Dual | · Isoprene synthase mutant ISPSLN in mitochondria/cytosol | 11.09 g/L | Bioreactor | Yao et al. (2018) | ||||
| · Complete MVA pathway in mitochondria/cytosol | ||||||||
| · Additional expression of MVD1 and IDI1 in mitochondria/cytosol | ||||||||
| · Diploid strain formation by mating 2 mitochondrial/cytosolic strains | ||||||||
| Amorpha-4,11-diene | COQ3, COX4 | Single | · Entire amorphadiene pathway targeted in mitochondria (only ADS tagged with COQ3-MTS) | 430 mg/L | Flask | Yuan & Ching (2016) | ||
| 8-Hydroxygeraniol | COX4 | Single | · Choice of a starting strain JHY65 for its improved respiratory growth and increased mitochondrial stability2 | 227 mg/L; | Bioreactor | Harvey et al. (20182), Yee et al. (2019) | ||
| · The geraniol pathway targeted in mitochondria (using Gg mFPS and ObGES) | ||||||||
| · Expression of ER-targeted CrG8H | ||||||||
| · Deletion of OYE2 and OYE3 | ||||||||
| Nepetalactol | · Expression of cytosolic CrGOR and CrISY for nepetalactol synthesis | 5.9 mg/L | Tube | |||||
| Linalool | COX4 | Dual | · Complete linalool pathway in both mitochondria and cytosol (using ERG20F96W/N127W and CoLIS) | 23.45 mg/L | Bioreactor | Zhang et al. (2020) | ||
| · Downregulation of endogenous ERG20 | ||||||||
| · Additional expression of a fusion CoLIS-ERG20F96W/N127W protein in both mitochondria and cytosol. | ||||||||
| Single | · The entire MVA pathway targeted in mitochondria | 23.8 mg/L | Flask | Zhang et al. (2022) | ||||
| · Mitochondrial expression of truncated AaLS1 and ERG20F96W/N127W | ||||||||
| Limonene | COX4 | Dual | · The entire limonene pathway targeted in mitochondria | 2.63 g/L | Bioreactor | Kong et al. (2023) | ||
| · The entire cytosolic limonene pathway with multiple copies of ERG20F96W/N127W and tLimS, and downregulation of ERG20 | ||||||||
| · Enhanced acetyl-CoA pool by knocking out CIT2 and MLS1 | ||||||||
| · Enhancing NADPH pool by overexpressing pentose phosphate (PP) pathway and deleting the NADPH-consuming pathway | ||||||||
| α-Santalene | MMF1, COX4, HSP60, LSC2, LDP1, ALD4 | Single | · Complete α-santalene pathway targeted in mitochondria with additional overexpression of tHMG1 | 41 mg/L | N/A | Dong et al. (2021) | ||
| Squalene | COX4 | Dual | · Complete MVA pathway targeted in mitochondria with additional expression of ERG19 | 21.1 g/L | Bioreactor | Zhu et al. (2021) | ||
| · Full squalene pathway expressed in cytosol (overexpression only for ERG10, tHMG1, ERG19, IDI1, ERG20, and ERG9) | ||||||||
| · Expression of ALD6, ASC1, ASC2 and ADH2 for improved cytosolic acetyl-CoA | ||||||||
| · Down regulation of ERG1 | ||||||||
| COX4 | Dual | · Complete MVA pathway targeted in mitochondria with regulation of genes by promoter screening | 230 mg/L | Flask | Yanagibashi et al. (2024a) | |||
| · Full squalene pathway in cytosol with additional copies of tHMG1 and ERG12 | ||||||||
| COX4 | Single | · Complete MVA pathway targeted in mitochondria | 707 nmol/g DCW | Flask | Yanagibashi et al. (2024b) | |||
| · Improved mitochondrial volume by deletion of MDM32 | ||||||||
| β-Carotene | COX4 | Single | · Expression of CrtYBXd, CrtIXd, and BTS1 (for β-carotene synthesis) | 1609 nmol/g DCW | Flask | |||
| 3-Hydroxypropionate | COX4 | Single | · Mitochondrial targeting of mutant ACC1 and dissected MCR | 71.09 g/L | Bioreactor | Zhang et al. (2023) | ||
| · Overexpression of IDP1 and POS5 for improved NADPH pool in mitochondria | ||||||||
| Itaconic acid | A. niger | ICDA, ACOA | Dual | · Itaconic acid pathway targeted in mitochondria (AtCadA and AnAcoA) | 1.4 g/L | Flask | Blumhoff et al. (2013) | |
| · Expression of itaconic acid pathway in cytosol (AtCadA and EcAcnA) | ||||||||
| Acetoin | C. glabrata | COX4 | Single | · Acetoin pathway targeted in mitochondria (BsALS and BaALDC) | 3.26 g/L | Flask | Li et al. (2015) | |
| · Overexpression of mitochondrial pyruvate carrier (ScMPC) | ||||||||
| Biofuel | Isobutanol | S. cerevisiae | COX4 | Single | · Mitochondrial targeting of ARO10, LlKiVD and AdhARE1, ADH7 | 635 mg/L | Tube | Avalos et al. (2013) |
| · Overexpression of mitochondrially native genes ILV2, ILV5, ILV3 | ||||||||
| Single | · Mitochondrial targeting of ARO10, LlAdhARE1 | 1.245 g/L | Plate | Hammer & Avalos (2017b) | ||||
| · Deletion of BAT1 and ILV6 | ||||||||
| Single | · Mitochondrial targeting of ARO10, LlAdhARE1 | 8.49 g/L | Bioreactor | Zhao et al. (2018) | ||||
| · Deletion of BAT1 | ||||||||
| 2-Methyl-1-butanol | Single | · Optogenetic regulation of ILV2 and PDC1 | 2.38 g/L | Bioreactor | ||||
| Isopentanol | COX4, CD9, COX6 | Single | · Mitochondrial targeting of LEU4 mutant, LEU1, LEU2 | 1.24 g/L | Plate | Hammer et al. (2020) | ||
| · Deletion of LEU4, LEU9, BAT1 and OAC1 | ||||||||
| n-Butanol | COX4, CYB2, CAT2 | Single | · Mitochondrial targeting of LEU4, LEU1, LEU2 and citramalate synthase (LiCim) | 1.05 g/L | Bioreactor | Shi et al. (2016) | ||
| · Overexpression of LEU9, cysteine desulfurase NFS1, ARO10 and ADH7 | ||||||||
| Fatty acid alkyl ester | COX4 | Single | · Mitochondrial targeting of ARO10 and ADH7 for improved isobutanol production | 230 mg/L | Flask | Teo et al. (2015) | ||
| · Overexpression of two wax ester synthase Ms Ws2 and Maqu_0168 | ||||||||
| · Deletion of OPI1 and RPD3 | ||||||||
| Isobutyrate | COX4 | Dual | · Mitochondrial targeting of ARO10 and ADH7, and ATF1 | 260.2 mg/L | Tube | Yuan et al. (2016) | ||
| 3-Methyl-1-butyl acetate | · ATF1 overexpression in cytosol | 296.1 mg/L | ||||||
| 2-Methyl-1-butyl acetate | · Overexpression of PYC2, MDH2, MAE1 | 289.6 mg/L | ||||||
*The single pathway refers to the entire biosynthetic pathway or partial downstream pathway, compartmentalized either in the cytosol or in a subcellular organelle; whereas the dual pathway refers to the entire biosynthetic pathway that is expressed in both the cytosol and a subcellular organelle.
Gg: Gallus gallus; Ob: Ocimum basilicum; Cr: Catharanthus roseus; Co: Cinnamomum osmophloeum; Aa: Actinidia arguta; Ll: Lactococcus lactis; Li: Leptospira interrogans; Ms: Marinobacter sp.; Bs: Bacillus subtilis; Ba: Bacillus amyloliquefaciens; Sc: Saccharomyces cerevisiae; At: Aspergillus terreus; An: Aspergillus niger; Ec: Escherichia coli
| Product | Yeast species | PTS type | Single/Dual pathway* | Engineering description | Titer | Scale | References | |
|---|---|---|---|---|---|---|---|---|
| Biochemical | Geraniol | S. cerevisiae | SKL | Single | · Compartmentalization of AgGPPS2 and ObGES in peroxisome | 2.75 mg/L | Plate | Gerke et al. (2020) |
| · Deletion of PEX30 and PEX32 for increased peroxisome number | ||||||||
| · Expression of truncated α-arrestin-like adaptor Bul1 for high tolerance to geraniol | ||||||||
| · The entire geraniol pathway targeted in peroxisome (using ERG20N127Wand ObGerS) with episomal plasmids | 5.52 g/L | Bioreactor | Dusséaux et al. (2020) | |||||
| Dual | · Machine learning (ML)-aided identification of the MVA pathway’s critical gene | 120 mg/L | Flask | Mukherjee et al. (2022) | ||||
| · The MVA pathway targeted in peroxisome | ||||||||
| · The dual MVA platform strain (diploid) created by mating peroxisomal and cytosolic strains | ||||||||
| · Overexpression of a fusion tObGES-ERG20WW | ||||||||
| ePTS1 | Single | · The entire geraniol pathway targeted in peroxisome (using ERG20F96W, N127W and tVoGES) | 9.5 g/L | Flask | Baker et al. (2024) | |||
| · ML-guided screen of peroxisome complexes for improved peroxisome capacity | ||||||||
| P. pastoris | ePTS1, PTS of TAL1-2 | Single | · HTS-based screen of 25 putative PTS, followed by directed evolution of selected 3 PTS1 with high targeting efficiency | 20 mg/L | Plate | Ye et al. (2024) | ||
| · Peroxisomal compartmentalization of geraniol pathway (only tCrGES tagged with newly identified PTS; the remaining proteins tagged with ePTS1). | ||||||||
| R-Limonene | R. toruloides | SKL, PTS2 (C4N11 of 3-KT) | Dual | · Peroxisomal targeting of the entire limonene pathway (two copies of NPPS::LS) | 1.05 g/L | Flask | Gao et al. (2024a) | |
| · Peroxisomal targeting of Nph17 and overexpressing Nph17 and ACCT for improved acetoacetyl-CoA in peroxisome and cytosol, respectively | ||||||||
| · Cytosolic overexpression of NPPS::LS | ||||||||
| · Overexpression of ACL for improved cytosolic acetyl-CoA | ||||||||
| S. cerevisiae | SKL | Single | · Peroxisomal targeting of the entire biosynthetic pathway targeted with episomal plasmids. | 2.58 g/L | Bioreactor | Dusséaux et al. (2020) | ||
| · R-limonene: ClLimS | ||||||||
| α-Pinene | · α-Pinene: PtPinS | 69.22 mg/L | Flask | |||||
| Sabinene | · Sabinene: SpSabS | 32.32 mg/L | ||||||
| Camphene | · Camphene: 1SeCamS | 5.77 mg/L | ||||||
| 8-Hydroxygeraniol | · 8-Hydroxygeraniol: ObGerS, CrG8OH, CrCPR | 25.11 mg/L | ||||||
| Canabigerolic acid (CBGA) | · CBGA: CsPT4 | 0.82 mg./L | ||||||
| Linalool | ePTS1 | Dual | · Improving catalytic efficiency toward GPP of linalool synthase through site-directed mutagenesis | 2.6 g/L | Bioreactor | Zhou et al. (2023) | ||
| · Complete linalool pathway targeting to peroxisome | ||||||||
| · Overexpression of tHMG1, IDI1, ERG20F96W, N127W, and McLIS with repression of native ERG20 for cytosolic linalool pathway | ||||||||
| · Mating peroxisomal and cytosolic strain to form the dual strain (diploid) | ||||||||
| β-Myrcene | SKL | Dual | · Source screening of β-myrcene synthase, with truncated QiMS as the best candidate | 142.64 mg/L | Bioreactor | Shu et al. (2024) | ||
| · Peroxisome localization of fusion enzyme tQiMSD436N-ERG20WW | ||||||||
| · Overexpression of tHMG1 and IDI in the cytosolic MVA pathway and replacement of native ERG20 by ERG20W | ||||||||
| · Enhancing cytosolic acetyl-CoA by overexpressing ADH2 and ALD6 | ||||||||
| α-Humulene | S. cerevisiae | ePTS1 | Dual | · Compartmentalization of entire α-humulene pathway in peroxisome | 1.73 g/L | Bioreactor | Zhang et al. (2020) | |
| · Overexpression of tHMG1 and ERG20 in cytosol | ||||||||
| · Down regulation of ERG9 by replacing its native promoter by HXT1p and adding PEST sequence | ||||||||
| · Additional overexpression of cytosolic ZzZSS1 in rDNA sites | ||||||||
| SKL | Dual | · ML-guided dual MVA platform strain (diploid) created by mating peroxisomal and cytosolic strains | ~22.5 mg/L | Flask | Mukherjee et al. (2022) | |||
| · Overexpression of cytosolic ERG20 and ZzZSS1 | ||||||||
| oPTS1* | Dual | · Develop an orthogonal transport system ScPEX5*-oPTS1* | 17.33 g/L | Bioreactor | Zhang et al. (2024) | |||
| · rDNA integration of the entire cytosolic and peroxisomal α-humulene pathway (ZSS1 as humulene synthase) | ||||||||
| · Overexpression of ANT1, IDP2, IDP3 (rDNA integration) for improved ATP and NADPH pool in peroxisome | ||||||||
| · Down regulation of ERG9 by replacing its native promoter by HXT1p and adding PEST sequence | ||||||||
| Y. lipolytica | PTS (GGGSSKL) | Single | · Localization of entire α-humulene pathway in peroxisome (NADH-HMG1 as HMG-CoA reductase; AcHS2 as humulene synthase) | 3.2 g/L | Bioreactor | Guo et al. (2021) | ||
| · Increasing ATP supply in peroxisome by overexpression of ANT1 | ||||||||
| · Adjusting the copy numbers of rate-limiting enzymes (one more copy of peroxisomal targeting NADH-HMG1 and AcHS2) | ||||||||
| · Upregulation of β-oxidation by overexpressing POT1 | ||||||||
| · Oleic acid as a possible C source for α-humulene accumulation | ||||||||
| PTS (GGGSSKL) | Dual | · Iterative integration of the entire cytosolic and peroxisomal α-humulene pathway (AcHS2 as humulene synthase) | 21.7 g/L | Bioreactor | Guo et al. (2022) | |||
| · Mediating copy number of cytosolic tHMG1 and AcHS2 | ||||||||
| · Improving cytosolic acetyl-CoA by overexpression of CoA‐acetylating aldehyde dehydrogenase (EcAAD) | ||||||||
| · Repression of ERG9 expression using Cu2+ repressible promoter | ||||||||
| Candida tropicalis | SKL | Single | · Multiple overexpression of the entire peroxisomal α-humulene pathway (double copies of the pathway) | 2.42 mg/L | Flask | Zhang et al. (2022) | ||
| α-Bisabolene | P. pastoris | SKL and LARF | Single | · Peroxisome isolation of the entire α-bisabolene pathway with fusion enzyme of FPPS-AgBIS; upregulation of acetyl-CoA-to-mevalonate pathway; and additional copy of separated FPPS and AgBIS | 1.1 g/L | Flask | Gao et al. (2024b) | |
| α-Farnesene | ePTS1 | Dual | · Overexpression of tHMG1, IDI1, ERG20 and MdAFS for cytosolic α-farnesene pathway | 2.56 g/L | Flask | Liu et al. (2021) | ||
| · Improving cytosolic acetyl-CoA pool by introducing pyruvate dehydrogenase (Ec cytoPDH) and ATP-dependent citrate lyase (YlACL) | ||||||||
| · Introduction of IUP pathway and downstream genes (2 copies of IDI, ERG20, MdAFS) into peroxisome | ||||||||
| β-Amyrin | S. cerevisiae | ePTS1 | Dual | · Peroxisome compartmentalization of squalene pathway | 2.6 g/L | Bioreactor | Du et al. (2022) | |
| · Double copies of downstream β-amyrin pathway (ERG1 and GgbAS1) in cytosol | ||||||||
| · Deregulation of ERG7 | ||||||||
| Squalene | ePTS1 | Dual | · The entire squalene pathway targeted in peroxisome with an additional copy of tHMG1 and IDI1 | 11 g/L | Bioreactor | Liu et al. (2020) | ||
| · Deletion of GPD1 and GPD2 | ||||||||
| · Overexpression of ANT1, IDP2 and IDP3 for improved ATP and NADPH pool in peroxisome | ||||||||
| · ACS1, YlACL1 and YlACL2 targeted to peroxisome for improved peroxisomal acetyl-CoA | ||||||||
| · Diploid strain formation by mating two peroxisomal and cytosolic strains | ||||||||
| SKL | Dual | · The dual MVA platform strain (diploid) created by mating peroxisomal and cytosolic MVA strains | 300 mg/L | Flask | Mukherjee et al. (2022) | |||
| · Overexpression of ERG9 and ERG20 | ||||||||
| · Prevention of squalene degradation by supplementation of terbinafine | ||||||||
| Y. lipolytica | SKL | Dual | · The complete squalene pathway targeted in peroxisome | 32.8 g/L (on glucose)/ 31.6 g/L (on acetate) | Bioreactor | Ma et al. (2024) | ||
| · Enhanced TAG-derived free fatty acids in peroxisome through overexpressing TlTGL, DGA1 and ACC1 | ||||||||
| · Improved cytosolic acetyl-CoA pool by overexpressing PYC1, YHM2 and MmACL | ||||||||
| · Upregulated β-oxidation by overexpressing POX2, MFE1 and POT1 | ||||||||
| · Increased size/number of peroxisomes by overexpressing PEX10 | ||||||||
| · Introduction of Acetyl-CoA shortcut in the peroxisome by targeting 2SeACS1L641P. | ||||||||
| Lycopene | P. pastoris | SKL | Single | · Peroxisomal compartmentalization of lycopene biosynthetic pathway (CrtE, CrtB, CrtI) | 73.9 mg/L | Bioreactor | Bhataya et al. (2009) | |
| Astaxanthin | Y. lipolytica | SKL | Single | · Peroxisomal targeting of the fusion enzyme PsCtrW-HpCrtZ | 58.7 mg/L | Flask | Ma et al. (2021) | |
| Protopanaxadiol | S. cerevisiae | PTS1, PTS2 | Single | · Regulating number and size of peroxisome through deletion of PEX11 and ATG36, and overexpression of PEX34 | 4.1 mg/L | Flask | Choi et al. (2022) | |
| · Peroxisomal compartmentalization of protopanaxadiol pathway (including tHMG1, ERG9, PgErg1, PgDs, PgPpds, and PgCpr) | ||||||||
| (S)-Norcoclaurine | ePTS1 | Single | · Targeting norcoclaurine synthase (NCS) in peroxisome | ~7 mg/L | Plate | Grewal et al. (2021) | ||
| · Controlling peroxisome biogenesis by engineering transcriptional factors ADR1, OAF1 and PIP2 | ||||||||
| Ophiobolins U | PTS of PEX15 | N/A | · Peroxisomal surface display of CAT2 for improved cytosolic acetyl-CoA | 128.9 mg/L | Flask | Zhang et al. (2024) | ||
| Indole-3-acetic acid | K. marxianus | PTS of PEX15 | Single | · Peroxisomal surface display of tryptophan-2-monooxygenase PsIaaM and indole-acetamide hydrolase PsIaaH | ~61.25 mg/L | Tube | Bassett et al. (2024) | |
| · Controlling peroxisome number and size by overexpression of PEX19 | ||||||||
| Triacetic acid lactone (TAL) | PTS of PEX15 | Single | · Peroxisomal surface display of CAT2, ACC1 and Gh2-PS | ~ 1.1 g/L | Tube | |||
| · Controlling peroxisome number and size by overexpression of PEX11 | ||||||||
| S. cerevisiae | PTS of PEX15 | Single | · Peroxisomal surface display of CAT2, ACC1S1157A and Gh2-PS | 0.77 g/L | Tube | Yocum et al. (2022) | ||
| pPTS (PTS of Psc60) | Dual | · Cytosolic expression and peroxisomal targeting of Gh2-PS and ACC1S686A, S659A, S1157A | 0.14 g/L | Plate | Lin et al. (2023) | |||
| · Increasing peroxisome proliferation by overexpression of ADR1, OAF1 and PIP2 | ||||||||
| Flaviolin | ePTS1 | Single | · Peroxisomal targeting of Sac_RppA and ACC1S686A, S659A, S1157A | N/A | N/A | |||
| Desmethylxanthohumol (DMX) | N/A | Single | · Compartmentalization of the complete MVA pathway in peroxisome | ~62.5 µg/L | Flask | Yang et al. (2024) | ||
| · Peroxisomal targeting of ACC1 and naringenin chalcone and DMX biosynthetic genes | ||||||||
| Amino acid (Lysine, Histidine) | S. japonicus | PTS1 and PTS2 | N/A | · Peroxisomal targeting of GPD2, LYS3 and HIS2 enzymes | N/A | N/A | Gu et al. (2023) | |
| · Regulating peroxisome size with PEX5 mutant and PEX11 | ||||||||
| Penicillin | Hansenula polymorpha | SKL | Single | · Peroxisomal targeting of penicillin pathway’s bottom genes, including PcIAT and PcPCL | 1 mg/L | Plate | Gidijala et al. (2009) | |
| PHA | S. cerevisiae | PTS of BnICL | Single | · Peroxisomal targeting of PaPHAC1 synthase | 0.45 g/g DCW | Flask | Poirier et al. (2001) | |
| Biofuel | Fatty acid ethyl esters (FAEEs) | Y. lipolytica | SKL | Single | · Peroxisomal targeting of wax ester synthase (AbAtfA) | 110.9 mg/L | Flask | Xu et al. (2016) |
| Fatty alcohols | S. cerevisiae | PTS2 | Single | · Peroxisomal targeting of fatty acyl-CoA reductase (TaFAR) | 1.3 g/L | Tube | Sheng et al. (2016) | |
| · Increasing peroxisomal importing rate by overexpression of PEX7 | ||||||||
| · Improved malonyl-CoA pool by overexpression of ACC1 | ||||||||
| PTS2 | Single | · Peroxisomal targeting of fatty acyl-CoA reductase MaFAR in fatty acyl-CoA overproducing strain | 193 mg/L | Flask | Zhou et al. (2016) | |||
| Alkanes | PTS1 and PTS2 | Single | · Peroxisomal targeting of free fatty acid (FFA) pathway including MmCAR, AnNPGA, 3SeADO, ferredoxin 3SeFd and ferredoxin reductase 3SeFNR | ~ 3.5 mg/L | Flask | |||
| · Increasing fatty acid production by deleting HFD1 and POX1 | ||||||||
| · Increasing peroxisome population by deletion of PEX31, 32 and overexpression of PEX34 | ||||||||
| Olefins | PTS2 | · Peroxisomal targeting of OleT with flavodoxin EcFldA and flavodoxin reductase EcFNR | ~ 0.20 mg/L | Flask | ||||
| α-Alkenes | P. pastoris | PTS1 and PTS2 | Single | · Peroxisomal targeting of decarboxylase PfUndB together with cofactor protein putidaredoxin–putidaredoxin reductase PpPdr/Pdx | 1.6 mg/L | Flask | Cai et al. (2022) | |
*The single pathway refers to the entire biosynthetic pathway or partial downstream pathway, compartmentalized either in the cytosol or in a subcellular organelle; whereas the dual pathway refers to the entire biosynthetic pathway that is expressed in both the cytosol and a subcellular organelle.
Yl: Yarrowia lipolytica; Ob: Ocimum basilicum; Ag: Abies grandis; Vo: Valeriana officinalis; Tl: Thermomyces lanuginosus; Mm: Mus musculus; HTS: high throughput screening; Cr: Catharanthus roseus; Cl: Citrus limon; Pt: Pinus taeda; Sp: Salvia pomifera; 1Se: Solanum elaeagnifolium; Cs: Cannabis sativa; 2Se: Salmonella enterica; Ps: Paracoccus sp.; Hp: Haematococcus pluvialis; Zz: Zingiber zerumbet; Sc: Saccharomyces cerevisiae; At: Arabidopsis thaliana; Ac: Aquilaria crassna; Ec: E. coli; Gg: Glycyrrhiza glabra; Qi: Quercus ilex; Md: Malus domestica; Pg: Panax ginseng; Sac: Streptomyces aculeolatus ; Gh: Gerbera hybrid; Ab: Acinetobacter baylyi ADP1; Ta: Tyto alba; Ma: Marinobacter aquaeolei; 3Se: Synechococcus elongates; Mm: Mycobacterium marinum; An: Aspergillus nidulans ; Pf: Pseudomonas fluorescens; Pp: Pseudomonas putida; Pc: Penicillium chrysogenum; Ps: Pseudomonas savastanoi; Pa: Pseudomonas aeruginosa; Bn: Brassica napus; NPPS::LS: the chimera of NPP synthase from Solanum lycopersicum and limonene synthase from Citrus limon
| Product | Yeast species | ER engineering | Titer | Scale | References | |
|---|---|---|---|---|---|---|
| Biochemical | Opiate (morphine) | S. cerevisiae | · ER targeting of NADPH-dependent aldo-keto reductase PsCOR1.3 with ER signal peptide from CNE1 | 3.1 mg/L | Plate | Thodey et al. (2014) |
| Ceramide NS | S. cerevisiae | · ER targeting of human sphingolipid desaturase (hDES1) with ER retention signal KKEK from SUR2 | N/A | N/A | Murakami et al. (2015) | |
| Trans-nootkatol | S. cerevisiae | · Regulating ER membrane proliferation by overexpression of ICE2 | 30 mg/L | Flask | Emmerstorfer et al. (2015) | |
| 1-Hydroxybufuralol | P. pastoris | N/A | N/A | |||
| β-Amyrin | S. cerevisiae | · Expansion of ER by deleting the phosphatidic acid phosphatase PAH1 | N/A | N/A | Arendt et al. (2017) | |
| Medicagenic acid | 27.1 mg/L | Flask | ||||
| Oleanane-type sapogenin | N/A | N/A | ||||
| Squalene | S. cerevisiae | · Expansion of ER by overexpression of transcriptional factor INO2 | 634 mg/L | Flask | Kim et al. (2019) | |
| Protopanaxadiol | S. cerevisiae | · Controlling gene dosage of tHMG1 (3 copies) for squalene production | 12.1 mg/L | Flask | ||
| Astaxanthin | Y. lipolytica | · ER targeting of the fusion enzyme β-carotene ketolase-hydroxylase (PsCtrW-HpCrtZ) with ER signal peptide KDEL | 53.2 mg/L | Flask | Ma et al. (2021) | |
| Human antibody (IgG) | S. cerevisiae | · Expansion of ER by deleting lipid-regulator OPI1 | 126 µg/L | Plate | de Ruijter et al. (2016) | |
| · Overexpression of peptidyl-prolyl isomerase CPR5 for enhanced secretion | ||||||
| · Modification of ER morphology and size by deleting lipid-regulator gene OPI1 and ER membrane curvature genes RTN1, RTN2 and YOP1 | 79 ng/mL | Plate | Niemelä et al. (2024) | |||
| Ovalbumin | S. cerevisiae | · Expression of OVA using signal peptide from INU1 | 116.3 mg/L | Bioreactor | Dong et al. (2024) | |
| · Co-expression of chaperone Kar2 and disulfide isomerase PDI for improved folding environment in ER | ||||||
| · Expansion of ER by deleting OPI1 and overexpressing transcriptional factors INO2 and INO4 | ||||||
| S-Scoulerine | S. cerevisiae | · ER targeting of berberine bridge enzyme (CyBBE) with ER C-terminal HDEL signal peptide (CyBBE_ERTS) | 113.1 mg/L | Flask | Jiao et al. (2024) | |
| · ER targeting of mammalian peroxiredoxin IV (PRDX4) with N- and C-terminal ERTS (α- mPRDX4_ERTS) for reducing toxicity of hydrogen peroxide | ||||||
| · Expansion of ER by deleting transcriptional regulator OPI1 of phospholipid biosynthesis | ||||||
| Biofuel | Fatty acid ethyl esters (FAEEs) | Y. lipolytica | · ER targeting of wax-ester synthase (AbAtfA) with ER peptide signal KDEL | 136.5 mg/L | Flask | Xu et al. (2016) |
| Fatty alkane | · ER targeting of fatty acyl-CoA reductase (AbACR1)-aldehyde deformylating oxygenase (PmADO) gene cluster with ER signal peptide KDEL | 16.8 mg/L | Flask | |||
| Fatty alcohol | · ER targeting of fatty acyl-CoA reductase (AbACR1) and aldehyde reductase (EcAHR) with ER signal peptide KDEL | 49.2 mg/L | Flask | |||
- Arendt P, Miettinen K, Pollier J, De Rycke R, Callewaert N, et al. 2017. An endoplasmic reticulum-engineered yeast platform for overproduction of triterpenoids. Metab Eng. 40: 165–175. ArticlePubMed
- Avalos JL, Fink GR, Stephanopoulos G. 2013. Compartmentalization of metabolic pathways in yeast mitochondria improves the production of branched-chain alcohols. Nat Biotechnol. 31(4): 335–341. ArticlePubMedPMCPDF
- Baker JJ, Shi J, Wang S, Mujica EM, Bianco S, et al. 2024. ML-enhanced peroxisome capacity enables compartmentalization of multienzyme pathway. Nat Chem Biol. https://doi.org/10.1038/s41589-024-01759-2. Article
- Bassett S, Suganda JC, Da Silva NA. 2024. Engineering peroxisomal surface display for enhanced biosynthesis in the emerging yeast Kluyveromyces marxianus. Metab Eng. 86: 326–336. ArticlePubMed
- Bhataya A, Schmidt-Dannert C, Lee PC. 2009. Metabolic engineering of Pichia pastoris X-33 for lycopene production. Process Biochem. 44(10): 1095–1102. Article
- Blumhoff ML, Steiger MG, Mattanovich D, Sauer M. 2013. Targeting enzymes to the right compartment: metabolic engineering for itaconic acid production by Aspergillus niger. Metab Eng. 19: 26–32. ArticlePubMed
- Cai P, Li Y, Zhai X, Yao L, Ma X, et al. 2022. Microbial synthesis of long-chain α-alkenes from methanol by engineering Pichia pastoris. Bioresour Bioprocess. 9(1): 58.ArticlePubMedPMCPDF
- Choi BH, Kang HJ, Kim SC, Lee PC. 2022. Organelle engineering in yeast: Enhanced production of protopanaxadiol through manipulation of peroxisome proliferation in Saccharomyces cerevisiae. Microorganisms. 10(3): 635.ArticlePubMedPMC
- Crater JS, Lievense JC. 2018. Scale-up of industrial microbial processes. FEMS Microbiol Lett. 365(13): fny115.Article
- de Ruijter JC, Koskela EV, Frey AD. 2016. Enhancing antibody folding and secretion by tailoring the Saccharomyces cerevisiae endoplasmic reticulum. Microb Cell Fact. 15: 87.ArticlePubMedPMC
- DeLoache WC, Russ ZN, Dueber JE. 2016. Towards repurposing the yeast peroxisome for compartmentalizing heterologous metabolic pathways. Nat Commun. 7: 11152.ArticlePubMedPMCPDF
- Dong X, Lin Y, Zhang J, Lv X, Liu L, et al. 2024. Modification of the endoplasmic reticulum to enhance ovalbumin secretion in Saccharomyces cerevisiae. J Agric Food Chem. 72(36): 19985–19993. ArticlePubMedLink
- Dong C, Shi Z, Huang L, Zhao H, Xu Z, et al. 2021. Cloning and characterization of a panel of mitochondrial targeting sequences for compartmentalization engineering in Saccharomyces cerevisiae. Biotechnol Bioeng. 118(11): 4269–4277. ArticlePubMedLink
- Du MM, Zhu ZT, Zhang GG, Zhao YQ, Gao B, et al. 2022. Engineering Saccharomyces cerevisiae for hyperproduction of β-amyrin by mitigating the inhibition effect of squalene on β-amyrin synthase. J Agric Food Chem. 70(1): 229–237. ArticlePubMed
- Dusséaux S, Wajn WT, Liu Y, Ignea C, Kampranis SC. 2020. Transforming yeast peroxisomes into microfactories for the efficient production of high-value isoprenoids. Proc Natl Acad Sci USA. 117(50): 31789–31799. ArticlePubMedPMC
- Emmerstorfer A, Wimmer-Teubenbacher M, Wriessnegger T, Leitner E, Müller M, et al. 2015. Over-expression of ICE2 stabilizes cytochrome P450 reductase in Saccharomyces cerevisiae and Pichia pastoris. Biotechnol J. 10(4): 623–635. ArticlePubMed
- Farhi M, Marhevka E, Masci T, Marcos E, Eyal Y, et al. 2011. Harnessing yeast subcellular compartments for the production of plant terpenoids. Metab Eng. 13(5): 474–481. ArticlePubMed
- Gao Q, Dong Y, Huang Y, Liu S, Zheng X, et al. 2024a. Dual-regulation in peroxisome and cytoplasm toward efficient limonene biosynthesis with Rhodotorula toruloides. ACS Synth Biol. 13(8): 2545–2554. ArticlePubMedLink
- Gao L, Hou R, Cai P, Yao L, Wu X, et al. 2024b. Engineering yeast peroxisomes for α-bisabolene production from sole methanol with the aid of proteomic analysis. JACS Au. 4(7): 2474–2483. ArticlePubMedPMCLink
- Gerke J, Frauendorf H, Schneider D, Wintergoller M, Hofmeister T, et al. 2020. Production of the fragrance geraniol in peroxisomes of a product-tolerant baker’s yeast. Front Bioeng Biotechnol. 8: 582052.ArticlePubMedPMC
- Gidijala L, Kiel JAKW, Douma RD, Seifar RM, van Gulik WM, et al. 2009. An engineered yeast efficiently secreting penicillin. PLoS One. 4(12): e8317. ArticlePubMedPMC
- Grewal PS, Samson JA, Baker JJ, Choi B, Dueber JE. 2021. Peroxisome compartmentalization of a toxic enzyme improves alkaloid production. Nat Chem Biol. 17(1): 96–103. ArticlePubMedPDF
- Grillitsch K, Connerth M, Köfeler H, Arrey TN, Rietschel B, et al. 2011. Lipid particles/droplets of the yeast Saccharomyces cerevisiae revisited: lipidome meets proteome. Biochim Biophys Acta. 1811(12): 1165–1176. ArticlePubMedPMC
- Gu Y, Alam S, Oliferenko S. 2023. Peroxisomal compartmentalization of amino acid biosynthesis reactions imposes an upper limit on compartment size. Nat Commun. 14(1): 5544.ArticlePubMedPMCPDF
- Guo Q, Li YW, Yan F, Li K, Wang YT, et al. 2022. Dual cytoplasmic-peroxisomal engineering for high-yield production of sesquiterpene α-humulene in Yarrowia lipolytica. Biotechnol Bioeng. 119(10): 2819–2830. ArticlePubMedLink
- Guo Q, Shi TQ, Peng QQ, Sun XM, Ji XJ, et al. 2021. Harnessing Yarrowia lipolytica peroxisomes as a subcellular factory for α-humulene overproduction. J Agric Food Chem. 69(46): 13831–13837. ArticlePubMed
- Hammer SK, Avalos JL. 2017a. Harnessing yeast organelles for metabolic engineering. Nat Chem Biol. 13(8): 823–832. ArticlePDF
- Hammer SK, Avalos JL. 2017b. Uncovering the role of branched-chain amino acid transaminases in Saccharomyces cerevisiae isobutanol biosynthesis. Metab Eng. 44: 302–312. Article
- Hammer SK, Zhang Y, Avalos JL. 2020. Mitochondrial compartmentalization confers specificity to the 2-ketoacid recursive pathway: increasing isopentanol production in Saccharomyces cerevisiae. ACS Synth Biol. 9(3): 546–555. ArticlePubMed
- Harvey CJB, Tang M, Schlecht U, Horecka J, Fischer CR, et al. 2018. HEx: a heterologous expression platform for the discovery of fungal natural products. Sci Adv. 4(4): eaar5459. ArticlePubMedPMC
- Huttanus HM, Feng X. 2017. Compartmentalized metabolic engineering for biochemical and biofuel production. Biotechnol J. 12(6): e1700038. ArticleLink
- Jiao X, Fu X, Li Q, Bu J, Liu X, et al. 2024. De novo production of protoberberine and benzophenanthridine alkaloids through metabolic engineering of yeast. Nat Commun. 15(1): 8759.ArticlePubMedPMCPDF
- Kim J, Hoang Nguyen Tran P, Lee SM. 2020. Current challenges and opportunities in non-native chemical production by engineered yeasts. Front Bioeng Biotechnol. 8: 594061.ArticlePubMedPMC
- Kim JE, Jang IS, Son SH, Ko YJ, Cho BK, et al. 2019. Tailoring the Saccharomyces cerevisiae endoplasmic reticulum for functional assembly of terpene synthesis pathway. Metab Eng. 56: 50–59. ArticlePubMed
- Kong X, Wu Y, Yu W, Liu Y, Li J, et al. 2023. Efficient synthesis of limonene in Saccharomyces cerevisiae using combinatorial metabolic engineering strategies. J Agric Food Chem. 71(20): 7752–7764. ArticlePubMedLink
- Li S, Liu L, Chen J. 2015. Compartmentalizing metabolic pathway in Candida glabrata for acetoin production. Metab Eng. 28: 1–7. ArticlePubMed
- Lian J, Si T, Nair NU, Zhao H. 2014. Design and construction of acetyl-CoA overproducing Saccharomyces cerevisiae strains. Metab Eng. 24: 139–149. ArticlePubMed
- Lin P, Fu Z, Liu X, Liu C, Bai Z, et al. 2023. Direct utilization of peroxisomal acetyl-CoA for the synthesis of polyketide compounds in Saccharomyces cerevisiae. ACS Synth Biol. 12(6): 1599–1607. ArticlePubMedLink
- Liu H, Chen SL, Xu JZ, Zhang WG. 2021. Dual regulation of cytoplasm and peroxisomes for improved α-farnesene production in recombinant Pichia pastoris. ACS Synth Biol. 10(6): 1563–1573. ArticlePubMed
- Liu GS, Li T, Zhou W, Jiang M, Tao XY, et al. 2020. The yeast peroxisome: a dynamic storage depot and subcellular factory for squalene overproduction. Metab Eng. 57: 151–161. ArticlePubMed
- Lombard J, Moreira D. 2011. Origins and early evolution of the mevalonate pathway of isoprenoid biosynthesis in the three domains of life. Mol Biol Evol. 28(1): 87–99. ArticlePubMed
- Lv X, Wang F, Zhou P, Ye L, Xie W, et al. 2016. Dual regulation of cytoplasmic and mitochondrial acetyl-CoA utilization for improved isoprene production in Saccharomyces cerevisiae. Nat Commun. 7: 12851.ArticlePubMedPMCPDF
- Lv X, Xie W, Lu W, Guo F, Gu J, et al. 2014. Enhanced isoprene biosynthesis in Saccharomyces cerevisiae by engineering of the native acetyl-CoA and mevalonic acid pathways with a push-pull-restrain strategy. J Biotechnol. 186: 128–136. ArticlePubMed
- Ma Y, Li J, Huang S, Stephanopoulos G. 2021. Targeting pathway expression to subcellular organelles improves astaxanthin synthesis in Yarrowia lipolytica. Metab Eng. 68: 152–161. ArticlePubMed
- Ma Y, Shang Y, Stephanopoulos G. 2024. Engineering peroxisomal biosynthetic pathways for maximization of triterpene production in Yarrowia lipolytica. Proc Natl Acad Sci USA. 121(5): e2314798121. ArticlePubMedPMC
- Malina C, Larsson C, Nielsen J. 2018. Yeast mitochondria: an overview of mitochondrial biology and the potential of mitochondrial systems biology. FEMS Yeast Res. 18(5): 10.Article
- Margeot A, Blugeon C, Sylvestre J, Vialette S, Jacq C, et al. 2002. In Saccharomyces cerevisiae, ATP2 mRNA sorting to the vicinity of mitochondria is essential for respiratory function. EMBO J. 21(24): 6893–6904. ArticlePubMedPMC
- Mukherjee M, Blair RH, Wang ZQ. 2022. Machine-learning guided elucidation of contribution of individual steps in the mevalonate pathway and construction of a yeast platform strain for terpenoid production. Metab Eng. 74: 139–149. ArticlePubMed
- Murakami S, Shimamoto T, Nagano H, Tsuruno M, Okuhara H, et al. 2015. Producing human ceramide-NS by metabolic engineering using yeast Saccharomyces cerevisiae. Sci Rep. 5: 16319.ArticlePubMedPMCPDF
- Niemelä LRK, Koskela EV, Frey AD. 2024. Modification of the endoplasmic reticulum morphology enables improved recombinant antibody expression in Saccharomyces cerevisiae. J Biotechnol. 387: 1–11. ArticlePubMed
- Okoro V, Azimov U, Munoz J. 2022. Recent advances in production of bioenergy carrying molecules, microbial fuels, and fuel design - A review. Fuel. 316: 123330.Article
- Poirier Y, Erard N, Petétot JM. 2001. Synthesis of polyhydroxyalkanoate in the peroxisome of Saccharomyces cerevisiae by using intermediates of fatty acid beta-oxidation. Appl Environ Microbiol. 67(11): 5254–5260. ArticlePubMedPMCLink
- Qiu XY, Xie SS, Min L, Wu XM, Zhu LY, et al. 2018. Spatial organization of enzymes to enhance synthetic pathways in microbial chassis: a systematic review. Microb Cell Fact. 17(1): 120.ArticlePubMedPMCPDF
- Radivojević T, Costello Z, Workman K, Garcia Martin H. 2020. A machine learning Automated Recommendation Tool for synthetic biology. Nat Commun. 11: 4879.ArticlePubMedPMC
- Schwarz DS, Blower MD. 2016. The endoplasmic reticulum: structure, function and response to cellular signaling. Cell Mol Life Sci. 73(1): 79–94. ArticlePubMedPDF
- Sheng J, Stevens J, Feng X. 2016. Pathway compartmentalization in peroxisome of Saccharomyces cerevisiae to produce versatile medium chain fatty alcohols. Sci Rep. 6: 26884.ArticlePubMedPMCPDF
- Shi S, Si T, Liu Z, Zhang H, Ang EL, et al. 2016. Metabolic engineering of a synergistic pathway for n-butanol production in Saccharomyces cerevisiae. Sci Rep. 6: 25675.ArticlePubMedPMCPDF
- Shu Y, Dong T, Zhou X, Wang H, Liu H, et al. 2024. Systematic engineering to enhance β-myrcene production in yeast. J Agric Food Chem. 72(35): 19395–19402. ArticlePubMedLink
- Sibirny AA. 2016. Yeast peroxisomes: structure, functions and biotechnological opportunities. FEMS Yeast Res. 16(4): fow038.ArticlePubMed
- Tao XY, Lin YC, Wang FQ, Liu QH, Ma YS, et al. 2022. Production of sesquiterpene patchoulol in mitochondrion-engineered Saccharomyces cerevisiae. Biotechnol Lett. 44(4): 571–580. ArticlePubMedPDF
- Teo WS, Ling H, Yu AQ, Chang MW. 2015. Metabolic engineering of Saccharomyces cerevisiae for production of fatty acid short- and branched-chain alkyl esters biodiesel. Biotechnol Biofuels. 8: 177.ArticlePubMedPMCPDF
- Thodey K, Galanie S, Smolke CD. 2014. A microbial biomanufacturing platform for natural and semisynthetic opioids. Nat Chem Biol. 10(10): 837–844. ArticlePubMedPMCPDF
- van Roermund CW, Hettema EH, van den Berg M, Tabak HF, Wanders RJ. 1999. Molecular characterization of carnitine-dependent transport of acetyl-CoA from peroxisomes to mitochondria in Saccharomyces cerevisiae and identification of a plasma membrane carnitine transporter, Agp2p. EMBO J. 18(21): 5843–5852. ArticlePubMedPMC
- Vizeacoumar FJ, Torres-Guzman JC, Bouard D, Aitchison JD, Rachubinski RA. 2004. Pex30p, Pex31p, and Pex32p form a family of peroxisomal integral membrane proteins regulating peroxisome size and number in Saccharomyces cerevisiae. Mol Biol Cell. 15(2): 665–677. ArticlePubMedPMC
- Weinert BT, Iesmantavicius V, Moustafa T, Schölz C, Wagner SA, et al. 2014. Acetylation dynamics and stoichiometry in Saccharomyces cerevisiae. Mol Syst Biol. 10(1): 716.ArticlePubMedPMCLink
- West M, Zurek N, Hoenger A, Voeltz GK. 2011. A 3D analysis of yeast ER structure reveals how ER domains are organized by membrane curvature. J Cell Biol. 193(2): 333–346. ArticlePubMedPMCPDF
- Xu P, Qiao K, Ahn WS, Stephanopoulos G. 2016. Engineering Yarrowia lipolytica as a platform for synthesis of drop-in transportation fuels and oleochemicals. Proc Natl Acad Sci USA. 113(39): 10848–10853. ArticlePubMedPMC
- Yanagibashi S, Bamba T, Kirisako T, Kondo A, Hasunuma T. 2024a. Beneficial effect of optimizing the expression balance of the mevalonate pathway introduced into the mitochondria on terpenoid production in Saccharomyces cerevisiae. J Biosci Bioeng. 137(1): 16–23. Article
- Yanagibashi S, Bamba T, Kirisako T, Kondo A, Hasunuma T. 2024b. The potency of mitochondria enlargement for mitochondria-mediated terpenoid production in yeast. Appl Microbiol Biotechnol. 108(1): 110.ArticlePDF
- Yang S, Chen R, Cao X, Wang G, Zhou YJ. 2024. De novo biosynthesis of the hops bioactive flavonoid xanthohumol in yeast. Nat Commun. 15(1): 253.ArticlePubMedPMCPDF
- Yao Z, Zhou P, Su B, Su S, Ye L, et al. 2018. Enhanced isoprene production by reconstruction of metabolic balance between strengthened precursor supply and improved isoprene synthase in Saccharomyces cerevisiae. ACS Synth Biol. 7(9): 2308–2316. ArticlePubMed
- Ye C, Hong H, Gao J, Li M, Gou Y, et al. 2024. Characterization and engineering of peroxisome targeting sequences for compartmentalization engineering in Pichia pastoris. Biotechnol Bioeng. 121(7): 2091–2105. ArticlePubMed
- Yee DA, DeNicola AB, Billingsley JM, Creso JG, Subrahmanyam V, et al. 2019. Engineered mitochondrial production of monoterpenes in Saccharomyces cerevisiae. Metab Eng. 55: 76–84. ArticlePubMedPMC
- Yocum HC, Bassett S, Da Silva NA. 2022. Enhanced production of acetyl-CoA-based products via peroxisomal surface display in Saccharomyces cerevisiae. Proc Natl Acad Sci USA. 119(48): e2214941119. ArticlePubMedPMC
- Yofe I, Soliman K, Chuartzman SG, Morgan B, Weill U, et al. 2017. Pex35 is a regulator of peroxisome abundance. J Cell Sci. 130(4): 791–804. ArticlePubMedPMCPDF
- Yuan J, Ching CB. 2016. Mitochondrial acetyl-CoA utilization pathway for terpenoid production. Metab Eng. 38: 303–309. ArticlePubMed
- Yuan J, Mishra P, Ching CB. 2016. Metabolically engineered Saccharomyces cerevisiae for branched-chain ester production. J Biotechnol. 239: 90–97. ArticlePubMed
- Zhang Y, Cao X, Wang J, Tang F. 2022. Enhancement of linalool production in Saccharomyces cerevisiae by utilizing isopentenol utilization pathway. Microb Cell Fact. 21: 212.ArticlePubMedPMCPDF
- Zhang C, Chen C, Bian X, Zhang J, Zhang Z, et al. 2024. Construction of an orthogonal transport system for Saccharomyces cerevisiae peroxisomes to efficiently produce sesquiterpenes. Metab Eng. 85: 84–93. ArticlePubMed
- Zhang C, Li M, Zhao GR, Lu W. 2020. Harnessing yeast peroxisomes and cytosol acetyl-CoA for sesquiterpene α-humulene production. J Agric Food Chem. 68(5): 1382–1389. ArticlePubMed
- Zhang Y, Su M, Chen Y, Wang Z, Nielsen J, et al. 2023. Engineering yeast mitochondrial metabolism for 3-hydroxypropionate production. Biotechnol Biofuels Bioprod. 16: 64.ArticlePubMedPMCPDF
- Zhang Y, Wang J, Cao X, Liu W, Yu H, et al. 2020. High-level production of linalool by engineered Saccharomyces cerevisiae harboring dual mevalonate pathways in mitochondria and cytoplasm. Enzyme Microb Technol. 134: 109462.ArticlePubMed
- Zhang C, Wu J, Sun Q, Ding S, Tao H, et al. 2024. De novo production of bioactive sesterterpenoid ophiobolins in Saccharomyces cerevisiae cell factories. Microb Cell Fact. 23: 129.ArticlePubMedPMCPDF
- Zhang L, Yang H, Xia Y, Shen W, Liu L, et al. 2022. Engineering the oleaginous yeast Candida tropicalis for α-humulene overproduction. Biotechnol Biofuels Bioprod. 15: 59.ArticlePubMedPMCPDF
- Zhang Q, Zeng W, Xu S, Zhou J. 2021. Metabolism and strategies for enhanced supply of acetyl-CoA in Saccharomyces cerevisiae. Bioresour Technol. 342: 125978.ArticlePubMed
- Zhao EM, Zhang Y, Mehl J, Park H, Lalwani MA, et al. 2018. Optogenetic regulation of engineered cellular metabolism for microbial chemical production. Nature. 555(7698): 683–687. ArticlePubMedPMCPDF
- Zhou YJ, Buijs NA, Zhu Z, Gómez DO, Boonsombuti A, et al. 2016. Harnessing yeast peroxisomes for biosynthesis of fatty-acid-derived biofuels and chemicals with relieved side-pathway competition. J Am Chem Soc. 138(47): 15368–15377. ArticlePubMed
- Zhou P, Zhou X, Yuan D, Fang X, Pang X, et al. 2023. Combining protein and organelle engineering for linalool overproduction in Saccharomyces cerevisiae. J Agric Food Chem. 71(26): 10133–10143. ArticlePubMedLink
- Zhu ZT, Du MM, Gao B, Tao XY, Zhao M, et al. 2021. Metabolic compartmentalization in yeast mitochondria: burden and solution for squalene overproduction. Metab Eng. 68: 232–245. ArticlePubMed
References
Figure & Data
References
Citations

- Peroxisome engineering in yeast: Advances, challenges, and prospects
Cuifang Ye, Xiaoqian Li, Tao Liu, Shiyu Li, Mengyu Zhang, Yao Zhao, Jintao Cheng, Guiling Yang, Peiwu Li
Biotechnology Advances.2026; 86: 108747. CrossRef - Building an expanded bio-based economy through synthetic biology
Andrea M. Garza Elizondo, Ilenne del Valle Kessra, Erica Teixeira Prates, Evan Komp, Elise K. Phillips, Nandhini Ashok, Daniel A. Jacobson, Erin G. Webb, Yannick J. Bomble, William G. Alexander, Joanna Tannous, Chung-Jui Tsai, Wayne A. Parrott, Xiaohan Ya
Biotechnology Advances.2026; 87: 108775. CrossRef - Productive chaos and precision engineering: decoupling discovery from manufacturing to revolutionize plant-inspired therapeutics
Dexter Achu Mosoh
Frontiers in Plant Science.2026;[Epub] CrossRef - Advancing microbial engineering through synthetic biology
Ki Jun Jeong
Journal of Microbiology.2025; 63(3): e2503100. CrossRef - Metabolic engineering strategies for constructing methylotrophic cell factories
Pei Zhou, Yang Sun, Yinbiao Xu, Yupeng Liu, Hua Li
Systems Microbiology and Biomanufacturing.2025; 5(4): 1371. CrossRef
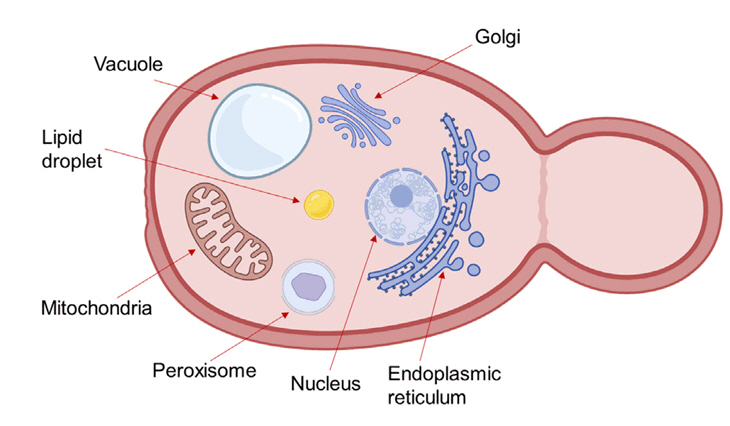
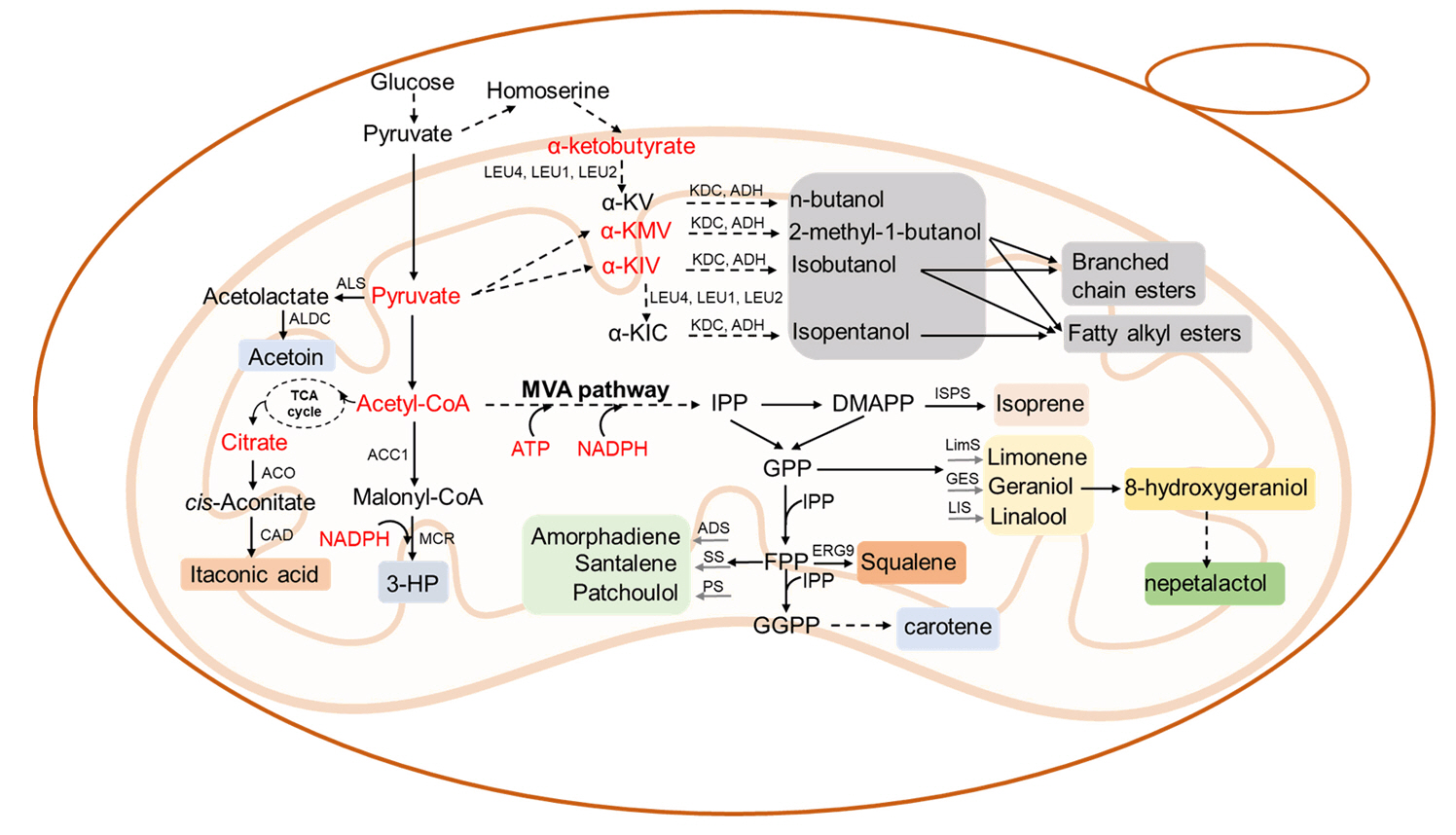
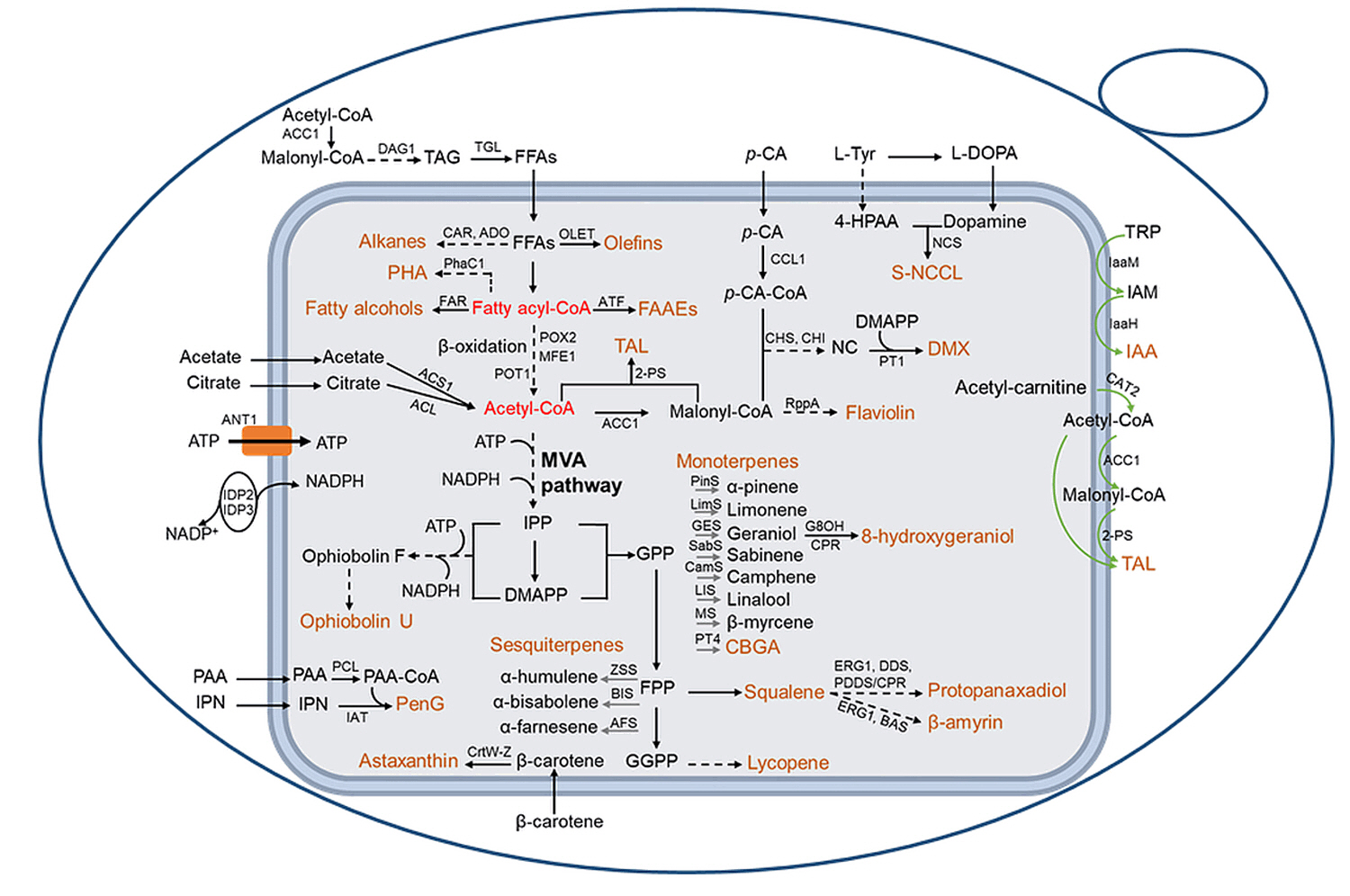
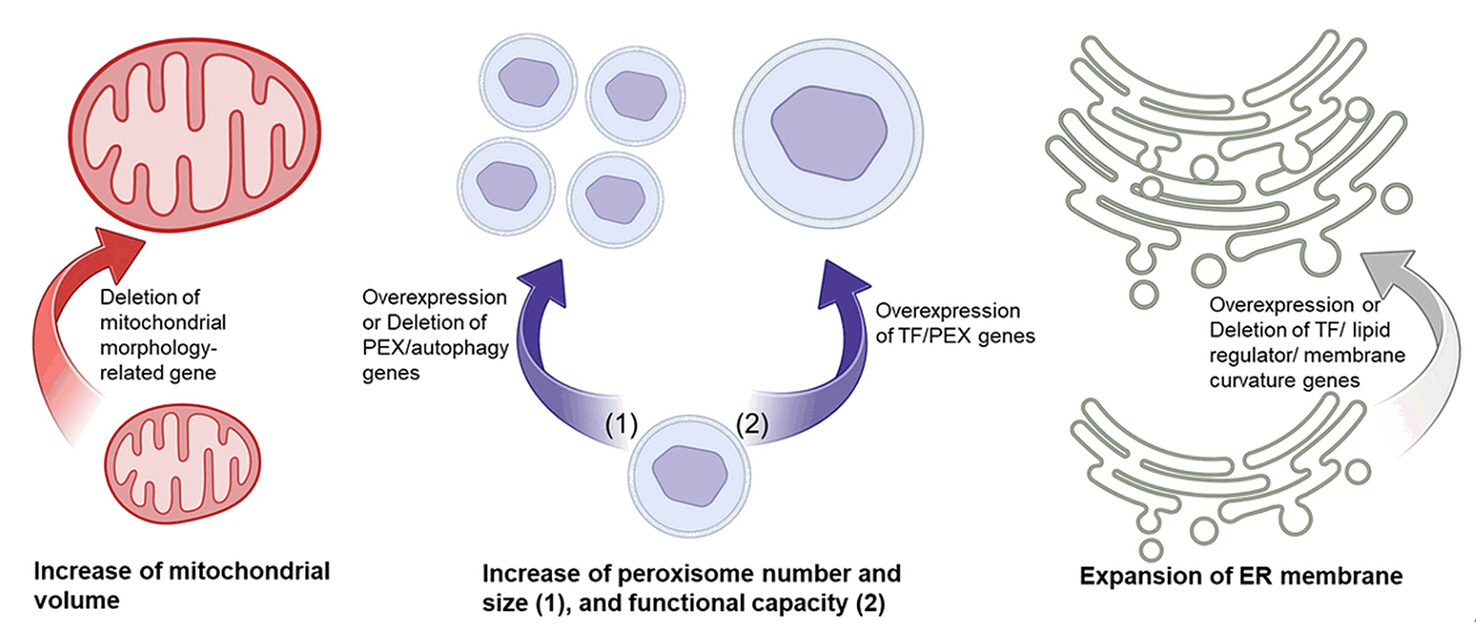

Fig. 1.
Fig. 2.
Fig. 3.
Fig. 4.
Fig. 5.
| Product | Yeast species | MTS type | Single/Dual pathway |
Engineering description | Titer | Scale | References | |
|---|---|---|---|---|---|---|---|---|
| Biochemical | Isoprene | S. cerevisiae | COX4 | Dual | · Complete MVA pathway in mitochondria (two copies of tHMG1) | 2.53 g/L | Bioreactor | Lv et al. ( |
| · Diploid strain formation by mating the mitochondrial engineered strain with YXM10 strain1 | ||||||||
| Dual | · Isoprene synthase mutant ISPSLN in mitochondria/cytosol | 11.09 g/L | Bioreactor | |||||
| · Complete MVA pathway in mitochondria/cytosol | ||||||||
| · Additional expression of MVD1 and IDI1 in mitochondria/cytosol | ||||||||
| · Diploid strain formation by mating 2 mitochondrial/cytosolic strains | ||||||||
| Amorpha-4,11-diene | COQ3, COX4 | Single | · Entire amorphadiene pathway targeted in mitochondria (only ADS tagged with COQ3-MTS) | 430 mg/L | Flask | |||
| 8-Hydroxygeraniol | COX4 | Single | · Choice of a starting strain JHY65 for its improved respiratory growth and increased mitochondrial stability2 | 227 mg/L; | Bioreactor | |||
| · The geraniol pathway targeted in mitochondria (using Gg mFPS and ObGES) | ||||||||
| · Expression of ER-targeted CrG8H | ||||||||
| · Deletion of OYE2 and OYE3 | ||||||||
| Nepetalactol | · Expression of cytosolic CrGOR and CrISY for nepetalactol synthesis | 5.9 mg/L | Tube | |||||
| Linalool | COX4 | Dual | · Complete linalool pathway in both mitochondria and cytosol (using ERG20F96W/N127W and CoLIS) | 23.45 mg/L | Bioreactor | |||
| · Downregulation of endogenous ERG20 | ||||||||
| · Additional expression of a fusion CoLIS-ERG20F96W/N127W protein in both mitochondria and cytosol. | ||||||||
| Single | · The entire MVA pathway targeted in mitochondria | 23.8 mg/L | Flask | |||||
| · Mitochondrial expression of truncated AaLS1 and ERG20F96W/N127W | ||||||||
| Limonene | COX4 | Dual | · The entire limonene pathway targeted in mitochondria | 2.63 g/L | Bioreactor | |||
| · The entire cytosolic limonene pathway with multiple copies of ERG20F96W/N127W and tLimS, and downregulation of ERG20 | ||||||||
| · Enhanced acetyl-CoA pool by knocking out CIT2 and MLS1 | ||||||||
| · Enhancing NADPH pool by overexpressing pentose phosphate (PP) pathway and deleting the NADPH-consuming pathway | ||||||||
| α-Santalene | MMF1, COX4, HSP60, LSC2, LDP1, ALD4 | Single | · Complete α-santalene pathway targeted in mitochondria with additional overexpression of tHMG1 | 41 mg/L | N/A | |||
| Squalene | COX4 | Dual | · Complete MVA pathway targeted in mitochondria with additional expression of ERG19 | 21.1 g/L | Bioreactor | |||
| · Full squalene pathway expressed in cytosol (overexpression only for ERG10, tHMG1, ERG19, IDI1, ERG20, and ERG9) | ||||||||
| · Expression of ALD6, ASC1, ASC2 and ADH2 for improved cytosolic acetyl-CoA | ||||||||
| · Down regulation of ERG1 | ||||||||
| COX4 | Dual | · Complete MVA pathway targeted in mitochondria with regulation of genes by promoter screening | 230 mg/L | Flask | ||||
| · Full squalene pathway in cytosol with additional copies of tHMG1 and ERG12 | ||||||||
| COX4 | Single | · Complete MVA pathway targeted in mitochondria | 707 nmol/g DCW | Flask | ||||
| · Improved mitochondrial volume by deletion of MDM32 | ||||||||
| β-Carotene | COX4 | Single | · Expression of CrtYBXd, CrtIXd, and BTS1 (for β-carotene synthesis) | 1609 nmol/g DCW | Flask | |||
| 3-Hydroxypropionate | COX4 | Single | · Mitochondrial targeting of mutant ACC1 and dissected MCR | 71.09 g/L | Bioreactor | |||
| · Overexpression of IDP1 and POS5 for improved NADPH pool in mitochondria | ||||||||
| Itaconic acid | A. niger | ICDA, ACOA | Dual | · Itaconic acid pathway targeted in mitochondria (AtCadA and AnAcoA) | 1.4 g/L | Flask | ||
| · Expression of itaconic acid pathway in cytosol (AtCadA and EcAcnA) | ||||||||
| Acetoin | C. glabrata | COX4 | Single | · Acetoin pathway targeted in mitochondria (BsALS and BaALDC) | 3.26 g/L | Flask | ||
| · Overexpression of mitochondrial pyruvate carrier (ScMPC) | ||||||||
| Biofuel | Isobutanol | S. cerevisiae | COX4 | Single | · Mitochondrial targeting of ARO10, LlKiVD and AdhARE1, ADH7 | 635 mg/L | Tube | |
| · Overexpression of mitochondrially native genes ILV2, ILV5, ILV3 | ||||||||
| Single | · Mitochondrial targeting of ARO10, LlAdhARE1 | 1.245 g/L | Plate | |||||
| · Deletion of BAT1 and ILV6 | ||||||||
| Single | · Mitochondrial targeting of ARO10, LlAdhARE1 | 8.49 g/L | Bioreactor | |||||
| · Deletion of BAT1 | ||||||||
| 2-Methyl-1-butanol | Single | · Optogenetic regulation of ILV2 and PDC1 | 2.38 g/L | Bioreactor | ||||
| Isopentanol | COX4, CD9, COX6 | Single | · Mitochondrial targeting of LEU4 mutant, LEU1, LEU2 | 1.24 g/L | Plate | |||
| · Deletion of LEU4, LEU9, BAT1 and OAC1 | ||||||||
| n-Butanol | COX4, CYB2, CAT2 | Single | · Mitochondrial targeting of LEU4, LEU1, LEU2 and citramalate synthase (LiCim) | 1.05 g/L | Bioreactor | |||
| · Overexpression of LEU9, cysteine desulfurase NFS1, ARO10 and ADH7 | ||||||||
| Fatty acid alkyl ester | COX4 | Single | · Mitochondrial targeting of ARO10 and ADH7 for improved isobutanol production | 230 mg/L | Flask | |||
| · Overexpression of two wax ester synthase Ms Ws2 and Maqu_0168 | ||||||||
| · Deletion of OPI1 and RPD3 | ||||||||
| Isobutyrate | COX4 | Dual | · Mitochondrial targeting of ARO10 and ADH7, and ATF1 | 260.2 mg/L | Tube | |||
| 3-Methyl-1-butyl acetate | · ATF1 overexpression in cytosol | 296.1 mg/L | ||||||
| 2-Methyl-1-butyl acetate | · Overexpression of PYC2, MDH2, MAE1 | 289.6 mg/L | ||||||
| Product | Yeast species | PTS type | Single/Dual pathway |
Engineering description | Titer | Scale | References | |
|---|---|---|---|---|---|---|---|---|
| Biochemical | Geraniol | S. cerevisiae | SKL | Single | · Compartmentalization of AgGPPS2 and ObGES in peroxisome | 2.75 mg/L | Plate | |
| · Deletion of PEX30 and PEX32 for increased peroxisome number | ||||||||
| · Expression of truncated α-arrestin-like adaptor Bul1 for high tolerance to geraniol | ||||||||
| · The entire geraniol pathway targeted in peroxisome (using ERG20N127Wand ObGerS) with episomal plasmids | 5.52 g/L | Bioreactor | ||||||
| Dual | · Machine learning (ML)-aided identification of the MVA pathway’s critical gene | 120 mg/L | Flask | |||||
| · The MVA pathway targeted in peroxisome | ||||||||
| · The dual MVA platform strain (diploid) created by mating peroxisomal and cytosolic strains | ||||||||
| · Overexpression of a fusion tObGES-ERG20WW | ||||||||
| ePTS1 | Single | · The entire geraniol pathway targeted in peroxisome (using ERG20F96W, N127W and tVoGES) | 9.5 g/L | Flask | ||||
| · ML-guided screen of peroxisome complexes for improved peroxisome capacity | ||||||||
| P. pastoris | ePTS1, PTS of TAL1-2 | Single | · HTS-based screen of 25 putative PTS, followed by directed evolution of selected 3 PTS1 with high targeting efficiency | 20 mg/L | Plate | |||
| · Peroxisomal compartmentalization of geraniol pathway (only tCrGES tagged with newly identified PTS; the remaining proteins tagged with ePTS1). | ||||||||
| R-Limonene | R. toruloides | SKL, PTS2 (C4N11 of 3-KT) | Dual | · Peroxisomal targeting of the entire limonene pathway (two copies of NPPS::LS) | 1.05 g/L | Flask | ||
| · Peroxisomal targeting of Nph17 and overexpressing Nph17 and ACCT for improved acetoacetyl-CoA in peroxisome and cytosol, respectively | ||||||||
| · Cytosolic overexpression of NPPS::LS | ||||||||
| · Overexpression of ACL for improved cytosolic acetyl-CoA | ||||||||
| S. cerevisiae | SKL | Single | · Peroxisomal targeting of the entire biosynthetic pathway targeted with episomal plasmids. | 2.58 g/L | Bioreactor | |||
| · R-limonene: ClLimS | ||||||||
| α-Pinene | · α-Pinene: PtPinS | 69.22 mg/L | Flask | |||||
| Sabinene | · Sabinene: SpSabS | 32.32 mg/L | ||||||
| Camphene | · Camphene: 1SeCamS | 5.77 mg/L | ||||||
| 8-Hydroxygeraniol | · 8-Hydroxygeraniol: ObGerS, CrG8OH, CrCPR | 25.11 mg/L | ||||||
| Canabigerolic acid (CBGA) | · CBGA: CsPT4 | 0.82 mg./L | ||||||
| Linalool | ePTS1 | Dual | · Improving catalytic efficiency toward GPP of linalool synthase through site-directed mutagenesis | 2.6 g/L | Bioreactor | |||
| · Complete linalool pathway targeting to peroxisome | ||||||||
| · Overexpression of tHMG1, IDI1, ERG20F96W, N127W, and McLIS with repression of native ERG20 for cytosolic linalool pathway | ||||||||
| · Mating peroxisomal and cytosolic strain to form the dual strain (diploid) | ||||||||
| β-Myrcene | SKL | Dual | · Source screening of β-myrcene synthase, with truncated QiMS as the best candidate | 142.64 mg/L | Bioreactor | |||
| · Peroxisome localization of fusion enzyme tQiMSD436N-ERG20WW | ||||||||
| · Overexpression of tHMG1 and IDI in the cytosolic MVA pathway and replacement of native ERG20 by ERG20W | ||||||||
| · Enhancing cytosolic acetyl-CoA by overexpressing ADH2 and ALD6 | ||||||||
| α-Humulene | S. cerevisiae | ePTS1 | Dual | · Compartmentalization of entire α-humulene pathway in peroxisome | 1.73 g/L | Bioreactor | ||
| · Overexpression of tHMG1 and ERG20 in cytosol | ||||||||
| · Down regulation of ERG9 by replacing its native promoter by HXT1p and adding PEST sequence | ||||||||
| · Additional overexpression of cytosolic ZzZSS1 in rDNA sites | ||||||||
| SKL | Dual | · ML-guided dual MVA platform strain (diploid) created by mating peroxisomal and cytosolic strains | ~22.5 mg/L | Flask | ||||
| · Overexpression of cytosolic ERG20 and ZzZSS1 | ||||||||
| oPTS1* | Dual | · Develop an orthogonal transport system ScPEX5*-oPTS1* | 17.33 g/L | Bioreactor | ||||
| · rDNA integration of the entire cytosolic and peroxisomal α-humulene pathway (ZSS1 as humulene synthase) | ||||||||
| · Overexpression of ANT1, IDP2, IDP3 (rDNA integration) for improved ATP and NADPH pool in peroxisome | ||||||||
| · Down regulation of ERG9 by replacing its native promoter by HXT1p and adding PEST sequence | ||||||||
| Y. lipolytica | PTS (GGGSSKL) | Single | · Localization of entire α-humulene pathway in peroxisome (NADH-HMG1 as HMG-CoA reductase; AcHS2 as humulene synthase) | 3.2 g/L | Bioreactor | |||
| · Increasing ATP supply in peroxisome by overexpression of ANT1 | ||||||||
| · Adjusting the copy numbers of rate-limiting enzymes (one more copy of peroxisomal targeting NADH-HMG1 and AcHS2) | ||||||||
| · Upregulation of β-oxidation by overexpressing POT1 | ||||||||
| · Oleic acid as a possible C source for α-humulene accumulation | ||||||||
| PTS (GGGSSKL) | Dual | · Iterative integration of the entire cytosolic and peroxisomal α-humulene pathway (AcHS2 as humulene synthase) | 21.7 g/L | Bioreactor | ||||
| · Mediating copy number of cytosolic tHMG1 and AcHS2 | ||||||||
| · Improving cytosolic acetyl-CoA by overexpression of CoA‐acetylating aldehyde dehydrogenase (EcAAD) | ||||||||
| · Repression of ERG9 expression using Cu2+ repressible promoter | ||||||||
| Candida tropicalis | SKL | Single | · Multiple overexpression of the entire peroxisomal α-humulene pathway (double copies of the pathway) | 2.42 mg/L | Flask | |||
| α-Bisabolene | P. pastoris | SKL and LARF | Single | · Peroxisome isolation of the entire α-bisabolene pathway with fusion enzyme of FPPS-AgBIS; upregulation of acetyl-CoA-to-mevalonate pathway; and additional copy of separated FPPS and AgBIS | 1.1 g/L | Flask | ||
| α-Farnesene | ePTS1 | Dual | · Overexpression of tHMG1, IDI1, ERG20 and MdAFS for cytosolic α-farnesene pathway | 2.56 g/L | Flask | |||
| · Improving cytosolic acetyl-CoA pool by introducing pyruvate dehydrogenase (Ec cytoPDH) and ATP-dependent citrate lyase (YlACL) | ||||||||
| · Introduction of IUP pathway and downstream genes (2 copies of IDI, ERG20, MdAFS) into peroxisome | ||||||||
| β-Amyrin | S. cerevisiae | ePTS1 | Dual | · Peroxisome compartmentalization of squalene pathway | 2.6 g/L | Bioreactor | ||
| · Double copies of downstream β-amyrin pathway (ERG1 and GgbAS1) in cytosol | ||||||||
| · Deregulation of ERG7 | ||||||||
| Squalene | ePTS1 | Dual | · The entire squalene pathway targeted in peroxisome with an additional copy of tHMG1 and IDI1 | 11 g/L | Bioreactor | |||
| · Deletion of GPD1 and GPD2 | ||||||||
| · Overexpression of ANT1, IDP2 and IDP3 for improved ATP and NADPH pool in peroxisome | ||||||||
| · ACS1, YlACL1 and YlACL2 targeted to peroxisome for improved peroxisomal acetyl-CoA | ||||||||
| · Diploid strain formation by mating two peroxisomal and cytosolic strains | ||||||||
| SKL | Dual | · The dual MVA platform strain (diploid) created by mating peroxisomal and cytosolic MVA strains | 300 mg/L | Flask | ||||
| · Overexpression of ERG9 and ERG20 | ||||||||
| · Prevention of squalene degradation by supplementation of terbinafine | ||||||||
| Y. lipolytica | SKL | Dual | · The complete squalene pathway targeted in peroxisome | 32.8 g/L (on glucose)/ 31.6 g/L (on acetate) | Bioreactor | |||
| · Enhanced TAG-derived free fatty acids in peroxisome through overexpressing TlTGL, DGA1 and ACC1 | ||||||||
| · Improved cytosolic acetyl-CoA pool by overexpressing PYC1, YHM2 and MmACL | ||||||||
| · Upregulated β-oxidation by overexpressing POX2, MFE1 and POT1 | ||||||||
| · Increased size/number of peroxisomes by overexpressing PEX10 | ||||||||
| · Introduction of Acetyl-CoA shortcut in the peroxisome by targeting 2SeACS1L641P. | ||||||||
| Lycopene | P. pastoris | SKL | Single | · Peroxisomal compartmentalization of lycopene biosynthetic pathway (CrtE, CrtB, CrtI) | 73.9 mg/L | Bioreactor | ||
| Astaxanthin | Y. lipolytica | SKL | Single | · Peroxisomal targeting of the fusion enzyme PsCtrW-HpCrtZ | 58.7 mg/L | Flask | ||
| Protopanaxadiol | S. cerevisiae | PTS1, PTS2 | Single | · Regulating number and size of peroxisome through deletion of PEX11 and ATG36, and overexpression of PEX34 | 4.1 mg/L | Flask | ||
| · Peroxisomal compartmentalization of protopanaxadiol pathway (including tHMG1, ERG9, PgErg1, PgDs, PgPpds, and PgCpr) | ||||||||
| (S)-Norcoclaurine | ePTS1 | Single | · Targeting norcoclaurine synthase (NCS) in peroxisome | ~7 mg/L | Plate | |||
| · Controlling peroxisome biogenesis by engineering transcriptional factors ADR1, OAF1 and PIP2 | ||||||||
| Ophiobolins U | PTS of PEX15 | N/A | · Peroxisomal surface display of CAT2 for improved cytosolic acetyl-CoA | 128.9 mg/L | Flask | |||
| Indole-3-acetic acid | K. marxianus | PTS of PEX15 | Single | · Peroxisomal surface display of tryptophan-2-monooxygenase PsIaaM and indole-acetamide hydrolase PsIaaH | ~61.25 mg/L | Tube | ||
| · Controlling peroxisome number and size by overexpression of PEX19 | ||||||||
| Triacetic acid lactone (TAL) | PTS of PEX15 | Single | · Peroxisomal surface display of CAT2, ACC1 and Gh2-PS | ~ 1.1 g/L | Tube | |||
| · Controlling peroxisome number and size by overexpression of PEX11 | ||||||||
| S. cerevisiae | PTS of PEX15 | Single | · Peroxisomal surface display of CAT2, ACC1S1157A and Gh2-PS | 0.77 g/L | Tube | |||
| pPTS (PTS of Psc60) | Dual | · Cytosolic expression and peroxisomal targeting of Gh2-PS and ACC1S686A, S659A, S1157A | 0.14 g/L | Plate | ||||
| · Increasing peroxisome proliferation by overexpression of ADR1, OAF1 and PIP2 | ||||||||
| Flaviolin | ePTS1 | Single | · Peroxisomal targeting of Sac_RppA and ACC1S686A, S659A, S1157A | N/A | N/A | |||
| Desmethylxanthohumol (DMX) | N/A | Single | · Compartmentalization of the complete MVA pathway in peroxisome | ~62.5 µg/L | Flask | |||
| · Peroxisomal targeting of ACC1 and naringenin chalcone and DMX biosynthetic genes | ||||||||
| Amino acid (Lysine, Histidine) | S. japonicus | PTS1 and PTS2 | N/A | · Peroxisomal targeting of GPD2, LYS3 and HIS2 enzymes | N/A | N/A | ||
| · Regulating peroxisome size with PEX5 mutant and PEX11 | ||||||||
| Penicillin | Hansenula polymorpha | SKL | Single | · Peroxisomal targeting of penicillin pathway’s bottom genes, including PcIAT and PcPCL | 1 mg/L | Plate | ||
| PHA | S. cerevisiae | PTS of BnICL | Single | · Peroxisomal targeting of PaPHAC1 synthase | 0.45 g/g DCW | Flask | ||
| Biofuel | Fatty acid ethyl esters (FAEEs) | Y. lipolytica | SKL | Single | · Peroxisomal targeting of wax ester synthase (AbAtfA) | 110.9 mg/L | Flask | |
| Fatty alcohols | S. cerevisiae | PTS2 | Single | · Peroxisomal targeting of fatty acyl-CoA reductase (TaFAR) | 1.3 g/L | Tube | ||
| · Increasing peroxisomal importing rate by overexpression of PEX7 | ||||||||
| · Improved malonyl-CoA pool by overexpression of ACC1 | ||||||||
| PTS2 | Single | · Peroxisomal targeting of fatty acyl-CoA reductase MaFAR in fatty acyl-CoA overproducing strain | 193 mg/L | Flask | ||||
| Alkanes | PTS1 and PTS2 | Single | · Peroxisomal targeting of free fatty acid (FFA) pathway including MmCAR, AnNPGA, 3SeADO, ferredoxin 3SeFd and ferredoxin reductase 3SeFNR | ~ 3.5 mg/L | Flask | |||
| · Increasing fatty acid production by deleting HFD1 and POX1 | ||||||||
| · Increasing peroxisome population by deletion of PEX31, 32 and overexpression of PEX34 | ||||||||
| Olefins | PTS2 | · Peroxisomal targeting of OleT with flavodoxin EcFldA and flavodoxin reductase EcFNR | ~ 0.20 mg/L | Flask | ||||
| α-Alkenes | P. pastoris | PTS1 and PTS2 | Single | · Peroxisomal targeting of decarboxylase PfUndB together with cofactor protein putidaredoxin–putidaredoxin reductase PpPdr/Pdx | 1.6 mg/L | Flask | ||
| Product | Yeast species | ER engineering | Titer | Scale | References | |
|---|---|---|---|---|---|---|
| Biochemical | Opiate (morphine) | S. cerevisiae | · ER targeting of NADPH-dependent aldo-keto reductase PsCOR1.3 with ER signal peptide from CNE1 | 3.1 mg/L | Plate | |
| Ceramide NS | S. cerevisiae | · ER targeting of human sphingolipid desaturase (hDES1) with ER retention signal KKEK from SUR2 | N/A | N/A | ||
| Trans-nootkatol | S. cerevisiae | · Regulating ER membrane proliferation by overexpression of ICE2 | 30 mg/L | Flask | ||
| 1-Hydroxybufuralol | P. pastoris | N/A | N/A | |||
| β-Amyrin | S. cerevisiae | · Expansion of ER by deleting the phosphatidic acid phosphatase PAH1 | N/A | N/A | ||
| Medicagenic acid | 27.1 mg/L | Flask | ||||
| Oleanane-type sapogenin | N/A | N/A | ||||
| Squalene | S. cerevisiae | · Expansion of ER by overexpression of transcriptional factor INO2 | 634 mg/L | Flask | ||
| Protopanaxadiol | S. cerevisiae | · Controlling gene dosage of tHMG1 (3 copies) for squalene production | 12.1 mg/L | Flask | ||
| Astaxanthin | Y. lipolytica | · ER targeting of the fusion enzyme β-carotene ketolase-hydroxylase (PsCtrW-HpCrtZ) with ER signal peptide KDEL | 53.2 mg/L | Flask | ||
| Human antibody (IgG) | S. cerevisiae | · Expansion of ER by deleting lipid-regulator OPI1 | 126 µg/L | Plate | ||
| · Overexpression of peptidyl-prolyl isomerase CPR5 for enhanced secretion | ||||||
| · Modification of ER morphology and size by deleting lipid-regulator gene OPI1 and ER membrane curvature genes RTN1, RTN2 and YOP1 | 79 ng/mL | Plate | ||||
| Ovalbumin | S. cerevisiae | · Expression of OVA using signal peptide from INU1 | 116.3 mg/L | Bioreactor | ||
| · Co-expression of chaperone Kar2 and disulfide isomerase PDI for improved folding environment in ER | ||||||
| · Expansion of ER by deleting OPI1 and overexpressing transcriptional factors INO2 and INO4 | ||||||
| S-Scoulerine | S. cerevisiae | · ER targeting of berberine bridge enzyme (CyBBE) with ER C-terminal HDEL signal peptide (CyBBE_ERTS) | 113.1 mg/L | Flask | ||
| · ER targeting of mammalian peroxiredoxin IV (PRDX4) with N- and C-terminal ERTS (α- mPRDX4_ERTS) for reducing toxicity of hydrogen peroxide | ||||||
| · Expansion of ER by deleting transcriptional regulator OPI1 of phospholipid biosynthesis | ||||||
| Biofuel | Fatty acid ethyl esters (FAEEs) | Y. lipolytica | · ER targeting of wax-ester synthase (AbAtfA) with ER peptide signal KDEL | 136.5 mg/L | Flask | |
| Fatty alkane | · ER targeting of fatty acyl-CoA reductase (AbACR1)-aldehyde deformylating oxygenase (PmADO) gene cluster with ER signal peptide KDEL | 16.8 mg/L | Flask | |||
| Fatty alcohol | · ER targeting of fatty acyl-CoA reductase (AbACR1) and aldehyde reductase (EcAHR) with ER signal peptide KDEL | 49.2 mg/L | Flask | |||
The single pathway refers to the entire biosynthetic pathway or partial downstream pathway, compartmentalized either in the cytosol or in a subcellular organelle; whereas the dual pathway refers to the entire biosynthetic pathway that is expressed in both the cytosol and a subcellular organelle. Gg:
The single pathway refers to the entire biosynthetic pathway or partial downstream pathway, compartmentalized either in the cytosol or in a subcellular organelle; whereas the dual pathway refers to the entire biosynthetic pathway that is expressed in both the cytosol and a subcellular organelle. Yl:
Ps:
Table 1.
Table 2.
Table 3.
TOP
 MSK
MSK
 ePub Link
ePub Link Cite this Article
Cite this Article






