- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Volume 63(3); 2025 > Article
-
Review
Harnessing organelle engineering to facilitate biofuels and biochemicals production in yeast - Phuong Hoang Nguyen Tran1,2, Taek Soon Lee1,2,*
-
Journal of Microbiology 2025;63(3):e2501006.
DOI: https://doi.org/10.71150/jm.2501006
Published online: March 28, 2025
1Joint BioEnergy Institute, Emeryville 94608, CA, USA
2Biological Systems & Engineering Division, Lawrence Berkeley National Laboratory, Berkeley 94720, CA, USA
- *Correspondence Taek Soon Lee tslee@lbl.gov
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Figure & Data
References
Citations

- Peroxisome engineering in yeast: Advances, challenges, and prospects
Cuifang Ye, Xiaoqian Li, Tao Liu, Shiyu Li, Mengyu Zhang, Yao Zhao, Jintao Cheng, Guiling Yang, Peiwu Li
Biotechnology Advances.2026; 86: 108747. CrossRef - Building an expanded bio-based economy through synthetic biology
Andrea M. Garza Elizondo, Ilenne del Valle Kessra, Erica Teixeira Prates, Evan Komp, Elise K. Phillips, Nandhini Ashok, Daniel A. Jacobson, Erin G. Webb, Yannick J. Bomble, William G. Alexander, Joanna Tannous, Chung-Jui Tsai, Wayne A. Parrott, Xiaohan Ya
Biotechnology Advances.2026; 87: 108775. CrossRef - Productive chaos and precision engineering: decoupling discovery from manufacturing to revolutionize plant-inspired therapeutics
Dexter Achu Mosoh
Frontiers in Plant Science.2026;[Epub] CrossRef - Advancing microbial engineering through synthetic biology
Ki Jun Jeong
Journal of Microbiology.2025; 63(3): e2503100. CrossRef - Metabolic engineering strategies for constructing methylotrophic cell factories
Pei Zhou, Yang Sun, Yinbiao Xu, Yupeng Liu, Hua Li
Systems Microbiology and Biomanufacturing.2025; 5(4): 1371. CrossRef
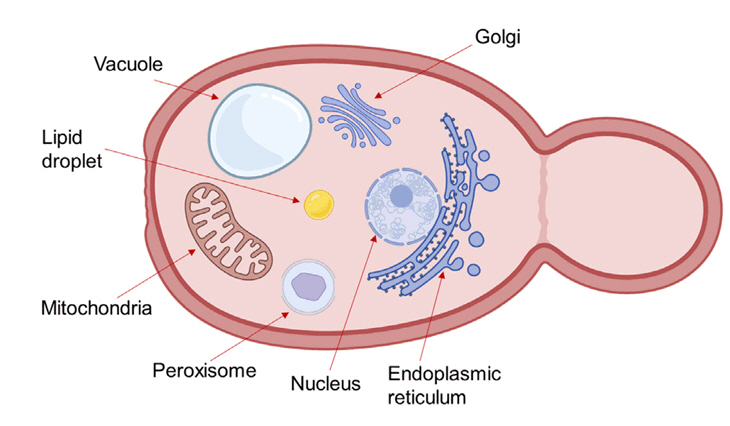
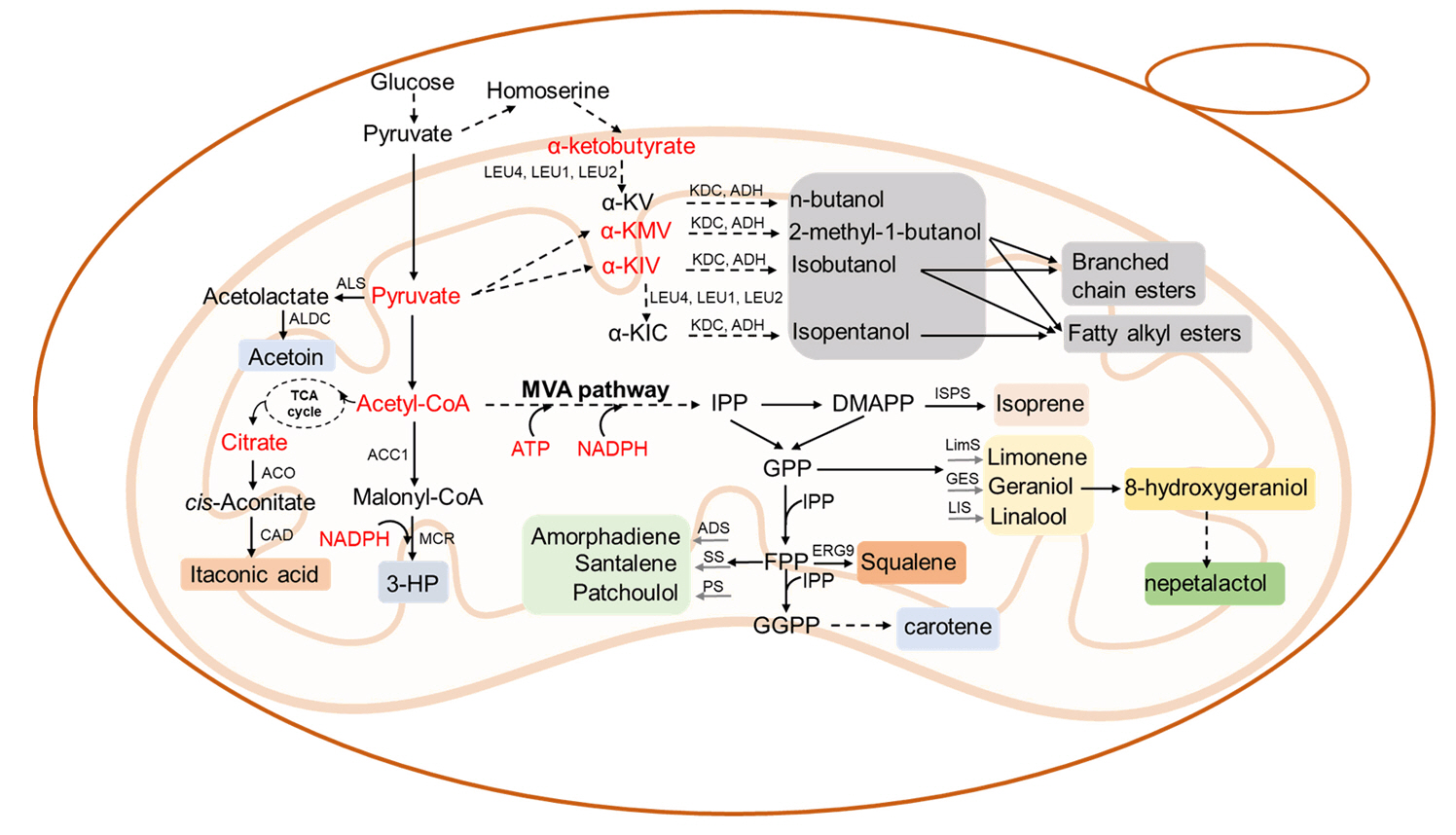
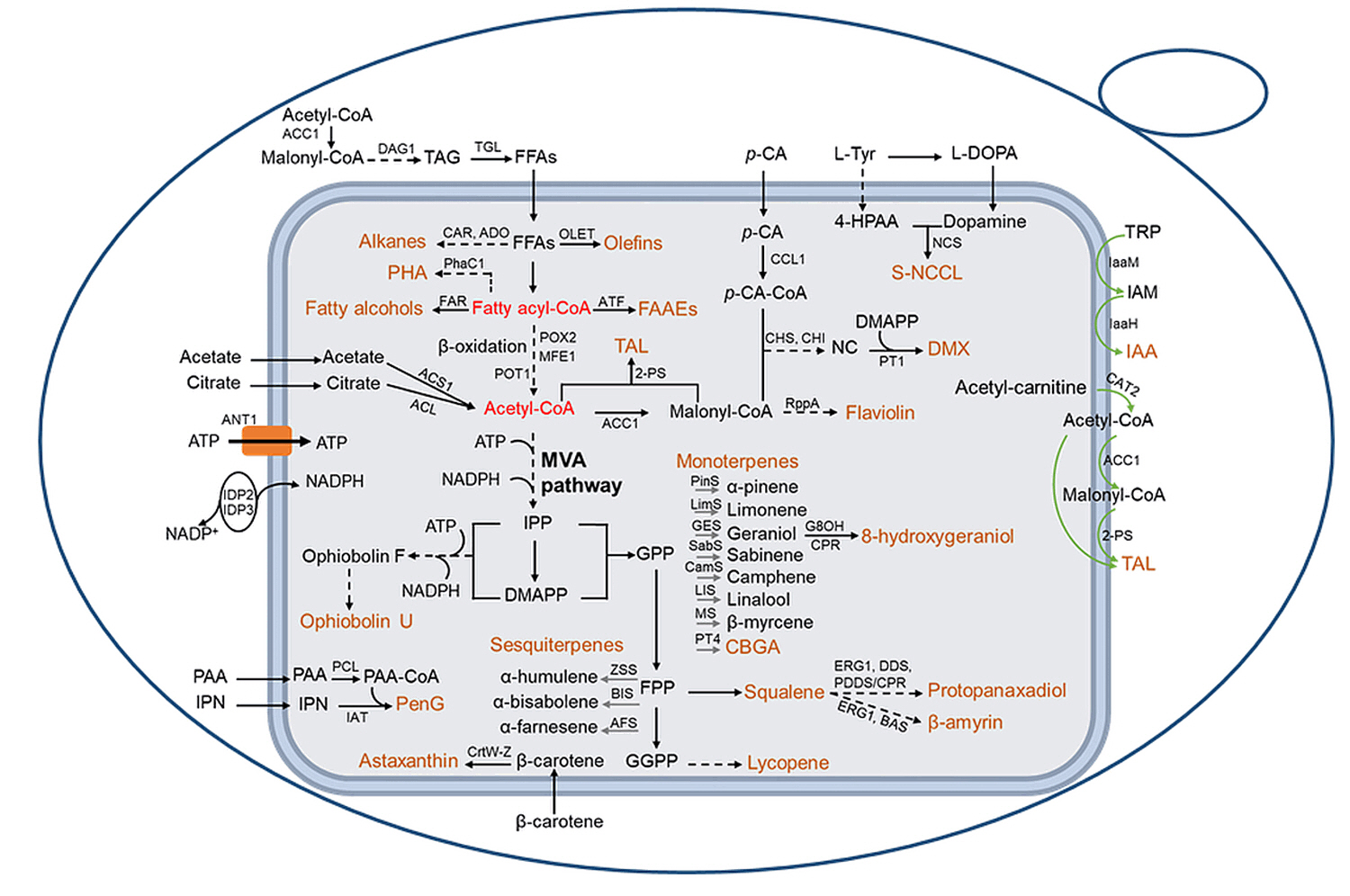
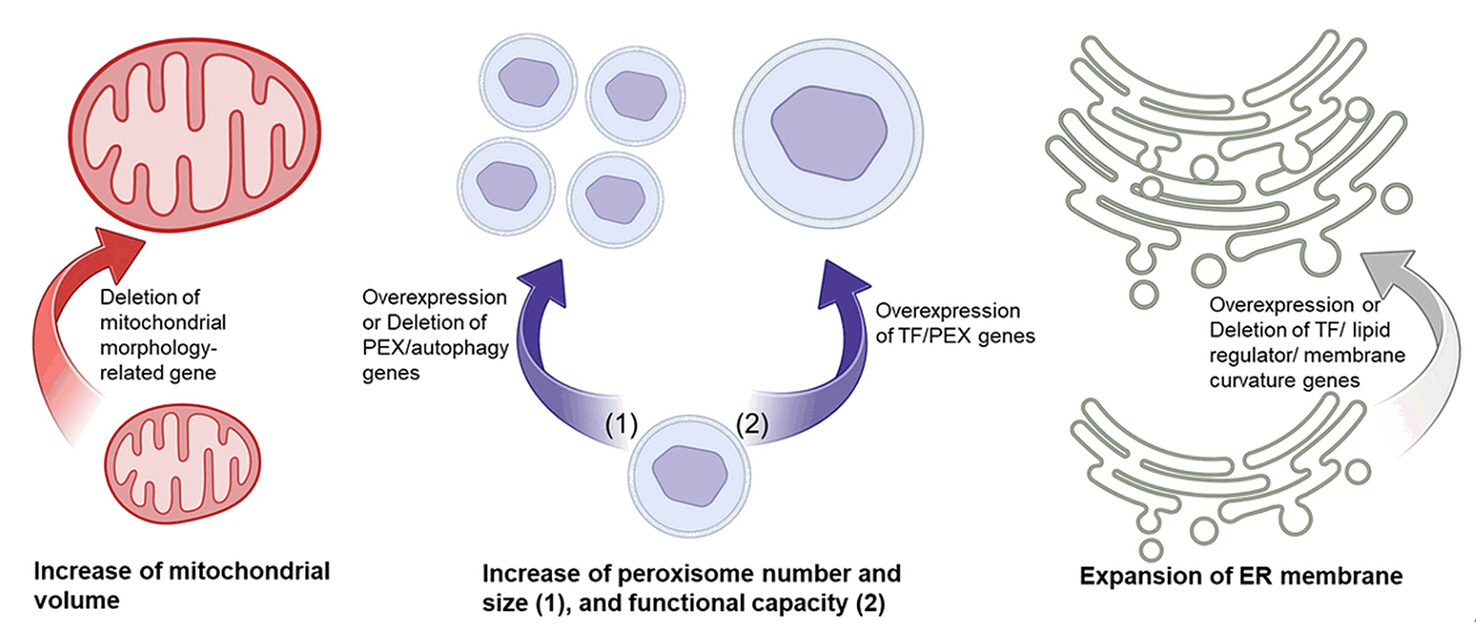

Fig. 1.
Fig. 2.
Fig. 3.
Fig. 4.
Fig. 5.
| Product | Yeast species | MTS type | Single/Dual pathway |
Engineering description | Titer | Scale | References | |
|---|---|---|---|---|---|---|---|---|
| Biochemical | Isoprene | S. cerevisiae | COX4 | Dual | · Complete MVA pathway in mitochondria (two copies of tHMG1) | 2.53 g/L | Bioreactor | Lv et al. ( |
| · Diploid strain formation by mating the mitochondrial engineered strain with YXM10 strain1 | ||||||||
| Dual | · Isoprene synthase mutant ISPSLN in mitochondria/cytosol | 11.09 g/L | Bioreactor | |||||
| · Complete MVA pathway in mitochondria/cytosol | ||||||||
| · Additional expression of MVD1 and IDI1 in mitochondria/cytosol | ||||||||
| · Diploid strain formation by mating 2 mitochondrial/cytosolic strains | ||||||||
| Amorpha-4,11-diene | COQ3, COX4 | Single | · Entire amorphadiene pathway targeted in mitochondria (only ADS tagged with COQ3-MTS) | 430 mg/L | Flask | |||
| 8-Hydroxygeraniol | COX4 | Single | · Choice of a starting strain JHY65 for its improved respiratory growth and increased mitochondrial stability2 | 227 mg/L; | Bioreactor | |||
| · The geraniol pathway targeted in mitochondria (using Gg mFPS and ObGES) | ||||||||
| · Expression of ER-targeted CrG8H | ||||||||
| · Deletion of OYE2 and OYE3 | ||||||||
| Nepetalactol | · Expression of cytosolic CrGOR and CrISY for nepetalactol synthesis | 5.9 mg/L | Tube | |||||
| Linalool | COX4 | Dual | · Complete linalool pathway in both mitochondria and cytosol (using ERG20F96W/N127W and CoLIS) | 23.45 mg/L | Bioreactor | |||
| · Downregulation of endogenous ERG20 | ||||||||
| · Additional expression of a fusion CoLIS-ERG20F96W/N127W protein in both mitochondria and cytosol. | ||||||||
| Single | · The entire MVA pathway targeted in mitochondria | 23.8 mg/L | Flask | |||||
| · Mitochondrial expression of truncated AaLS1 and ERG20F96W/N127W | ||||||||
| Limonene | COX4 | Dual | · The entire limonene pathway targeted in mitochondria | 2.63 g/L | Bioreactor | |||
| · The entire cytosolic limonene pathway with multiple copies of ERG20F96W/N127W and tLimS, and downregulation of ERG20 | ||||||||
| · Enhanced acetyl-CoA pool by knocking out CIT2 and MLS1 | ||||||||
| · Enhancing NADPH pool by overexpressing pentose phosphate (PP) pathway and deleting the NADPH-consuming pathway | ||||||||
| α-Santalene | MMF1, COX4, HSP60, LSC2, LDP1, ALD4 | Single | · Complete α-santalene pathway targeted in mitochondria with additional overexpression of tHMG1 | 41 mg/L | N/A | |||
| Squalene | COX4 | Dual | · Complete MVA pathway targeted in mitochondria with additional expression of ERG19 | 21.1 g/L | Bioreactor | |||
| · Full squalene pathway expressed in cytosol (overexpression only for ERG10, tHMG1, ERG19, IDI1, ERG20, and ERG9) | ||||||||
| · Expression of ALD6, ASC1, ASC2 and ADH2 for improved cytosolic acetyl-CoA | ||||||||
| · Down regulation of ERG1 | ||||||||
| COX4 | Dual | · Complete MVA pathway targeted in mitochondria with regulation of genes by promoter screening | 230 mg/L | Flask | ||||
| · Full squalene pathway in cytosol with additional copies of tHMG1 and ERG12 | ||||||||
| COX4 | Single | · Complete MVA pathway targeted in mitochondria | 707 nmol/g DCW | Flask | ||||
| · Improved mitochondrial volume by deletion of MDM32 | ||||||||
| β-Carotene | COX4 | Single | · Expression of CrtYBXd, CrtIXd, and BTS1 (for β-carotene synthesis) | 1609 nmol/g DCW | Flask | |||
| 3-Hydroxypropionate | COX4 | Single | · Mitochondrial targeting of mutant ACC1 and dissected MCR | 71.09 g/L | Bioreactor | |||
| · Overexpression of IDP1 and POS5 for improved NADPH pool in mitochondria | ||||||||
| Itaconic acid | A. niger | ICDA, ACOA | Dual | · Itaconic acid pathway targeted in mitochondria (AtCadA and AnAcoA) | 1.4 g/L | Flask | ||
| · Expression of itaconic acid pathway in cytosol (AtCadA and EcAcnA) | ||||||||
| Acetoin | C. glabrata | COX4 | Single | · Acetoin pathway targeted in mitochondria (BsALS and BaALDC) | 3.26 g/L | Flask | ||
| · Overexpression of mitochondrial pyruvate carrier (ScMPC) | ||||||||
| Biofuel | Isobutanol | S. cerevisiae | COX4 | Single | · Mitochondrial targeting of ARO10, LlKiVD and AdhARE1, ADH7 | 635 mg/L | Tube | |
| · Overexpression of mitochondrially native genes ILV2, ILV5, ILV3 | ||||||||
| Single | · Mitochondrial targeting of ARO10, LlAdhARE1 | 1.245 g/L | Plate | |||||
| · Deletion of BAT1 and ILV6 | ||||||||
| Single | · Mitochondrial targeting of ARO10, LlAdhARE1 | 8.49 g/L | Bioreactor | |||||
| · Deletion of BAT1 | ||||||||
| 2-Methyl-1-butanol | Single | · Optogenetic regulation of ILV2 and PDC1 | 2.38 g/L | Bioreactor | ||||
| Isopentanol | COX4, CD9, COX6 | Single | · Mitochondrial targeting of LEU4 mutant, LEU1, LEU2 | 1.24 g/L | Plate | |||
| · Deletion of LEU4, LEU9, BAT1 and OAC1 | ||||||||
| n-Butanol | COX4, CYB2, CAT2 | Single | · Mitochondrial targeting of LEU4, LEU1, LEU2 and citramalate synthase (LiCim) | 1.05 g/L | Bioreactor | |||
| · Overexpression of LEU9, cysteine desulfurase NFS1, ARO10 and ADH7 | ||||||||
| Fatty acid alkyl ester | COX4 | Single | · Mitochondrial targeting of ARO10 and ADH7 for improved isobutanol production | 230 mg/L | Flask | |||
| · Overexpression of two wax ester synthase Ms Ws2 and Maqu_0168 | ||||||||
| · Deletion of OPI1 and RPD3 | ||||||||
| Isobutyrate | COX4 | Dual | · Mitochondrial targeting of ARO10 and ADH7, and ATF1 | 260.2 mg/L | Tube | |||
| 3-Methyl-1-butyl acetate | · ATF1 overexpression in cytosol | 296.1 mg/L | ||||||
| 2-Methyl-1-butyl acetate | · Overexpression of PYC2, MDH2, MAE1 | 289.6 mg/L | ||||||
| Product | Yeast species | PTS type | Single/Dual pathway |
Engineering description | Titer | Scale | References | |
|---|---|---|---|---|---|---|---|---|
| Biochemical | Geraniol | S. cerevisiae | SKL | Single | · Compartmentalization of AgGPPS2 and ObGES in peroxisome | 2.75 mg/L | Plate | |
| · Deletion of PEX30 and PEX32 for increased peroxisome number | ||||||||
| · Expression of truncated α-arrestin-like adaptor Bul1 for high tolerance to geraniol | ||||||||
| · The entire geraniol pathway targeted in peroxisome (using ERG20N127Wand ObGerS) with episomal plasmids | 5.52 g/L | Bioreactor | ||||||
| Dual | · Machine learning (ML)-aided identification of the MVA pathway’s critical gene | 120 mg/L | Flask | |||||
| · The MVA pathway targeted in peroxisome | ||||||||
| · The dual MVA platform strain (diploid) created by mating peroxisomal and cytosolic strains | ||||||||
| · Overexpression of a fusion tObGES-ERG20WW | ||||||||
| ePTS1 | Single | · The entire geraniol pathway targeted in peroxisome (using ERG20F96W, N127W and tVoGES) | 9.5 g/L | Flask | ||||
| · ML-guided screen of peroxisome complexes for improved peroxisome capacity | ||||||||
| P. pastoris | ePTS1, PTS of TAL1-2 | Single | · HTS-based screen of 25 putative PTS, followed by directed evolution of selected 3 PTS1 with high targeting efficiency | 20 mg/L | Plate | |||
| · Peroxisomal compartmentalization of geraniol pathway (only tCrGES tagged with newly identified PTS; the remaining proteins tagged with ePTS1). | ||||||||
| R-Limonene | R. toruloides | SKL, PTS2 (C4N11 of 3-KT) | Dual | · Peroxisomal targeting of the entire limonene pathway (two copies of NPPS::LS) | 1.05 g/L | Flask | ||
| · Peroxisomal targeting of Nph17 and overexpressing Nph17 and ACCT for improved acetoacetyl-CoA in peroxisome and cytosol, respectively | ||||||||
| · Cytosolic overexpression of NPPS::LS | ||||||||
| · Overexpression of ACL for improved cytosolic acetyl-CoA | ||||||||
| S. cerevisiae | SKL | Single | · Peroxisomal targeting of the entire biosynthetic pathway targeted with episomal plasmids. | 2.58 g/L | Bioreactor | |||
| · R-limonene: ClLimS | ||||||||
| α-Pinene | · α-Pinene: PtPinS | 69.22 mg/L | Flask | |||||
| Sabinene | · Sabinene: SpSabS | 32.32 mg/L | ||||||
| Camphene | · Camphene: 1SeCamS | 5.77 mg/L | ||||||
| 8-Hydroxygeraniol | · 8-Hydroxygeraniol: ObGerS, CrG8OH, CrCPR | 25.11 mg/L | ||||||
| Canabigerolic acid (CBGA) | · CBGA: CsPT4 | 0.82 mg./L | ||||||
| Linalool | ePTS1 | Dual | · Improving catalytic efficiency toward GPP of linalool synthase through site-directed mutagenesis | 2.6 g/L | Bioreactor | |||
| · Complete linalool pathway targeting to peroxisome | ||||||||
| · Overexpression of tHMG1, IDI1, ERG20F96W, N127W, and McLIS with repression of native ERG20 for cytosolic linalool pathway | ||||||||
| · Mating peroxisomal and cytosolic strain to form the dual strain (diploid) | ||||||||
| β-Myrcene | SKL | Dual | · Source screening of β-myrcene synthase, with truncated QiMS as the best candidate | 142.64 mg/L | Bioreactor | |||
| · Peroxisome localization of fusion enzyme tQiMSD436N-ERG20WW | ||||||||
| · Overexpression of tHMG1 and IDI in the cytosolic MVA pathway and replacement of native ERG20 by ERG20W | ||||||||
| · Enhancing cytosolic acetyl-CoA by overexpressing ADH2 and ALD6 | ||||||||
| α-Humulene | S. cerevisiae | ePTS1 | Dual | · Compartmentalization of entire α-humulene pathway in peroxisome | 1.73 g/L | Bioreactor | ||
| · Overexpression of tHMG1 and ERG20 in cytosol | ||||||||
| · Down regulation of ERG9 by replacing its native promoter by HXT1p and adding PEST sequence | ||||||||
| · Additional overexpression of cytosolic ZzZSS1 in rDNA sites | ||||||||
| SKL | Dual | · ML-guided dual MVA platform strain (diploid) created by mating peroxisomal and cytosolic strains | ~22.5 mg/L | Flask | ||||
| · Overexpression of cytosolic ERG20 and ZzZSS1 | ||||||||
| oPTS1* | Dual | · Develop an orthogonal transport system ScPEX5*-oPTS1* | 17.33 g/L | Bioreactor | ||||
| · rDNA integration of the entire cytosolic and peroxisomal α-humulene pathway (ZSS1 as humulene synthase) | ||||||||
| · Overexpression of ANT1, IDP2, IDP3 (rDNA integration) for improved ATP and NADPH pool in peroxisome | ||||||||
| · Down regulation of ERG9 by replacing its native promoter by HXT1p and adding PEST sequence | ||||||||
| Y. lipolytica | PTS (GGGSSKL) | Single | · Localization of entire α-humulene pathway in peroxisome (NADH-HMG1 as HMG-CoA reductase; AcHS2 as humulene synthase) | 3.2 g/L | Bioreactor | |||
| · Increasing ATP supply in peroxisome by overexpression of ANT1 | ||||||||
| · Adjusting the copy numbers of rate-limiting enzymes (one more copy of peroxisomal targeting NADH-HMG1 and AcHS2) | ||||||||
| · Upregulation of β-oxidation by overexpressing POT1 | ||||||||
| · Oleic acid as a possible C source for α-humulene accumulation | ||||||||
| PTS (GGGSSKL) | Dual | · Iterative integration of the entire cytosolic and peroxisomal α-humulene pathway (AcHS2 as humulene synthase) | 21.7 g/L | Bioreactor | ||||
| · Mediating copy number of cytosolic tHMG1 and AcHS2 | ||||||||
| · Improving cytosolic acetyl-CoA by overexpression of CoA‐acetylating aldehyde dehydrogenase (EcAAD) | ||||||||
| · Repression of ERG9 expression using Cu2+ repressible promoter | ||||||||
| Candida tropicalis | SKL | Single | · Multiple overexpression of the entire peroxisomal α-humulene pathway (double copies of the pathway) | 2.42 mg/L | Flask | |||
| α-Bisabolene | P. pastoris | SKL and LARF | Single | · Peroxisome isolation of the entire α-bisabolene pathway with fusion enzyme of FPPS-AgBIS; upregulation of acetyl-CoA-to-mevalonate pathway; and additional copy of separated FPPS and AgBIS | 1.1 g/L | Flask | ||
| α-Farnesene | ePTS1 | Dual | · Overexpression of tHMG1, IDI1, ERG20 and MdAFS for cytosolic α-farnesene pathway | 2.56 g/L | Flask | |||
| · Improving cytosolic acetyl-CoA pool by introducing pyruvate dehydrogenase (Ec cytoPDH) and ATP-dependent citrate lyase (YlACL) | ||||||||
| · Introduction of IUP pathway and downstream genes (2 copies of IDI, ERG20, MdAFS) into peroxisome | ||||||||
| β-Amyrin | S. cerevisiae | ePTS1 | Dual | · Peroxisome compartmentalization of squalene pathway | 2.6 g/L | Bioreactor | ||
| · Double copies of downstream β-amyrin pathway (ERG1 and GgbAS1) in cytosol | ||||||||
| · Deregulation of ERG7 | ||||||||
| Squalene | ePTS1 | Dual | · The entire squalene pathway targeted in peroxisome with an additional copy of tHMG1 and IDI1 | 11 g/L | Bioreactor | |||
| · Deletion of GPD1 and GPD2 | ||||||||
| · Overexpression of ANT1, IDP2 and IDP3 for improved ATP and NADPH pool in peroxisome | ||||||||
| · ACS1, YlACL1 and YlACL2 targeted to peroxisome for improved peroxisomal acetyl-CoA | ||||||||
| · Diploid strain formation by mating two peroxisomal and cytosolic strains | ||||||||
| SKL | Dual | · The dual MVA platform strain (diploid) created by mating peroxisomal and cytosolic MVA strains | 300 mg/L | Flask | ||||
| · Overexpression of ERG9 and ERG20 | ||||||||
| · Prevention of squalene degradation by supplementation of terbinafine | ||||||||
| Y. lipolytica | SKL | Dual | · The complete squalene pathway targeted in peroxisome | 32.8 g/L (on glucose)/ 31.6 g/L (on acetate) | Bioreactor | |||
| · Enhanced TAG-derived free fatty acids in peroxisome through overexpressing TlTGL, DGA1 and ACC1 | ||||||||
| · Improved cytosolic acetyl-CoA pool by overexpressing PYC1, YHM2 and MmACL | ||||||||
| · Upregulated β-oxidation by overexpressing POX2, MFE1 and POT1 | ||||||||
| · Increased size/number of peroxisomes by overexpressing PEX10 | ||||||||
| · Introduction of Acetyl-CoA shortcut in the peroxisome by targeting 2SeACS1L641P. | ||||||||
| Lycopene | P. pastoris | SKL | Single | · Peroxisomal compartmentalization of lycopene biosynthetic pathway (CrtE, CrtB, CrtI) | 73.9 mg/L | Bioreactor | ||
| Astaxanthin | Y. lipolytica | SKL | Single | · Peroxisomal targeting of the fusion enzyme PsCtrW-HpCrtZ | 58.7 mg/L | Flask | ||
| Protopanaxadiol | S. cerevisiae | PTS1, PTS2 | Single | · Regulating number and size of peroxisome through deletion of PEX11 and ATG36, and overexpression of PEX34 | 4.1 mg/L | Flask | ||
| · Peroxisomal compartmentalization of protopanaxadiol pathway (including tHMG1, ERG9, PgErg1, PgDs, PgPpds, and PgCpr) | ||||||||
| (S)-Norcoclaurine | ePTS1 | Single | · Targeting norcoclaurine synthase (NCS) in peroxisome | ~7 mg/L | Plate | |||
| · Controlling peroxisome biogenesis by engineering transcriptional factors ADR1, OAF1 and PIP2 | ||||||||
| Ophiobolins U | PTS of PEX15 | N/A | · Peroxisomal surface display of CAT2 for improved cytosolic acetyl-CoA | 128.9 mg/L | Flask | |||
| Indole-3-acetic acid | K. marxianus | PTS of PEX15 | Single | · Peroxisomal surface display of tryptophan-2-monooxygenase PsIaaM and indole-acetamide hydrolase PsIaaH | ~61.25 mg/L | Tube | ||
| · Controlling peroxisome number and size by overexpression of PEX19 | ||||||||
| Triacetic acid lactone (TAL) | PTS of PEX15 | Single | · Peroxisomal surface display of CAT2, ACC1 and Gh2-PS | ~ 1.1 g/L | Tube | |||
| · Controlling peroxisome number and size by overexpression of PEX11 | ||||||||
| S. cerevisiae | PTS of PEX15 | Single | · Peroxisomal surface display of CAT2, ACC1S1157A and Gh2-PS | 0.77 g/L | Tube | |||
| pPTS (PTS of Psc60) | Dual | · Cytosolic expression and peroxisomal targeting of Gh2-PS and ACC1S686A, S659A, S1157A | 0.14 g/L | Plate | ||||
| · Increasing peroxisome proliferation by overexpression of ADR1, OAF1 and PIP2 | ||||||||
| Flaviolin | ePTS1 | Single | · Peroxisomal targeting of Sac_RppA and ACC1S686A, S659A, S1157A | N/A | N/A | |||
| Desmethylxanthohumol (DMX) | N/A | Single | · Compartmentalization of the complete MVA pathway in peroxisome | ~62.5 µg/L | Flask | |||
| · Peroxisomal targeting of ACC1 and naringenin chalcone and DMX biosynthetic genes | ||||||||
| Amino acid (Lysine, Histidine) | S. japonicus | PTS1 and PTS2 | N/A | · Peroxisomal targeting of GPD2, LYS3 and HIS2 enzymes | N/A | N/A | ||
| · Regulating peroxisome size with PEX5 mutant and PEX11 | ||||||||
| Penicillin | Hansenula polymorpha | SKL | Single | · Peroxisomal targeting of penicillin pathway’s bottom genes, including PcIAT and PcPCL | 1 mg/L | Plate | ||
| PHA | S. cerevisiae | PTS of BnICL | Single | · Peroxisomal targeting of PaPHAC1 synthase | 0.45 g/g DCW | Flask | ||
| Biofuel | Fatty acid ethyl esters (FAEEs) | Y. lipolytica | SKL | Single | · Peroxisomal targeting of wax ester synthase (AbAtfA) | 110.9 mg/L | Flask | |
| Fatty alcohols | S. cerevisiae | PTS2 | Single | · Peroxisomal targeting of fatty acyl-CoA reductase (TaFAR) | 1.3 g/L | Tube | ||
| · Increasing peroxisomal importing rate by overexpression of PEX7 | ||||||||
| · Improved malonyl-CoA pool by overexpression of ACC1 | ||||||||
| PTS2 | Single | · Peroxisomal targeting of fatty acyl-CoA reductase MaFAR in fatty acyl-CoA overproducing strain | 193 mg/L | Flask | ||||
| Alkanes | PTS1 and PTS2 | Single | · Peroxisomal targeting of free fatty acid (FFA) pathway including MmCAR, AnNPGA, 3SeADO, ferredoxin 3SeFd and ferredoxin reductase 3SeFNR | ~ 3.5 mg/L | Flask | |||
| · Increasing fatty acid production by deleting HFD1 and POX1 | ||||||||
| · Increasing peroxisome population by deletion of PEX31, 32 and overexpression of PEX34 | ||||||||
| Olefins | PTS2 | · Peroxisomal targeting of OleT with flavodoxin EcFldA and flavodoxin reductase EcFNR | ~ 0.20 mg/L | Flask | ||||
| α-Alkenes | P. pastoris | PTS1 and PTS2 | Single | · Peroxisomal targeting of decarboxylase PfUndB together with cofactor protein putidaredoxin–putidaredoxin reductase PpPdr/Pdx | 1.6 mg/L | Flask | ||
| Product | Yeast species | ER engineering | Titer | Scale | References | |
|---|---|---|---|---|---|---|
| Biochemical | Opiate (morphine) | S. cerevisiae | · ER targeting of NADPH-dependent aldo-keto reductase PsCOR1.3 with ER signal peptide from CNE1 | 3.1 mg/L | Plate | |
| Ceramide NS | S. cerevisiae | · ER targeting of human sphingolipid desaturase (hDES1) with ER retention signal KKEK from SUR2 | N/A | N/A | ||
| Trans-nootkatol | S. cerevisiae | · Regulating ER membrane proliferation by overexpression of ICE2 | 30 mg/L | Flask | ||
| 1-Hydroxybufuralol | P. pastoris | N/A | N/A | |||
| β-Amyrin | S. cerevisiae | · Expansion of ER by deleting the phosphatidic acid phosphatase PAH1 | N/A | N/A | ||
| Medicagenic acid | 27.1 mg/L | Flask | ||||
| Oleanane-type sapogenin | N/A | N/A | ||||
| Squalene | S. cerevisiae | · Expansion of ER by overexpression of transcriptional factor INO2 | 634 mg/L | Flask | ||
| Protopanaxadiol | S. cerevisiae | · Controlling gene dosage of tHMG1 (3 copies) for squalene production | 12.1 mg/L | Flask | ||
| Astaxanthin | Y. lipolytica | · ER targeting of the fusion enzyme β-carotene ketolase-hydroxylase (PsCtrW-HpCrtZ) with ER signal peptide KDEL | 53.2 mg/L | Flask | ||
| Human antibody (IgG) | S. cerevisiae | · Expansion of ER by deleting lipid-regulator OPI1 | 126 µg/L | Plate | ||
| · Overexpression of peptidyl-prolyl isomerase CPR5 for enhanced secretion | ||||||
| · Modification of ER morphology and size by deleting lipid-regulator gene OPI1 and ER membrane curvature genes RTN1, RTN2 and YOP1 | 79 ng/mL | Plate | ||||
| Ovalbumin | S. cerevisiae | · Expression of OVA using signal peptide from INU1 | 116.3 mg/L | Bioreactor | ||
| · Co-expression of chaperone Kar2 and disulfide isomerase PDI for improved folding environment in ER | ||||||
| · Expansion of ER by deleting OPI1 and overexpressing transcriptional factors INO2 and INO4 | ||||||
| S-Scoulerine | S. cerevisiae | · ER targeting of berberine bridge enzyme (CyBBE) with ER C-terminal HDEL signal peptide (CyBBE_ERTS) | 113.1 mg/L | Flask | ||
| · ER targeting of mammalian peroxiredoxin IV (PRDX4) with N- and C-terminal ERTS (α- mPRDX4_ERTS) for reducing toxicity of hydrogen peroxide | ||||||
| · Expansion of ER by deleting transcriptional regulator OPI1 of phospholipid biosynthesis | ||||||
| Biofuel | Fatty acid ethyl esters (FAEEs) | Y. lipolytica | · ER targeting of wax-ester synthase (AbAtfA) with ER peptide signal KDEL | 136.5 mg/L | Flask | |
| Fatty alkane | · ER targeting of fatty acyl-CoA reductase (AbACR1)-aldehyde deformylating oxygenase (PmADO) gene cluster with ER signal peptide KDEL | 16.8 mg/L | Flask | |||
| Fatty alcohol | · ER targeting of fatty acyl-CoA reductase (AbACR1) and aldehyde reductase (EcAHR) with ER signal peptide KDEL | 49.2 mg/L | Flask | |||
The single pathway refers to the entire biosynthetic pathway or partial downstream pathway, compartmentalized either in the cytosol or in a subcellular organelle; whereas the dual pathway refers to the entire biosynthetic pathway that is expressed in both the cytosol and a subcellular organelle. Gg:
The single pathway refers to the entire biosynthetic pathway or partial downstream pathway, compartmentalized either in the cytosol or in a subcellular organelle; whereas the dual pathway refers to the entire biosynthetic pathway that is expressed in both the cytosol and a subcellular organelle. Yl:
Ps:
Table 1.
Table 2.
Table 3.
TOP
 MSK
MSK
 ePub Link
ePub Link Cite this Article
Cite this Article






