- Skip Navigation
- Skip to contents
- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Volume 63(3); 2025 > Review
-
Review
Recent advances in the Design-Build-Test-Learn (DBTL) cycle for systems metabolic engineering of Corynebacterium glutamicum - Subeen Jeon†, Yu Jung Sohn†, Haeyoung Lee, Ji Young Park, Dojin Kim, Eun Seo Lee, Si Jae Park*
-
Journal of Microbiology 2025;63(3):e2501021.
DOI: https://doi.org/10.71150/jm.2501021
Published online: March 28, 2025
Department of Chemical Engineering and Materials Science, Graduate Program in System Health Science and Engineering, Ewha Womans University, Seoul 03760, Republic of Korea
- *Correspondence Si Jae Park parksj93@ewha.ac.kr
- †These authors contributed equally to this work.
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
ABSTRACT
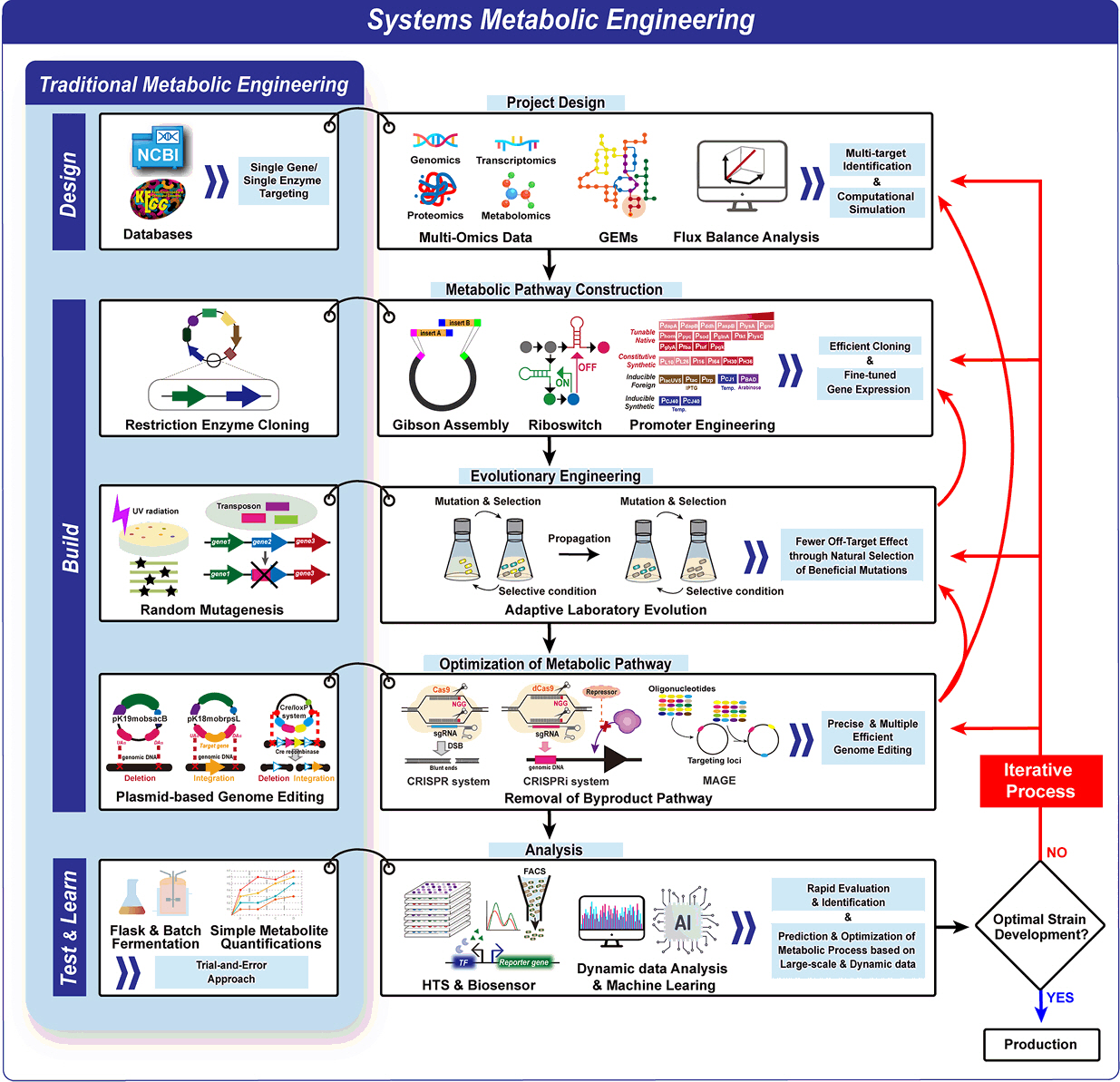
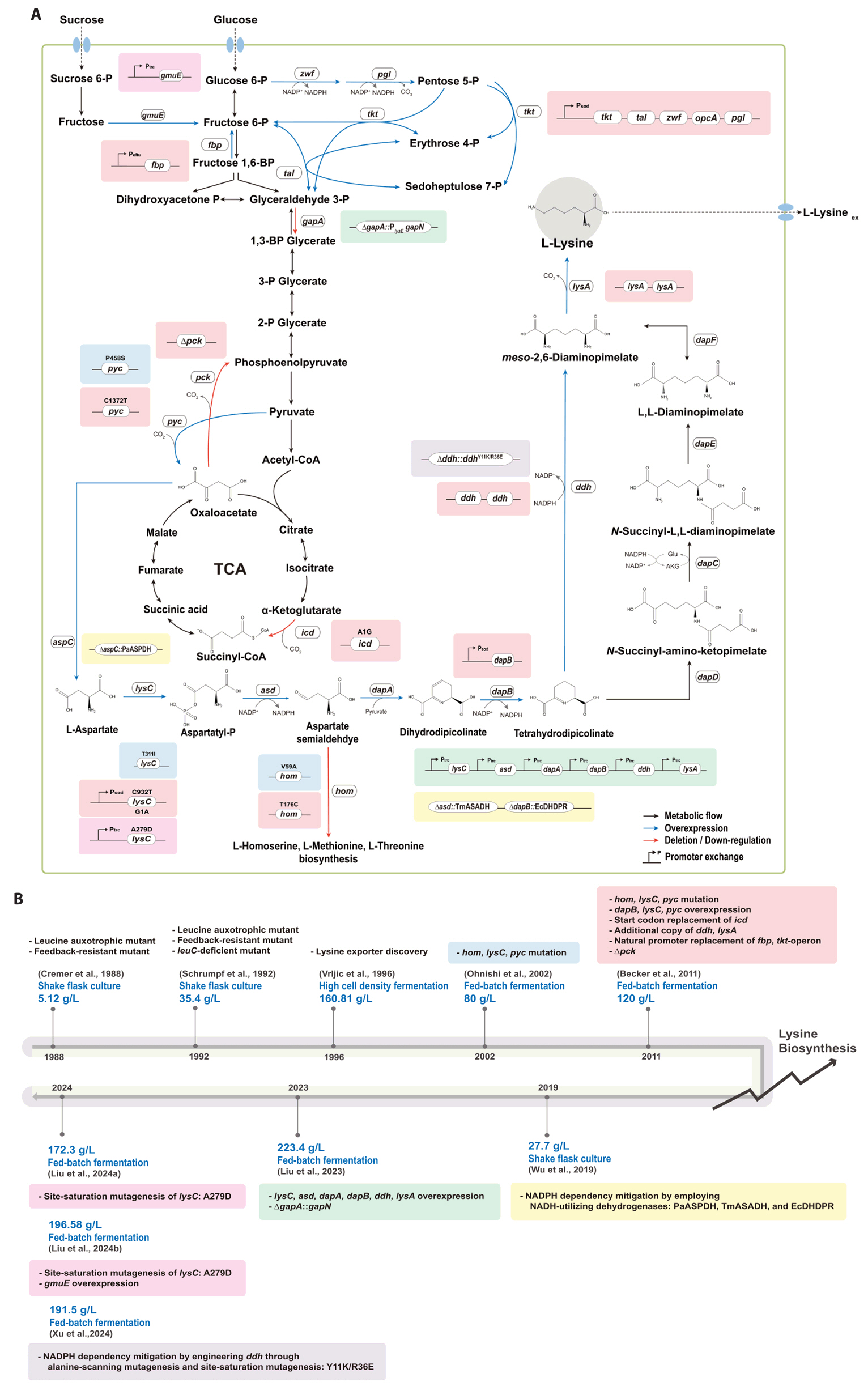
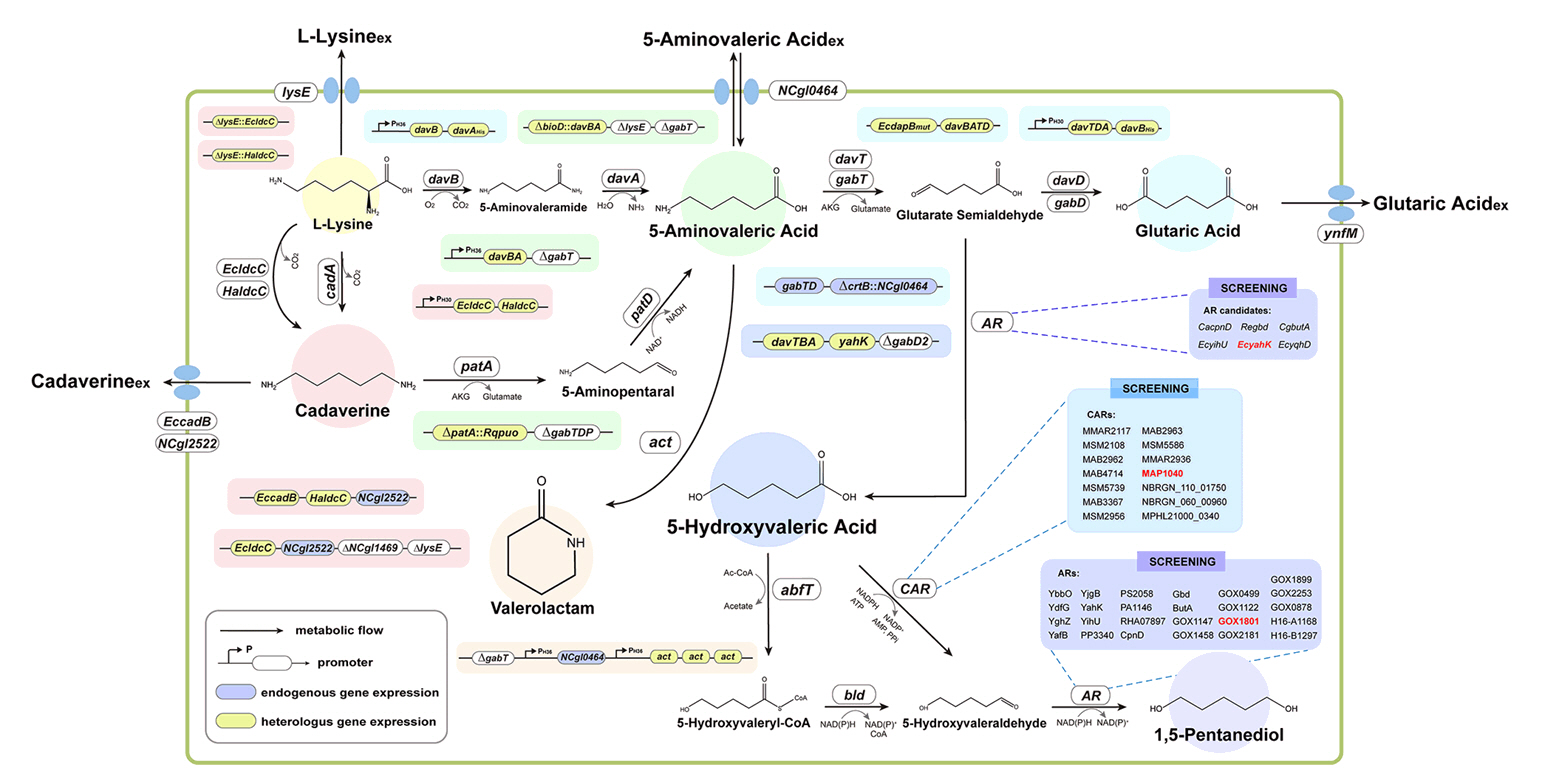
- Existing microbial engineering strategies—encompassing metabolic engineering, systems biology, and systems metabolic engineering—have significantly enhanced the potential of microbial cell factories as sustainable alternatives to the petrochemical industry by optimizing metabolic pathways. Recently, systems metabolic engineering, which integrates tools from synthetic biology, enzyme engineering, omics technology, and evolutionary engineering, has been successfully developed. By leveraging modern engineering strategies within the Design-Build-Test-Learn (DBTL) cycle framework, these advancements have revolutionized the biosynthesis of valuable compounds. This review highlights recent progress in the metabolic engineering of Corynebacterium glutamicum, a versatile microbial platform, achieved through various approaches from traditional metabolic engineering to advanced systems metabolic engineering, all within the DBTL cycle. A particular focus is placed C5 platform chemicals derived from L-lysine, one of the key amino acid production pathways of C. glutamicum. The development of DBTL cycle-based metabolic engineering strategies for this process is discussed.
- Keywords: Design-Build-Test-Learn (DBTL) cycle, metabolic engineering, systems metabolic engineering, Corynebacterium glutamicum, L-lysine, C5 chemicals
Introduction
From Traditional to Advanced Metabolic Engineering: Tools and Strategies for C. glutamicum Optimization Through the DBTL Cycle
Broadening the Metabolic Potential of C. glutamicum: From L-lysine production to Specialty Chemical Biosynthesis
Conclusion
Acknowledgments
This work was supported by the Development of next-generation biorefinery platform technologies for leading bio-based chemicals industry project (2022M3J5A1056072) and the Development of platform technologies of microbial cell factories for the next-generation biorefineries project (2022M3J5A1056117) from National Research Foundation (NRF) funded by the Korean government (MSIT), the Development of synthetic microbial platform systems for one step-one pot synthesis of next-generation biodegradable biopolymers (NRF-2022M3J4A1053696) from NRF supported by MSIT and by R&D Program of MOTIE/KEIT (RS-2024-00467186).
| Product | Host strain | Engineering technique | Titer (g/L) | Scale | References |
|---|---|---|---|---|---|
| Cadaverine | C. glutamicum PKC | √ Chromosomal integration of E. coli derived ldcC with a strong synthetic H30 promoter at the lysE site | 103.78 | Fed-batch | Kim et al. (2018) |
| C. glutamicum PKC | √ Chromosomal integration of H. alvei derived ldcC with a strong synthetic H30 promoter at the lysE site | 125 | Fed-batch | Kim et al. (2019a) | |
| C. glutamicum KCTC 1857 | √ Co-expression of dr1558 and cadA | 10.3 | Fed-batch | Kang and Choi (2021) | |
| C. glutamicum KCTC 1857 | √ Co-expression of dr1558 and ldcC | 25.1 | Fed-batch | Kang and Choi (2022) | |
| GTA | C. glutamicum KCTC 1857 | √ Introduction of glutarate biosynthesis pathway by expressing davTDBA genes | 24.5 | Fed-batch | Kim et al. (2018) |
| √ Gene modification of davB with an N-terminal His6-tag | |||||
| C. glutamicum BE (C. glutamicum KCTC 12390BP) | √ Identification and expression of 11 target genes for increasing L-lysine supply through gene deletion/integration/substitution along with system-wide analyses | 105.3 | Fed-batch | Han et al. (2020) | |
| √ Overexpression of ynfM | |||||
| C. glutamicum GRLys1 | √ Introduction of glutarate biosynthesis pathway by expressing ldcC, patDA, gabTDStu | 25 | Fed-batch | Pérez-García et al. (2018) | |
| √ Gene deletion of sugR, ldhA, snaA, cgmA, and gdh | |||||
| C. glutamicum GSLA2 Δgdh | √ Introduction of glutarate biosynthesis pathway by expressing gltBE686Q, ldcC, patDA, tetA(Z)Δ21bp-gabTDP134L | 22.7 | Fed-batch | Prell et al. (2021) | |
| √ Adaptive laboratory evolution | |||||
| 5-AVA | C. glutamicum BE | √ Introduction of 5-AVA biosynthesis pathway by expressing P. putida derived davB and davA | 33.1 | Fed-batch | Shin et al. (2016) |
| √ Overexpression of davA by fusing it with His6-Tag at its N-Terminal | |||||
| C. glutamicum GRLys1 | √ Introduction of 5-AVA biosynthesis pathway by expressing E. coil derived ldcC, patA and patD | 5.1 | Shake-flask | Jorge et al. (2017) | |
| √ Gene deletion of sugR, ldhA, snaA, cgmA, and gabTDP | |||||
| 5-HV | C. glutamicum PKC | √ Introduction of 5-HV biosynthesis pathway by expressing P. putida derived davTBA and E. coil derived yahK | 52.1 | Fed-batch | Sohn et al. (2021) |
| √ Gene deletion of gabD | |||||
| 1,5-PDO | C. glutamicum PKC ΔgabD2 | √ Introduction of 1,5-PDO biosynthesis pathway by expressing M. marinum derived carboxylic acid reductase (CAR) and G. oxydans derived GOX1801 | 43.4 | Fed-batch | Sohn et al. (2024) |
| √ Chromosomal integration of PH30DavBHisA expression cassette at the site of lysE | |||||
| √ Enzyme engineering of CAR | |||||
| VL | C. glutamicum XT1 | √ Introduction of valerolactam biosynthesis pathway by expressing P. putida derived davBA and C. propionicum derived act | 12.33 | Fed-batch | Zhao et al. (2023) |
| √ Dynamic upregulation system using engineered ChnR-B1/Pb-E1 biosensor system | |||||
| C. glutamicum GA16 ΔgabT | √ Gene expression down regulation of gdh using sRNA knock-down system | 76.1 | Fed-batch | Han and Lee (2023) | |
| √ Identification and engineering of 5-AVA transporter genes | |||||
| √ Chromosomal integration of multiple copies of act |
- Adkins J, Jordan J, Nielsen DR. 2013. Engineering Escherichia coli for renewable production of the 5‐carbon polyamide building‐blocks 5‐aminovalerate and glutarate. Biotechnol Bioeng. 110(6): 1726–1734. ArticlePubMed
- Becker J, Zelder O, Hafner S, Schroder H, Wittmann C. 2011. From zero to hero--design-based systems metabolic engineering of Corynebacterium glutamicum for L-lysine production. Metab Eng. 13(2): 159–168. ArticlePubMed
- Bendt AK, Burkovski A, Schaffer S, Bott M, Farwick M, et al. 2003. Towards a phosphoproteome map of Corynebacterium glutamicum. Proteomics. 3(8): 1637–1646. ArticlePubMed
- Binder S, Siedler S, Marienhagen J, Bott M, Eggeling L, et al. 2013. Recombineering in Corynebacterium glutamicum combined with optical nanosensors: a general strategy for fast producer strain generation. Nucleic Acids Res. 41(12): 6360–6369.ArticlePubMedPMC
- Chae TU, Ko YS, Hwang KS, Lee SY. 2017. Metabolic engineering of Escherichia coli for the production of four-, five- and six-carbon lactams. Metab Eng. 41: 82–91. ArticlePubMed
- Chai M, Deng C, Chen Q, Lu W, Liu Y, et al. 2021. Synthetic biology toolkits and metabolic engineering applied in Corynebacterium glutamicum for biomanufacturing. ACS Synth Biol. 10(12): 3237–3250. ArticlePubMed
- Chen W, Zhang Y, Yeo WS, Bae T, Ji Q. 2017. Rapid and efficient genome editing in staphylococcus aureus by using an engineered CRISPR/cas9 system. J Am Chem Soc. 139(10): 3790–3795. ArticlePubMed
- Chen Z, Bommareddy RR, Frank D, Rappert S, Zeng AP. 2014. Deregulation of feedback inhibition of phosphoenolpyruvate carboxylase for improved lysine production in Corynebacterium glutamicum. Appl Environ Microbiol. 80(4): 1388–1393. ArticlePubMedPMCLink
- Cheng J, Chen P, Song A, Wang D, Wang Q. 2018. Expanding lysine industry: Industrial biomanufacturing of lysine and its derivatives. J Ind Microbiol Biotechnol. 45(8): 719–734. ArticlePubMedPDF
- Cho JS, Yang D, Prabowo CPS, Ghiffary MR, Han T, et al. 2023. Targeted and high-throughput gene knockdown in diverse bacteria using synthetic sRNAs. Nat Commun. 14: 2359.ArticlePubMedPMCPDF
- Choi KR, Jang WD, Yang D, Cho JS, Park D, et al. 2019. Systems metabolic engineering strategies: Integrating systems and synthetic biology with metabolic engineering. Trends Biotechnol. 37(8): 817–837. ArticlePubMed
- Chuang HY, Hofree M, Ideker T. 2010. A decade of systems biology. Annu Rev Cell Dev Biol. 26: 721–744. ArticlePubMedPMC
- Cleto S, Jensen JV, Wendisch VF, Lu TK. 2016. Corynebacterium glutamicum metabolic engineering with CRISPR interference (CRISPRi). ACS Synth Biol. 5(5): 375–385. ArticlePubMedPMC
- Cremer J, Treptow C, Eggeling L, Sahm H. 1988. Regulation of enzymes of lysine biosynthesis in Corynebacterium glutamicum. J Gen Microbiol. 134(12): 3221–3229. ArticlePubMed
- Demain AL. 2000. Microbial biotechnology. Trends Biotechnol. 18(1): 26–31.ArticlePubMed
- Desai SC, Macrin AD, Senthilvelan T, Panda RC. 2024. Identification of genes associated with accelerated biological ageing through computational analysis: a systematic review. Biotechnol Bioprocess Eng. 29(4): 636–649. ArticlePDF
- Dragosits M, Mattanovich D. 2013. Adaptive laboratory evolution -- principles and applications for biotechnology. Microb Cell Fact. 12: 64.ArticlePubMedPMC
- Eggeling L, Bott M. 2015. A giant market and a powerful metabolism: L-lysine provided by Corynebacterium glutamicum. Appl Microbiol Biotechnol. 99(8): 3387–3394. ArticlePubMedPDF
- Eikmanns BJ, Eggeling L, Sahm H. 1993. Molecular aspects of lysine, threonine, and isoleucine biosynthesis in Corynebacterium glutamicum. Antonie van Leeuwenhoek. 64(2): 145–163. ArticlePubMedPDF
- Esvelt KM, Wang HH. 2013. Genome‐scale engineering for systems and synthetic biology. Mol Syst Biol. 9(1): 641.ArticlePubMedPMCLink
- Felix F, Letti LAJ, Vinicius de Melo Pereira G, Bonfim PGB, Soccol VT, et al. 2019. L-lysine production improvement: A review of the state of the art and patent landscape focusing on strain development and fermentation technologies. Crit Rev Biotechnol. 39(8): 1031–1055. ArticlePubMed
- Gilbert LA, Larson MH, Morsut L, Liu Z, Brar GA, et al. 2013. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell. 154(2): 442–451. ArticlePubMedPMC
- Godara A, Kao KC. 2020. Adaptive laboratory evolution for growth coupled microbial production. World J Microbiol Biotechnol. 36(11): 175.ArticlePubMedPDF
- Gonzalez-Ramos D, Gorter de Vries AR, Grijseels SS, van Berkum MC, Swinnen S, et al. 2016. A new laboratory evolution approach to select for constitutive acetic acid tolerance in saccharomyces cerevisiae and identification of causal mutations. Biotechnol Biofuels. 9: 173.ArticlePubMedPMC
- Gorshkova NV, Lobanova JS, Tokmakova IL, Smirnov SV, Akhverdyan VZ, et al. 2018. Mu-driven transposition of recombinant mini-mu unit DNA in the Corynebacterium glutamicum chromosome. Appl Microbiol Biotechnol. 102(6): 2867–2884. ArticlePubMedPMCPDF
- Gupta A, Lee SG, Sung BH, Lee DH, Cho BK, et al. 2024. Advancing biofoundry development: strategies and challenges. Biotechnol Bioprocess Eng. 29(5): 771–778. ArticlePDF
- Han T, Kim GB, Lee SY. 2020. Glutaric acid production by systems metabolic engineering of an L-lysine-overproducing Corynebacterium glutamicum. Proc Natl Acad Sci USA. 117(48): 30328–30334. ArticlePubMedPMC
- Han T, Lee SY. 2023. Metabolic engineering of Corynebacterium glutamicum for the high-level production of valerolactam, a nylon-5 monomer. Metab Eng. 79: 78–85. ArticlePubMed
- Haupka C, Delépine B, Irla M, Heux S, Wendisch VF. 2020. Flux enforcement for fermentative production of 5-aminovalerate and glutarate by Corynebacterium glutamicum. Catalysts. 10(9): 1065.Article
- Hsu PD, Lander ES, Zhang F. 2014. Development and applications of CRISPR-cas9 for genome engineering. Cell. 157(6): 1262–1278. ArticlePubMedPMC
- Huang H, Chai C, Li N, Rowe P, Minton NP, et al. 2016. CRISPR/cas9-based efficient genome editing in Clostridium ljungdahlii, an autotrophic gas-fermenting bacterium. ACS Synth Biol. 5(12): 1355–1361. ArticlePubMed
- Hwang Y, Noh MH, Jung GY. 2024. Recent advancements in flavonoid production through engineering microbial systems. Biotechnol Bioprocess Eng. 29(5): 792–805. ArticlePDF
- Ikeda M, Nakagawa S. 2003. The Corynebacterium glutamicum genome: features and impacts on biotechnological processes. Appl Microbiol Biotechnol. 62: 99–109.ArticlePubMedPDF
- Inui M, Tsuge Y, Suzuki N, Vertes AA, Yukawa H. 2005. Isolation and characterization of a native composite transposon, tn 14751, carrying 17.4 kilobases of Corynebacterium glutamicum chromosomal DNA. Appl Environ Microbiol. 71(1): 407–416. ArticlePubMedPMCLink
- Jäger W, Schafer A, Puhler A, Labes G, Wohlleben W. 1992. Expression of the Bacillus subtilis sacB gene leads to sucrose sensitivity in the gram-positive bacterium Corynebacterium glutamicum but not in Streptomyces lividans. J Bacteriol. 174(16): 5462–5465. ArticlePubMedPMCLink
- Jorge JM, Pérez-García F, Wendisch VF. 2017. A new metabolic route for the fermentative production of 5-aminovalerate from glucose and alternative carbon sources. Bioresour Technol. 245: 1701–1709. ArticlePubMed
- Jung HJ, Shin Y, Hwang JH, Shin N, Kim HJ, et al. 2024. Establishment of an optimized electroporation method for Halomonas sp. YK44 and its application in the coproduction of PHB and isobutanol. Biotechnol Bioprocess Eng. 29(2): 339–351. ArticlePDF
- Kang S, Choi J. 2021. Enhanced cadaverine production by recombinant Corynebacterium glutamicum with a heterologous dr1558 regulator at low pH condition. Process Biochem. 111: 63–70. Article
- Kang S, Choi J. 2022. Production of cadaverine in recombinant Corynebacterium glutamicum overexpressing lysine decarboxylase (ldcC) and response regulator dr1558. Appl Biochem Biotechnol. 194(2): 1013–1024. ArticlePubMedPDF
- Kiefer P, Heinzle E, Zelder O, Wittmann C. 2004. Comparative metabolic flux analysis of lysine-producing Corynebacterium glutamicum cultured on glucose or fructose. Appl Environ Microbiol. 70(1): 229–239. ArticlePubMedPMCLink
- Kim HT, Baritugo K, Hyun SM, Khang TU, Sohn YJ, et al. 2019a. Development of metabolically engineered Corynebacterium glutamicum for enhanced production of cadaverine and its use for the synthesis of bio-polyamide 510. ACS Sustain Chem Eng. 8(1): 129–138. Article
- Kim HT, Baritugo K, Oh YH, Hyun SM, Khang TU, et al. 2018. Metabolic engineering of Corynebacterium glutamicum for the high-level production of cadaverine that can be used for the synthesis of biopolyamide 510. ACS Sustain Chem Eng. 6(4): 5296–5305. Article
- Kim HT, Khang TU, Baritugo K, Hyun SM, Kang KH, et al. 2019b. Metabolic engineering of Corynebacterium glutamicum for the production of glutaric acid, a C5 dicarboxylic acid platform chemical. Metab Eng. 51: 99–109. Article
- Kim GY, Kim J, Park G, Kim HJ, Yang J, et al. 2023. Synthetic biology tools for engineering Corynebacterium glutamicum. Comput Struct Biotechnol J. 21: 1955–1965. ArticlePubMedPMC
- Kim T, Ko M, Rha E, Kim H, Lee H. 2024. Microfluidics-driven high-throughput phenotyping and screening in synthetic biology: from single cells to cell-free systems. Biotechnol. Bioprocess Eng. 29(1): 25–33. ArticlePDF
- Kind S, Jeong WK, Schroder H. 2010. Systems-wide metabolic pathway engineering in Corynebacterium glutamicum for bio-based production of diaminopentane. Metab Eng. 12(4): 341–351.ArticlePubMed
- Kind S, Neubauer S, Becker J, Yamamoto M, Völkert M, et al. 2014. From zero to hero-production of bio-based nylon from renewable resources using engineered Corynebacterium glutamicum. Metab Eng. 25: 113–123. ArticlePubMed
- Kromer JO, Sorgenfrei O, Klopprogge K, Heinzle E, Wittmann C. 2004. In-depth profiling of lysine-producing Corynebacterium glutamicum by combined analysis of the transcriptome, metabolome, and fluxome. J Bacteriol. 186(6): 1769–1784. ArticlePubMedPMCLink
- Larson MH, Gilbert LA, Wang X, Lim WA, Weissman JS, et al. 2013. CRISPR interference (CRISPRi) for sequence-specific control of gene expression. Nat Protoc. 8: 2180–2196. ArticlePubMedPMCPDF
- Lee SY, Kim HU, Chae TU, Cho JS, Kim JW, et al. 2019. A comprehensive metabolic map for production of bio-based chemicals. Nat Catal. 2: 18–33. ArticlePDF
- Lee JW, Kim TY, Jang YS, Choi S, Lee SY. 2011. Systems metabolic engineering for chemicals and materials. Trends Biotechnol. 29(8): 370–378. ArticlePubMed
- Lee WH, Pathanibul P, Quarterman J, Jo JH, Han NS, et al. 2012. Whole cell biosynthesis of a functional oligosaccharide, 2'-fucosyllactose, using engineered Escherichia coli. Microb Cell Fact. 11: 48.ArticlePubMedPMCPDF
- Lee J, Shin H, Lee KH, Lee H, Lee G, et al. 2024. Component analysis and utilization strategy of brown macroalgae as promising feedstock for sugar platform-based marine biorefinery. Biotechnol. Bioprocess Eng. 29(2): 377–386. ArticlePDF
- Li M, Li D, Huang Y, Liu M, Wang H, et al. 2014. Improving the secretion of cadaverine in Corynebacterium glutamicum by cadaverine–lysine antiporter. J Ind Microbiol Biotechnol. 41(4): 701–709.ArticlePubMedPDF
- Lindner SN, Niederholtmeyer H, Schmitz K, Schoberth SM, Wendisch VF, et al. 2010. Polyphosphate/ATP-dependent NAD kinase of Corynebacterium glutamicum: biochemical properties and impact of ppnK overexpression on lysine production. Appl Microbiol Biotechnol. 87(2): 583–593.ArticlePubMedPDF
- Liu Z, Liu J, Zhang F, Zhang W. 2024a. Modifying Corynebacterium glutamicum by metabolic engineering for efficient synthesis of L-lysine. Syst Microbiol Biomanuf. 5: 288–299. ArticlePDF
- Liu J, Ou Y, Xu JZ, Rao ZM, Zhang WG. 2023. L-lysine production by systems metabolic engineering of an NADPH auto-regulated Corynebacterium glutamicum. Bioresour Technol. 387: 129701.ArticlePubMed
- Liu WL, Wang HY, Li M, Xu GT, Xu JZ. 2024b. The 279th residue of aspartate kinase in Corynebacterium glutamicum is important for relieving the feedback inhibition by L-lysine and L-threonine. Mol Catal. 569: 114652.Article
- Liu J, Xu JZ, Rao ZM, Zhang WG. 2022. Industrial production of L-lysine in Corynebacterium glutamicum: Progress and prospects. Microbiol Res. 262: 127101.ArticlePubMed
- Liu C, Zhang L, Liu H, Cheng K. 2017. Delivery strategies of the CRISPR-cas9 gene-editing system for therapeutic applications. J Control Release. 266: 17–26. ArticlePubMedPMC
- Luntz MG, Zhdanova NI, Bourd GI. 1986. Transport and excretion of L-lysine in Corynebacterium glutamicum. Microbiology. 132(8): 2137–2146. Article
- Mans R, Daran JG, Pronk JT. 2018. Under pressure: Evolutionary engineering of yeast strains for improved performance in fuels and chemicals production. Curr Opin Biotechnol. 50: 47–56. ArticlePubMed
- Mimitsuka T, Sawai H, Hatsu M, Yamada K. 2007. Metabolic engineering of Corynebacterium glutamicum for cadaverine fermentation. Biosci Biotechnol Biochem. 71(9): 2130–2135. ArticlePubMed
- Mormann S, Lömker A, Rückert C, Gaigalat L, Tauch A, et al. 2006. Random mutagenesis in Corynebacterium glutamicum ATCC 13032 using an IS 6100-based transposon vector identified the last unknown gene in the histidine biosynthesis pathway. BMC Genom. 7: 205.ArticlePubMedPMCPDF
- Nakazawa S, Imaichi O, Kogure T, Kubota T, Toyoda K, et al. 2021. History-driven genetic modification design technique using a domain-specific lexical model for the acceleration of DBTL cycles for microbial cell factories. ACS Synth Biol. 10(9): 2308–2317.ArticlePubMedLink
- Naseri G, Koffas MA. 2020. Application of combinatorial optimization strategies in synthetic biology. Nat Commun. 11(1): 2446.ArticlePubMedPMCPDF
- Oh YH, Choi JW, Kim EY, Song BK, Jeong KJ, et al. 2015. Construction of synthetic promoter-based expression cassettes for the production of cadaverine in recombinant Corynebacterium glutamicum. Appl Biochem Biotechnol. 176(7): 2065–2075. ArticlePubMedPDF
- Ohnishi J, Katahira R, Mitsuhashi S, Kakita S, Ikeda M. 2005. A novel gnd mutation leading to increased L-lysine production in Corynebacterium glutamicum. FEMS Microbiol Lett. 242(2): 265–274. ArticlePubMed
- Ohnishi J, Mitsuhashi S, Hayashi M, Ando S, Yokoi H, et al. 2002. A novel methodology employing Corynebacterium glutamicum genome information to generate a new L-lysine-producing mutant. Appl Microbiol Biotechnol. 58(2): 217–223. ArticlePubMedPDF
- Park SJ, Kim EY, Noh W, Park HM, Oh YH, et al. 2013. Metabolic engineering of Escherichia coli for the production of 5-aminovalerate and glutarate as C5 platform chemicals. Metab Eng. 16: 42–47. ArticlePubMed
- Paquet D, Kwart D, Chen A, Sproul A, Jacob S, et al. 2016. Efficient introduction of specific homozygous and heterozygous mutations using CRISPR/cas9. Nature. 533: 125–129. ArticlePubMedPDF
- Peng F, Wang X, Sun Y, Dong G, Yang Y, et al. 2017. Efficient gene editing in Corynebacterium glutamicum using the CRISPR/cas9 system. Microb Cell Fact. 16: 201.ArticlePubMedPMCPDF
- Pérez-García F, Jorge JM, Dreyszas A, Risse JM, Wendisch VF. 2018. Efficient production of the dicarboxylic acid glutarate by Corynebacterium glutamicum via a novel synthetic pathway. Front Microbiol. 9: 2589.ArticlePubMedPMC
- Plachý J, Ulbert S, Pelechová J, Krfmphanzl V. 1985. Fermentation production of L-homoserine by Corynebacterium sp. and its possible use in the preparation of threonine and lysine. Folia Microbiol. 30(6): 485–492. ArticlePDF
- Prell C, Busche T, Ruckert C, Nolte L, Brandenbusch C, et al. 2021. Adaptive laboratory evolution accelerated glutarate production by Corynebacterium glutamicum. Microb Cell Fact. 20: 97.ArticlePubMedPMCPDF
- Quast K, Bathe B, Pühler A, Kalinowski J. 1999. The Corynebacterium glutamicum insertion sequence ISCg2 prefers conserved target sequences located adjacent to genes involved in aspartate and glutamate metabolism. Mol Gen Genet. 262(3): 568–578. ArticlePubMedPDF
- Richardson CD, Ray GJ, DeWitt MA, Curie GL, Corn JE. 2016. Enhancing homology-directed genome editing by catalytically active and inactive CRISPR-cas9 using asymmetric donor DNA. Nat Biotechnol. 34: 339–344. ArticlePubMedPDF
- Rohles CM, Giesselmann G, Kohlstedt M, Wittmann C, Becker J. 2016. Systems metabolic engineering of Corynebacterium glutamicum for the production of the carbon-5 platform chemicals 5-aminovalerate and glutarate. Microb Cell Fact. 15: 154.ArticlePubMedPMCPDF
- Rohles CM, Gläser L, Kohlstedt M, Gießelmann G, Pearson S, et al. 2018. A bio-based route to the carbon-5 chemical glutaric acid and to bionylon-6, 5 using metabolically engineered Corynebacterium glutamicum. Green Chem. 20: 4662–4674. Article
- Rohles C, Pauli S, Gießelmann G, Kohlstedt M, Becker J, et al. 2022. Systems metabolic engineering of Corynebacterium glutamicum eliminates all by-products for selective and high-yield production of the platform chemical 5-aminovalerate. Metab Eng. 73: 168–181. ArticlePubMed
- Sandberg TE, Salazar MJ, Weng LL, Palsson BO, Feist AM, et al. 2019. The emergence of adaptive laboratory evolution as an efficient tool for biological discovery and industrial biotechnology. Metab Eng. 56: 1–16.ArticlePubMedPMC
- Schäfer A, Tauch A, Jager W, Kalinowski J, Thierbach G, et al. 1994. Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: Selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene. 145(1): 69–73. ArticlePubMed
- Schrumpf B, Eggeling L, Sahm H. 1992. Isolation and prominent characteristics of an L-lysine hyperproducing strain of Corynebacterium glutamicum. Appl Environ Microbiol. 37: 566–571. ArticlePDF
- Schrumpf B, Schwarzer A, Kalinowski J, Puhler A, Eggeling L, et al. 1991. A functionally split pathway for lysine synthesis in Corynebacterium glutamicium. J Bacteriol. 173(14): 4510–4516. ArticlePubMedPMCLink
- Shah AH, Hameed A, Ahmad S, Khan GM. 2002. Optimization of culture conditions for L-lysine fermentation by Corynebacterium glutamicum. Biol Sci. 2(3): 151–156.Article
- Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, et al. 2014. Genome-scale CRISPR-cas9 knockout screening in human cells. Science. 343(6166): 84–87. ArticlePubMedPMC
- Shin JH, Park SH, Oh YH, Choi JW, Lee MH, et al. 2016. Metabolic engineering of Corynebacterium glutamicum for enhanced production of 5-aminovaleric acid. Microb Cell Fact. 15: 174.ArticlePubMedPMCPDF
- Sohn YJ, Hwang SY, Lee H, Jeon S, Park JY, et al. 2024. Metabolic engineering of Corynebacterium glutamicum for high-level production of 1,5-pentanediol, a C5 diol platform chemical. Adv Sci. doi.org/10.1002/advs.202412670. Article
- Sohn YJ, Kang M, Baritugo K, Son J, Kang KH, et al. 2021. Fermentative high-level production of 5-hydroxyvaleric acid by metabolically engineered Corynebacterium glutamicum. ACS Sustain Chem Eng. 9(6): 2523–2533. Article
- Sohn YJ, Kang M, Ryu M, Lee S, Kang KH, et al. 2022. Development of a bio-chemical route to C5 plasticizer synthesis using glutaric acid produced by metabolically engineered Corynebacterium glutamicum. Green Chem. 24: 1590–1602. Article
- Son J, Sohn YJ, Baritugo K, Jo SY, Song HM, et al. 2023. Recent advances in microbial production of diamines, aminocarboxylic acids, and diacids as potential platform chemicals and bio-based polyamides monomers. Biotechnol Adv. 62: 108070.ArticlePubMed
- Stella RG, Gertzen CG, Smits SH, Gätgens C, Polen T, et al. 2021. Biosensor-based growth-coupling and spatial separation as an evolution strategy to improve small molecule production of Corynebacterium glutamicum. Metab Eng. 68: 162–173.ArticlePubMed
- Stella RG, Wiechert J, Noack S, Frunzke J. 2019. Evolutionary engineering of Corynebacterium glutamicum. Biotechnol J. 14(9): e1800444. ArticlePubMedLink
- Suzuki N, Okai N, Nonaka H, Tsuge Y, Inui M, et al. 2006. High-throughput transposon mutagenesis of Corynebacterium glutamicum and construction of a single-gene disruptant mutant library. Appl Environ Microbiol. 72(5): 3750–3755. ArticlePubMedPMCLink
- Suzuki N, Tsuge Y, Inui M, Yukawa H. 2005. Cre/loxP-mediated deletion system for large genome rearrangements in Corynebacterium glutamicum. Appl Microbiol Biotechnol. 67(2): 225–233. ArticlePubMedPDF
- Tan Y, Xu D, Li Y, Wang X. 2012. Construction of a novel sacB-based system for marker-free gene deletion in Corynebacterium glutamicum. Plasmid. 67(1): 44–52. ArticlePubMed
- Tauch A, Kassing F, Kalinowski J, Pühler A. 1995. The Corynebacterium xerosis composite transposon Tn5432 consists of two identical insertion sequences, designated IS1249, flanking the erythromycin resistance gene ermCX. Plasmid. 34(2): 119–131. ArticlePubMed
- Tosaka O, Koichi T, Hirose Y. 1978. Production of L-lysine by leucine auxotrophs derived from AEC resistant mutant of Brevibacterium lactofermentum. Agric Biol Chem. 42(6): 1181–1186. Article
- Vertès AA, Inui M, Kobayashi M, Kurusu Y, Yukawa H. 1994. Isolation and characterization of IS31831, a transposable element from Corynebacterium glutamicum. Mol Microbiol. 11(4): 739–746. ArticlePubMed
- von Tiedemann P, Anwar S, Kemmer‐Jonas U, Asadi K, Frey H. 2020. Synthesis and solution processing of nylon‐5 ferroelectric thin films: The renaissance of odd‐nylons? Macromol Chem Phys. 221(5): 1900468.Article
- Vrljic M, Sahm H, Eggeling L. 1996. A new type of transporter with a new type of cellular function: L-lysine export from Corynebacterium glutamicum. Mol Microbiol. 22(5): 815–826. ArticlePubMed
- Wang G, Li Q, Zhang Z, Yin X, Wang B, et al. 2023. Recent progress in adaptive laboratory evolution of industrial microorganisms. J Ind Microbiol Biotechnol. 50(1): kuac023.ArticlePubMedPMCPDF
- Wen J, Sun W, Leng G, Li D, Feng C, et al. 2024. Enhanced fermentative γ-aminobutyric acid production by a metabolic engineered Corynebacterium glutamicum. Biotechnol Bioprocess Eng. 29: 129–140. ArticlePDF
- Wu W, Zhang Y, Liu D, Chen Z. 2019. Efficient mining of natural NADH-utilizing dehydrogenases enables systematic cofactor engineering of lysine synthesis pathway of Corynebacterium glutamicum. Metab Eng. 52: 77–86. ArticlePubMed
- Xu JZ, Liu WL, Zhang WG, Lu C. 2024. Creation of a NADH-dependent meso-diaminopimelate dehydrogenase by site-saturation mutagenesis for producing L-lysine in Corynebacterium glutamicum. Mol Catal. 564: 114351.Article
- Zerbini F, Zanella I, Fraccascia D, Konig E, Irene C, et al. 2017. Large scale validation of an efficient CRISPR/Cas-based multi gene editing protocol in Escherichia coli. Microb Cell Fact. 16: 68.ArticlePubMedPMCPDF
- Zhang Y, Cai J, Shang X, Wang B, Liu S, et al. 2017. A new genome-scale metabolic model of Corynebacterium glutamicum and its application. Biotechnol Biofuels. 10: 169.ArticlePubMedPMCPDF
- Zhao X, Wu Y, Feng T, Shen J, Lu H, et al. 2023. Dynamic upregulation of the rate-limiting enzyme for valerolactam biosynthesis in Corynebacterium glutamicum. Metab Eng. 77: 89–99. ArticlePubMed
- Zhao G, Zhang D, Zhou B, Li Z, Liu G, et al. 2024. Fine-regulating the carbon flux of L-isoleucine producing Corynebacterium glutamicum WM001 for efficient L-threonine production. ACS Synth Biol. 13(10): 3446–3460. ArticlePubMedLink
References
Supplementary Information
References
Citations

- Designing prokaryotic gene expression regulatory elements: From genomic mining to artificial intelligence-driven generation
Xuan Zhou, Wenyan Cao, Chao Huang, Xiaojuan Zhang, Shenghu Zhou, Yu Deng
Biotechnology Advances.2026; 87: 108781. CrossRef - Green bioconversion of insoluble chitin: chitinase development pathways via multi-strategy synergy
Zhi-Ping Sai, Yi-Rui Yin, Li-Quan Yang, Jia-Hui Wang, Xin-Yi Yang, Fu-Xian Liu, Xin Jing, Yi Zhang, Yu-Da Li, Peng Sang, Zheng-Feng Yang
Bioresources and Bioprocessing.2026;[Epub] CrossRef - Transformer‐Based Prediction of Sec‐ and Tat‐Type Signal Peptides for Enhanced Bacterial Protein Secretion
Seongmo Kang, Seong Min Lee, Ryu Hong Park, Gunhyeong Lee, Je Hyeon Lee, Ki Jun Jeong, Hyun Uk Kim
Biotechnology Journal.2026;[Epub] CrossRef - Engineering of Corynebacterium glutamicum for the enhanced production of optically pure (meso)-2,3-butanediol
Eun Seo Song, Kyeong Ho Kim, Joon Young Lee, Ki Jun Jeong
Bioresources and Bioprocessing.2026;[Epub] CrossRef - Synthetic biology development of microbial strains for liquid biofuel production
Kun Li, Jinsong Xuan, Yingang Feng
Biotechnology Advances.2026; 90: 108891. CrossRef - Fungal factories for feeding the future: mycoproteins as a sustainable protein source at the interface of food, health, and biotechnology innovation
Cristina Firincă, Mariana Constantin, Iuliana Răut, Lucian-Gabriel Zamfir, Maria-Luiza Jecu, Mihaela Doni, Ana-Maria Gurban
Sustainable Food Technology.2026;[Epub] CrossRef - Advancing microbial engineering through synthetic biology
Ki Jun Jeong
Journal of Microbiology.2025; 63(3): e2503100. CrossRef - Time-Series Metabolome and Transcriptome Analyses Reveal the Genetic Basis of Vanillin Biosynthesis in Vanilla
Zeyu Dong, Shaoguan Zhao, Yizhang Xing, Fan Su, Fei Xu, Lei Fang, Zhiyuan Zhang, Qingyun Zhao, Fenglin Gu
Plants.2025; 14(13): 1922. CrossRef - Systems and Synthetic Biology Approaches for Optimizing Microbial Cell Factories
Jongoh Shin, Myung Hyun Noh, Seung-Ho Baek, Jonghyeok Shin, Jung Ho Ahn, Sung Sun Yim, Sungho Jang, Hyun Gyu Lim
KSBB Journal.2025; 40(3): 214. CrossRef - Digital to Biological Translation: How the Algorithmic Data-Driven Design Reshapes Synthetic Biology
Abdul Manan, Nabila Qayyum, Rajath Ramachandran, Naila Qayyum, Sidra Ilyas
SynBio.2025; 3(4): 17. CrossRef
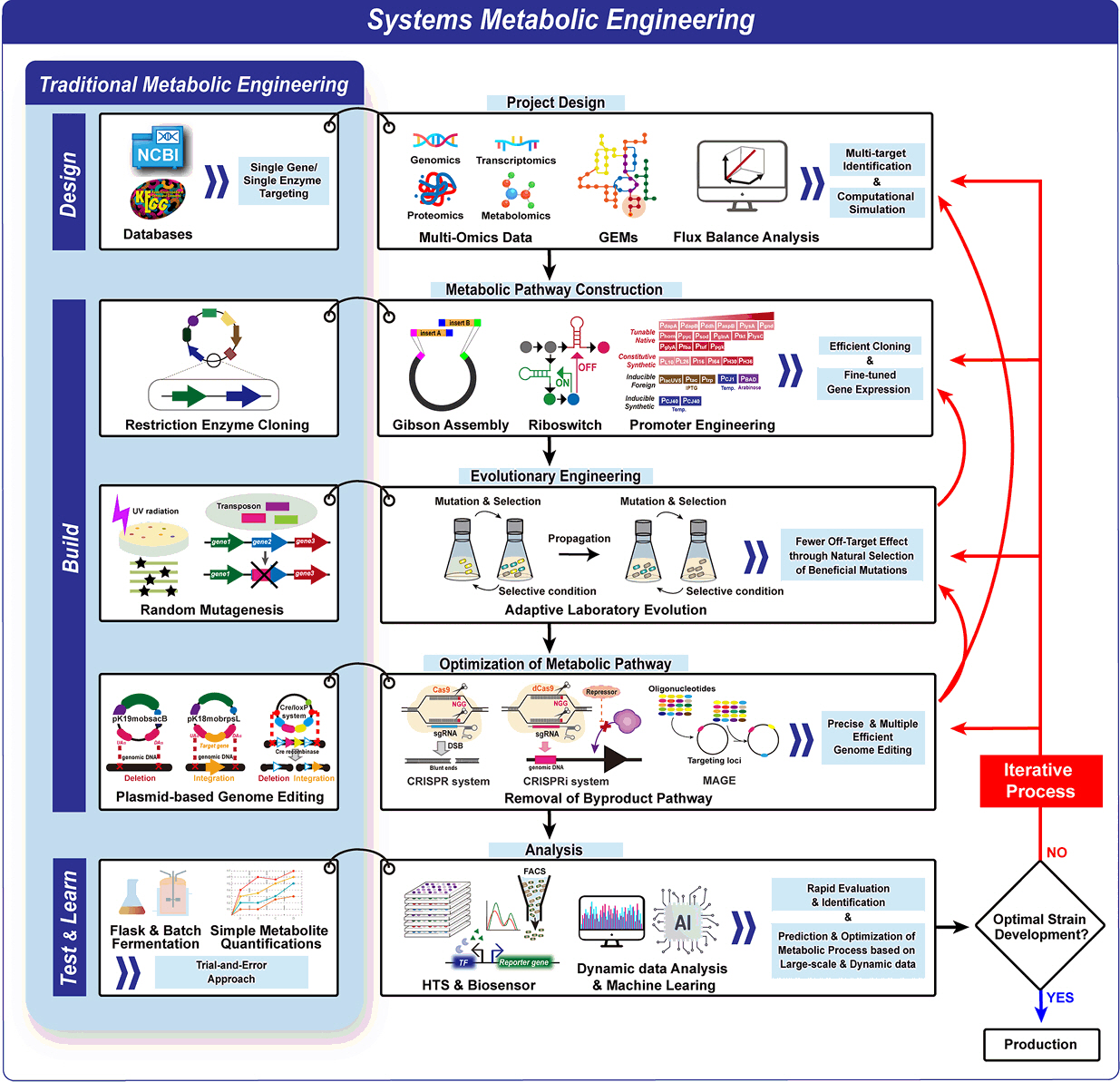
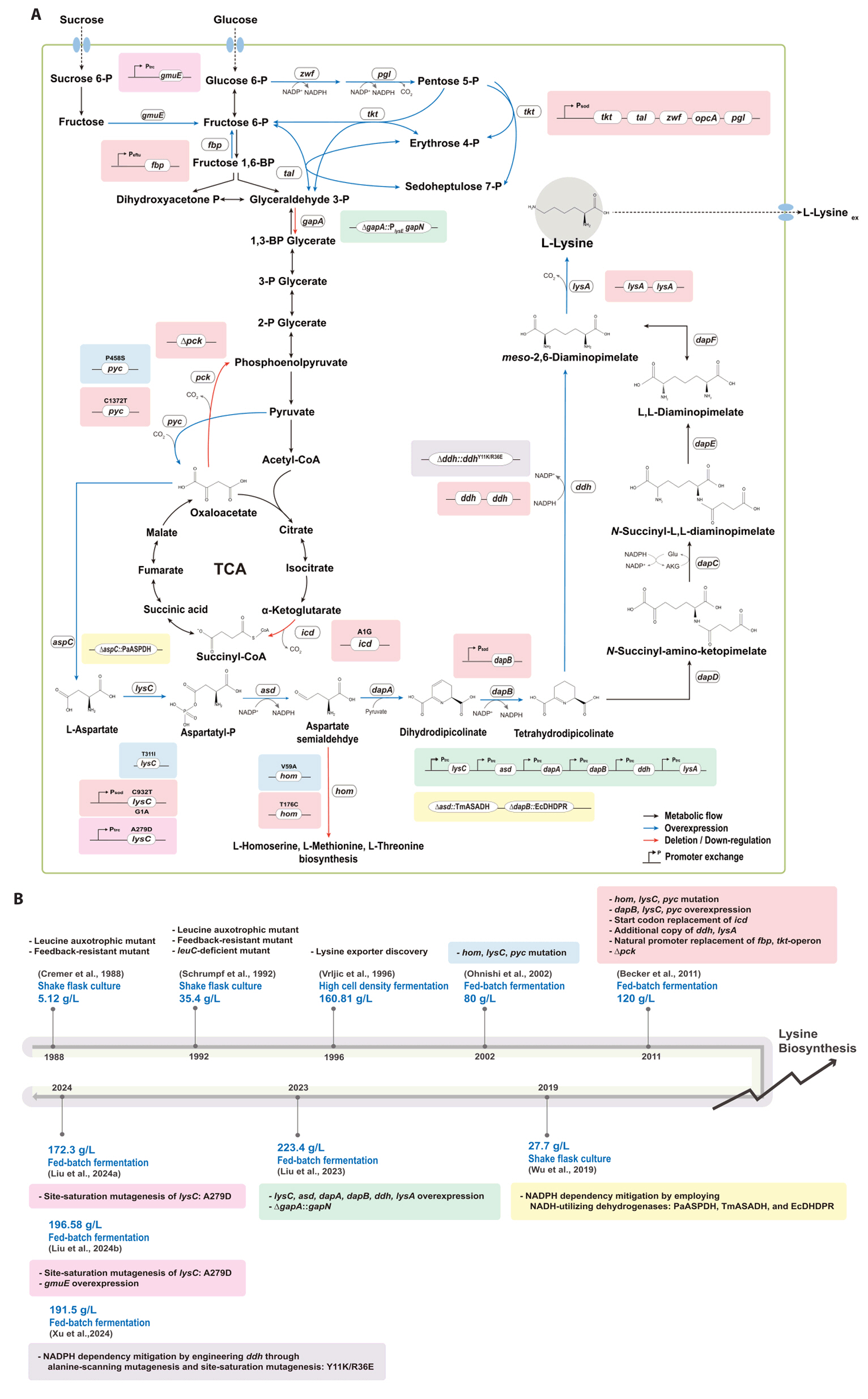
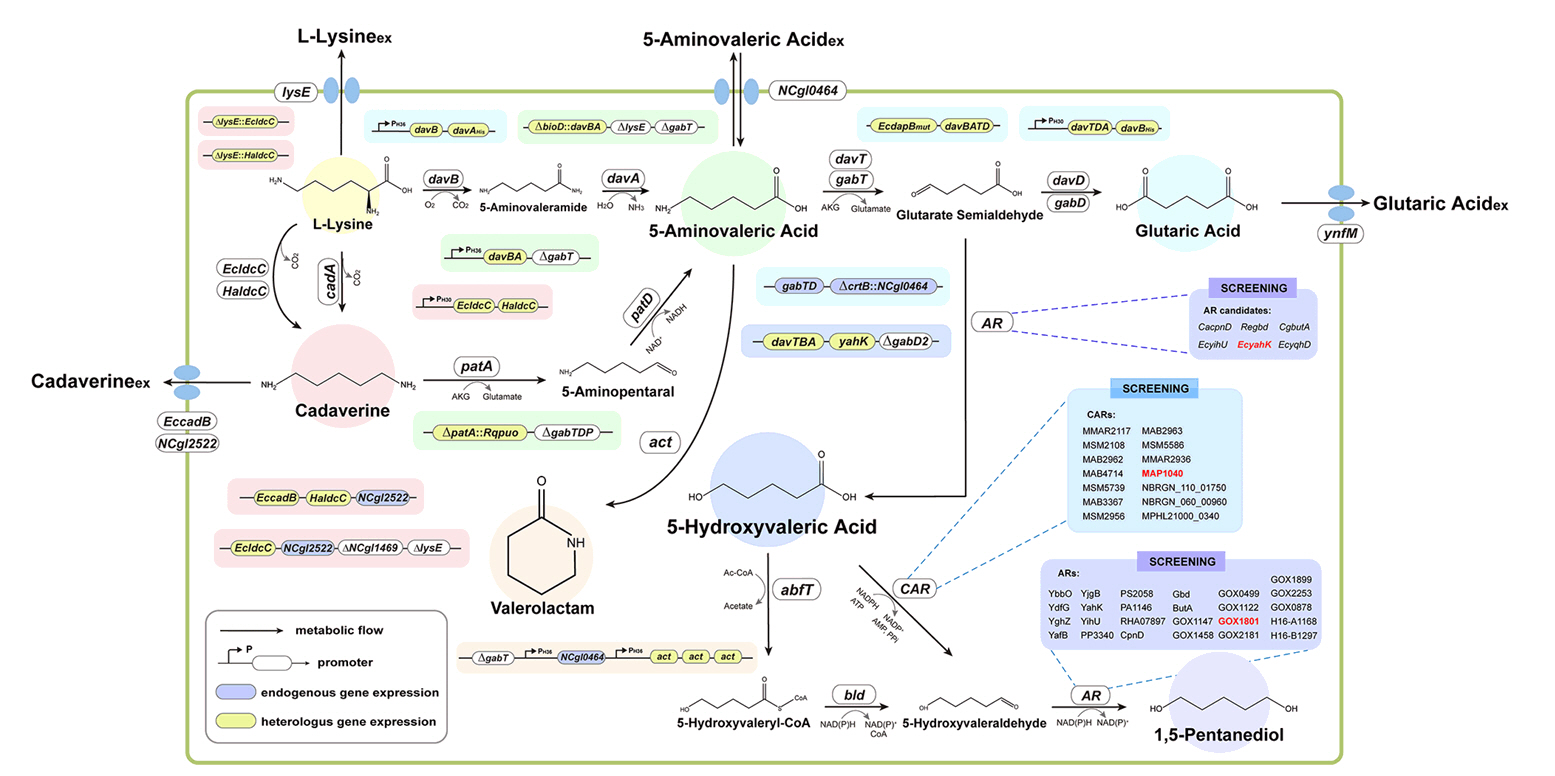
Fig. 1.
Fig. 2.
Fig. 3.
| Product | Host strain | Engineering technique | Titer (g/L) | Scale | References |
|---|---|---|---|---|---|
| Cadaverine | C. glutamicum PKC | √ Chromosomal integration of E. coli derived ldcC with a strong synthetic H30 promoter at the lysE site | 103.78 | Fed-batch | |
| C. glutamicum PKC | √ Chromosomal integration of H. alvei derived ldcC with a strong synthetic H30 promoter at the lysE site | 125 | Fed-batch | ||
| C. glutamicum KCTC 1857 | √ Co-expression of dr1558 and cadA | 10.3 | Fed-batch | ||
| C. glutamicum KCTC 1857 | √ Co-expression of dr1558 and ldcC | 25.1 | Fed-batch | ||
| GTA | C. glutamicum KCTC 1857 | √ Introduction of glutarate biosynthesis pathway by expressing davTDBA genes | 24.5 | Fed-batch | |
| √ Gene modification of davB with an N-terminal His6-tag | |||||
| C. glutamicum BE (C. glutamicum KCTC 12390BP) | √ Identification and expression of 11 target genes for increasing L-lysine supply through gene deletion/integration/substitution along with system-wide analyses | 105.3 | Fed-batch | ||
| √ Overexpression of ynfM | |||||
| C. glutamicum GRLys1 | √ Introduction of glutarate biosynthesis pathway by expressing ldcC, patDA, gabTDStu | 25 | Fed-batch | ||
| √ Gene deletion of sugR, ldhA, snaA, cgmA, and gdh | |||||
| C. glutamicum GSLA2 Δgdh | √ Introduction of glutarate biosynthesis pathway by expressing gltBE686Q, ldcC, patDA, tetA(Z)Δ21bp-gabTDP134L | 22.7 | Fed-batch | ||
| √ Adaptive laboratory evolution | |||||
| 5-AVA | C. glutamicum BE | √ Introduction of 5-AVA biosynthesis pathway by expressing P. putida derived davB and davA | 33.1 | Fed-batch | |
| √ Overexpression of davA by fusing it with His6-Tag at its N-Terminal | |||||
| C. glutamicum GRLys1 | √ Introduction of 5-AVA biosynthesis pathway by expressing E. coil derived ldcC, patA and patD | 5.1 | Shake-flask | ||
| √ Gene deletion of sugR, ldhA, snaA, cgmA, and gabTDP | |||||
| 5-HV | C. glutamicum PKC | √ Introduction of 5-HV biosynthesis pathway by expressing P. putida derived davTBA and E. coil derived yahK | 52.1 | Fed-batch | |
| √ Gene deletion of gabD | |||||
| 1,5-PDO | C. glutamicum PKC ΔgabD2 | √ Introduction of 1,5-PDO biosynthesis pathway by expressing M. marinum derived |
43.4 | Fed-batch | |
| √ Chromosomal integration of PH30DavBHisA expression cassette at the site of lysE | |||||
| √ Enzyme engineering of CAR | |||||
| VL | C. glutamicum XT1 | √ Introduction of valerolactam biosynthesis pathway by expressing P. putida derived davBA and C. propionicum derived act | 12.33 | Fed-batch | |
| √ Dynamic upregulation system using engineered ChnR-B1/Pb-E1 biosensor system | |||||
| C. glutamicum GA16 ΔgabT | √ Gene expression down regulation of gdh using sRNA knock-down system | 76.1 | Fed-batch | ||
| √ Identification and engineering of 5-AVA transporter genes | |||||
| √ Chromosomal integration of multiple copies of act |
Table 1.
TOP
 MSK
MSK
 ePub Link
ePub Link Cite this Article
Cite this Article




