- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Volume 63(3); 2025 > Review
-
Review
Synthetic biology strategies for sustainable bioplastic production by yeasts - Huong-Giang Le1,2, Yongjae Lee3, Sun-Mi Lee3,*
-
Journal of Microbiology 2025;63(3):e2501022.
DOI: https://doi.org/10.71150/jm.2501022
Published online: March 28, 2025
1Division of Energy and Environment Technology, University of Science and Technology (UST), Daejeon 34113, Republic of Korea
2Clean Energy Research Center, Korea Institute of Science and Technology (KIST), Seoul 02792, Republic of Korea
3Division of Environmental Science and Ecological Engineering, Korea University, Seoul 02841, Republic of Korea
- *Correspondence Sun-Mi Lee sunmilee@korea.ac.kr
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
ABSTRACT
- The increasing environmental concerns regarding conventional plastics have led to a growing demand for sustainable alternatives, such as biodegradable plastics. Yeast cell factories, specifically Saccharomyces cerevisiae and Yarrowia lipolytica, have emerged as promising platforms for bioplastic production due to their scalability, robustness, and ease of manipulation. This review highlights synthetic biology approaches aimed at developing yeast cell factories to produce key biodegradable plastics, including polylactic acid (PLA), polyhydroxyalkanoates (PHAs), and poly (butylene adipate-co-terephthalate) (PBAT). We explore recent advancements in engineered yeast strains that utilize various synthetic biology strategies, such as the incorporation of new genetic elements at the gene, pathway, and cellular system levels. The combined efforts of metabolic engineering, protein engineering, and adaptive evolution have enhanced strain efficiency and maximized product yields. Additionally, this review addresses the importance of integrating computational tools and machine learning into the Design-Build-Test-Learn cycle for strain development. This integration aims to facilitate strain development while minimizing effort and maximizing performance. However, challenges remain in improving strain robustness and scaling up industrial production processes. By combining advanced synthetic biology techniques with computational approaches, yeast cell factories hold significant potential for the sustainable and scalable production of bioplastics, thus contributing to a greener bioeconomy.
Introduction
Future Perspectives
Conclusion
Acknowledgments
This work is supported by the National Research Foundation of Korea (NRF) grant funded by the Ministry of Science and ICT (Information & Communication Technology) [grant number RS-2022-00156252] and the Korea University Grant [grant number K2406011].
| Strains | Substrate | Engineering Strategies | Tools | Culture condition | Titer (g/L) | Productivity (g/L/h) | References |
|---|---|---|---|---|---|---|---|
| Saccharomyces cerevisiae SP1130 | Glucose | Express heterologous lactate dehydrogenase (LDH) genes, attenuating several key pathway genes including glycerol-3-phosphate dehydrogenase 1 (GPD1) and cytochrome-c oxidoreductase 2 (CYB2). Knocking out the pyruvate decarboxylase 1 (PDC1) and alcohol dehydrogenase 1 (ADH1) to disrupt the ethanol production pathway, redirect metabolic flux towards lactic acid production. Introduce an alternative pathway to produce acetyl-CoA using the acetylating acetaldehyde dehydrogenase (A-ALD) gene from E. coli to compensate for the attenuated ADH1 | Heterologous gene expression, Introduction of a synthetic pathway | Fed-batch fermentation | 142 | 3.55 | Song et al. (2016) |
| S. cerevisiae T165R | Cane juice | Integrate six copies of L-LDH into the genome of host cell | Multi-integration of a heterologous gene | 1-L jar fermenter | L-LA 122 | - | Saitoh et al. (2005) |
| S. cerevisiae S.c-NO.2-100 | Glucose | Rewire the biosynthetic pathway for LA production by screening and introducing L-LDH, reducing ethanol accumulation by deleting PDC1, ADH1, introducing bacterial acetyl-CoA synthesis pathway. Strain evolution for acid tolerance by adaptive evolution. Delete NADH-consuming enzymes (NDE1/2) to enhance the cofactor availability. Overexpress bacterial transporters JEN1 and remodel the cell membrane by deleting SAM2 for efficient LA export | Heterologous gene expression, Introduction of heterologous transporter, ALE | 5-L batch bioreactor | L-LA 121.5 | 1.69 | Zhu et al. (2022) |
| S. cerevisiae BK01 | Acetate-rich buckwheat husk hydrolysates | Adaptive laboratory evolution (ALE) on 8% lactic acid on S. cerevisiae SR8LDH strain which was previously constructed by the introduction of LDH gene from Lactobacillus acidophilus ATCC4356 under the control of the PGK1 promoter. The mutant BK01 was characterized by genome sequencing and metabolomic profiling | ALE, metabolomic profiling | 20 mL fermentation | L-LA 119 | 1.24 | Jang et al. (2021) |
| S. cerevisiae ASc- d789M | Glucose | Screen D-LDH from bacteria, overexpress LpDLDH, delete glycerol production genes GPD1, GPD2, and lactate dehydrogenase gene DLD1 to minimize by-products, downregulation ADH1 using the L-methionine repressible promoter to minimize the impact on growth. Created an intraspecific hybrid strain by mating the engineered strain with a weak acid-tolerant wild-type strain BCC39850 to improve acid tolerance and D-LA production | Gene knockout by CRISPR/Cas9, CRISPR/Cas12a, yeast mating | Fed-batch fermentation | D-LA 23.41 ± 1.65 | 0.81 | Watcharawipas et al. (2021) |
| S. cerevisiae | Glucose | Genome evolution applied a method combining GMES and CRISPR-δ integration (GMES/CRISPR) to integrate 13 bacterial genes including HXT7, HXK2, PGI1, PFK1, PFK2, FBA1, TPI1, TDH3, PGK1, GPM1, ENO2, and PYK2, and d-lactate dehydrogenase (d-LDH) into genomic DNA | Genome evolution by GMES and CRISPR-δ integration, overlap extension PCR, transcriptome analysis, real-time PCR | Semi-neutralizing fermentation | D-LA 52.2 | 2.17 | Mitsui et al. (2020) |
| S. cerevisiae JHY5330 | Glucose | Introduce D-lactate dehydrogenase gene (ldhA) from bacteria, reduce D-LA consumption by deleting DLD1 encoding D-lactate dehydrogenase and JEN1 encoding monocarboxylate transporter, eliminate ethanol production gene PDC1 and ADH1, glycerol production genes GPD1 and GPD2, improve strain LA tolerance by ALE and overexpression of HAA1 encoding a transcriptional activator involved in weak acid stress response | ALE, gene knockout, PCR-mediated method using the Cre/loxP system, quantitative reverse transcription PCR (qRT-PCR) | Fed-batch fermentation | D-LA 112 | 2.2 | Baek et al. (2016) |
| S. cerevisiae JHY5610 | Glucose | Employed ALE for LA-tolerance of host strain, coupled with introduction of LA production gene Lm. ldhA, and deletion of ethanol and glycerol production genes (ADH, GPD) and D-LA degradation genes to redirect metabolic flux toward LA production | ALE, gene knockout, whole genome sequencing | Fed-batch fermentation | D-LA 82.6 | 1.5 | Baek et al. (2017) |
| S. cerevisiae | Glucose | Rewire metabolic fluxes toward the production of L-lactic acid by deleting PDC1, CYB2, GPD1, and replacing LDH. Re-engineer the intracellular redox balance by deleting NDE1 and NDE2 encoding isoenzymes of the external NADH dehydrogenase | Gene knockout, re-engineer intracellular redox | Fed-batch | L-LA 117 | - | Lee et al. (2015) |
| Pichia pastoris | Methanol | Screen LDH genes, co-utilize LDH cofactors, knockout DLD1, DLD2 and CYB2 to block LA consumption pathway, and construct a coordinated dual pathway in cytoplasm and mitochondria | Site-directed mutation, CRISPR-Cas9, fluorescence microscopy analysis | Fed-batch fermentation | L-LA 4.2 | 0.0126 | Wu et al. (2025) |
| Pichia kudriavzevii NG7 | Glucose | Replace the pyruvate decarboxylase 1 gene (PDC1) with the d-lactate dehydrogenase gene (d-LDH) derived from Lactobacillus plantarum, improve strain tolerance by ALE | ALE, whole genome sequencing | Bioreactor fed-batch | D-LA 154 | 4.16 | Park et al. (2018) |
| Candida boidinii | Glucose | Disrupted PDC1 to reduce ethanol production, introduce and express Bovine L-LDH gene under the PDC1 promoter and optimize fermentation condition | Heterologous gene expression | Batch fermentation | L-LA 85.9 | 1.79 | Osawa et al. (2009) |
| S. cerevisiae | Glucose | Poly (D-lactic acid) (PDLA) and copolymer P (LA-3HB) were produced by express PHA synthase PhaC1437Ps6-19, propionyl-CoA transferase Pct540Cp, acetyl-CoA acetyltransferase PhaA, and acetoacetyl-CoA reductase PhaB1 | Gibson Assembly, modular cloning method (MoClo), GC-MS, NMR analysis | Erlenmeyer baffled flasks fermentation | PDLA 0.73% (CDW) | - | Ylinen et al. (2021) |
| Yarrowia lipolytica | Glucose, racemic lactic acid | Disrupt lactic acid consumption pathway, introduce heterologous pathway for PDLA production containing propionyl-CoA transferase (pct) from Clostridium propionicum to convert lactic acid into lactyl-CoA, and an evolved polyhydroxyalkanoic acid PHA synthase polymerizing lactyl-CoA into PDLA | Site-directed mutagenesis, UHPLC-FTMS | Erlenmeyer baffled flasks fermentation | PDLA 26 mg/g-DCW | - | Lajus et al. (2020) |
| Strains | Substrate | Engineering Strategies | Tools | Culture condition | Titer | References |
|---|---|---|---|---|---|---|
| E. coli BL21 (DE3) | Glycerol | Reconstruct five-step reverse adipate-degradation pathway (RADP) from Thermobifida fusca in E. coli, then overexpress 5-Carboxy-2-pentenoyl-CoA reductase (Tfu_1647) - the gene coding for the rate-limiting step in the RADP, delete succinate-CoA ligase gene (sucD) to eliminate competitive | Introduction of a synthetic pathway, Gibson assembly, CRISPR/Cas9, gene overexpression, SDS-PAGE | Fed-batch fermentation in 5-L bioreactor | 68.0 g/L | Zhao et al. (2018) |
| Candida spp. | Fatty acid | Rewire the ω- then the β-oxidation pathway | - | Two-stage fed-batch process | 50 g/L | Beardslee & Picataggio (2012) |
| Candida tropicalis | C12 methyl laurate | Create a mutant strain of C. tropicali by deleting AOX genes encoding acyl-CoA oxidases to produce AA via ω- and the β-oxidation pathway | Gene deletion | Bioreactor, fed-batch | 12.1 g/L | Ju et al. (2020) |
| S. cerevisiae | Glucose | Use the reverse adipate degradation pathway (RADP) from Thermobifida fusca, co-expressing genes of Tfu_ 0875, Tfu_2399, Tfu_0067, Tfu_1647, Tfu_2576, and Tfu_ 2576 in S. cerevisiae | Introduction of a synthetic pathway, Gibson assembly, overlap extension PCR | Fed-batch bioreactor | 10.09 mg/L | Zhang et al. (2020) |
| S. cerevisiae | Glucose | Rewire the cis, cis-muconic acid pathway by expressing enoate reductases (ERs) from Bacillus coagulans to convert ccMA into AA | Introduction of a synthetic pathway, Gibson assembly | Three stage fermentations | 2.59 mg/L, ccMA > 284 mg/L | Raj et al. (2018) |
| Y. lipolytica | Fatty acid methyl esters | Regulate the β-oxidation pathway by disrupting acyl-CoA oxidases (POX1 and POX3) and enhance ω-oxidation through overexpression of ALK5, CPR1, and FAO1, two-phase bioprocess to enhance AA production | Gene deletion, two-phase bioprocess | 20 ml in 250 ml baffled flask | 1176.2 mg/L | Lee et al. (2024) |
| Strains | Substrate | Engineering Strategies | Tools | Culture condition | Titer (g/L) | Productivity (g/L/h) | References |
|---|---|---|---|---|---|---|---|
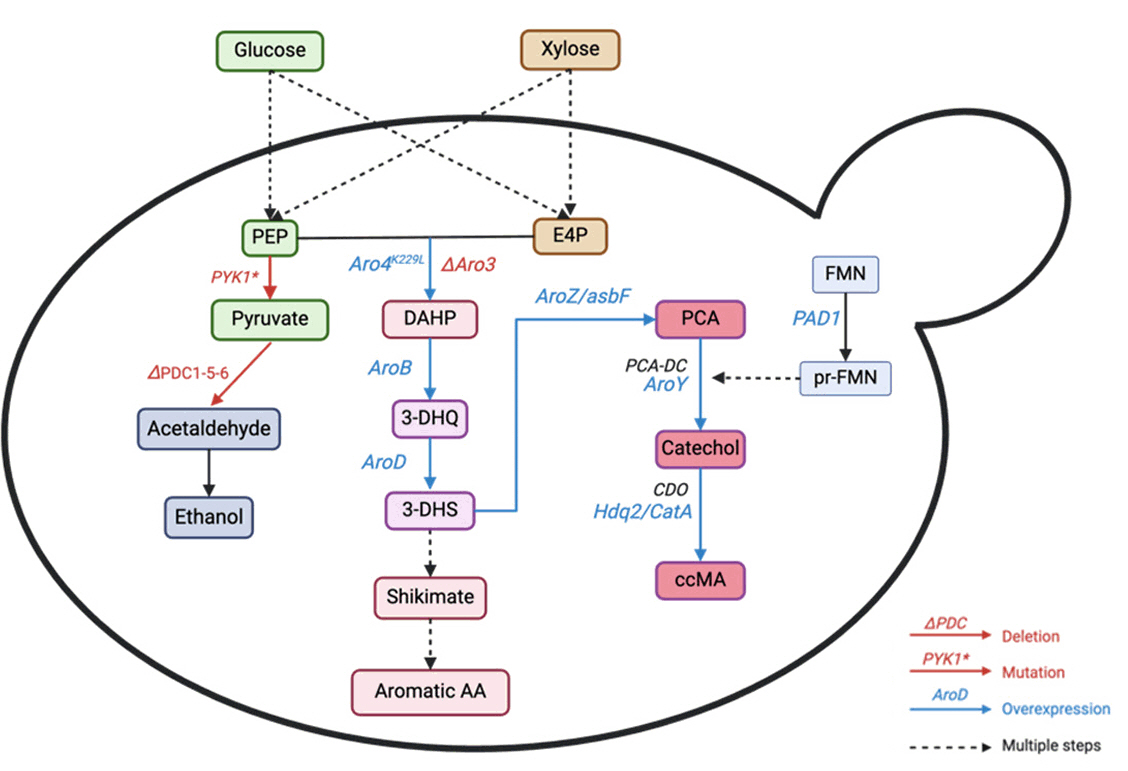
| S. cerevisiae | Glucose | Employed ccMA-biosensor, coupled with GFP expression to screen UV-mutagenesis libraries for ccMA producing yeast, then overexpress genes encoding for PCA decarboxylase and AROM protein and restore URA3 prototrophy. | UV-mutagenesis, fluorescence activated cell sorting (FACS), heterologous gene expression | Fed-batch | 20.8 | 0.139 | Wang et al. (2020) |
| S. cerevisiae | Glucose | Rewiring the shikimate pathway flux and enhancing the phosphoenolpyruvate supply by mutating PYK1 (A336S) to limit PEP-to-pyruvate conversion and overexpressing PYC1 and PCK1 to redirect pyruvate to PEP, overexpress DAHP synthase (ScARO4K229L), 3-dehydroquinate synthase (EcaroB), and 3-dehydroquinase | Cas9-assisted approach, Gibson assembly, USER cloning, EasyClone method | 2L fermenter | 22.5 | 0.19 | Wang et al. (2022) |
| S. cerevisiae | Glucose, xylose, mixture | Eliminated feedback inhibition in the shikimate pathway, insert heterologous pathway for MA production using 3-dehydroshikimate (DHS), protocatechuic acid decarboxylase (PCAD) and oxygen-consuming catechol 1,2-dioxygenase (CDO), delete PDC to eliminate ethanol production, minimized PCA production by enhancing PCAD overexpression and production of its co-factor | Introduction of a synthetic pathway, genome editing using CRISPR/Cas9 | 300 ml shake flasks with 40 mL YP medium | 4.5 | - | Nicolai et al. (2021) |
| S. cerevisiae | Glucose and xylose | Eliminates C2 dependency and ethanol production by ALE and deleting PDC1, PDC5, and PDC6, improving MA tolerance for host strain, promoting growth and production of target product by internal deletion of MTH1, MTH1ΔM41-T78, overexpressing PAD1 as well as reduces side production of intermediate protocatechuic acid (PCA) | Introduction of a synthetic pathway, enzyme engineering, ALE | Fed-batch | 9.3 | 0.100 | Tönjes et al. (2024) |
| S. cerevisiae | Glucose | Combine adaptive laboratory evolution (ALE) and rational metabolic engineering, improve flux by truncation of ARO1 and overexpress an endogenous aromatic decarboxylase | ALE, Gibson assembly | Fed-batch bioreactor | 2.1 | - | Leavitt et al. (2017) |
| S. cerevisiae | Glucose | Employ the computational tool YEASTRACT for predicting novel transcriptional repressors and OptForce strain-design for identifying non-intuitive pathway interventions, strating from glycolytic and pentose phosphate pathway pathway, follow by engineer ARO2, ARO3, ARO4, and the pentafunctional ARO, delete ric1 to increase transcription of ARO2, ARO3, ARO4 | Computational tool for predicting engineering target, Introduction of a synthetic pathway Gibson assembly | - | 0.32 | - | Suástegui et al. (2017) |
| S. cerevisiae | Glucose, xylose (supplement 1 g/L catechol) | Overexpress xylose isomerase gene from Bacteroides valgutus and pentose phosphate pathway genes from S. cerevisiae, then imported a three step ccMA production pathway from E. coli with AroZ-Neu, AroY-Com, and CatA-Cup genes, further overexpression of gene Aro1 (with a stop codon of AroE) and feedback-resistant Aro4opt mutant gene, ALE to improve xylose fermentation and ccMA | Introduction of a synthetic pathway, Enzyme engineering, ALE | Batch, shake flask | 0.424 (1.286 with catechol supplemented) | - | Liu et al. (2020) |
| S. cerevisiae | Glucose, supplement amino acids | Balance MA pathway performance with aromatic amino acid prototrophy by destabilizing Aro1, delete ARO4 and ARO3, introduce AroZ, AroY, Hqd2 CatA, Pad1. | CRISPR-mediated homology-directed repair (HDR) | Fed-batch bioreactor | 5.1 | - | Pyne et al. (2018) |
| S. cerevisiae | Glucose | Construct cassettes using USER cloning-ligation-PCR, express a three-step heterologous pathway using KpAroY.B, KpAroY.Ciso, CaCatA, engineering TKL1, ZWF1, Aro1AroEΔ, Aro4K229L | Introduction of a synthetic pathway, CRISPR/Cas9, in vivo recombination USER assembly and RNA interference | Batch, shake flask | 0.8 | - | Kildegaard et al. (2019) |
| Pichia occidentalis | Glucose | Screen host strain with high tolerance to MA, develop genome editing toolkit to engineer the non-conventional yeast and introduce the heterologous pathway for ccMA production | CRISPR-Cas9, antibiotic marker recycling, high-throughput screening | Fed-batch | 38.8 | 0.511 | Pyne et al. (2023) |
| Strains | Substrate | Engineering Strategies | Tools | Culture condition | Titer (mg/L) | Yield (mg/g) | References |
|---|---|---|---|---|---|---|---|
| E. coli | Glucose, xylose | Knocked out ndh gene. Introduced arabinose/xylose transport proteins from Bacillus subtilis (xylA, xylB, and araE). Incorporated the PHB biosynthesis pathway from C. necator | Modification of ribosome binding sites | Fed-batch culture | PHB 21010 | - | Huo et al. (2017) |
| S. cerevisiae | Silver grass, glucose, xylose | Constructed a polycistronic PHB cassette with phaA, phaB, and phaC using 2A peptide sequences. Introduced Cas9-expressing plasmid, gRNA, and polycistronic PHB cassette into S. cerevisiae via electroporation. Integrated the polycistronic PHB cassette into the genome | CRISPR-Cas9 mediated genome editing, 2A peptide sequence integration | Serum bottle, microaerobic | PHB 40.3 | 0.93 mg/g silver grass | Tran et al. (2023) |
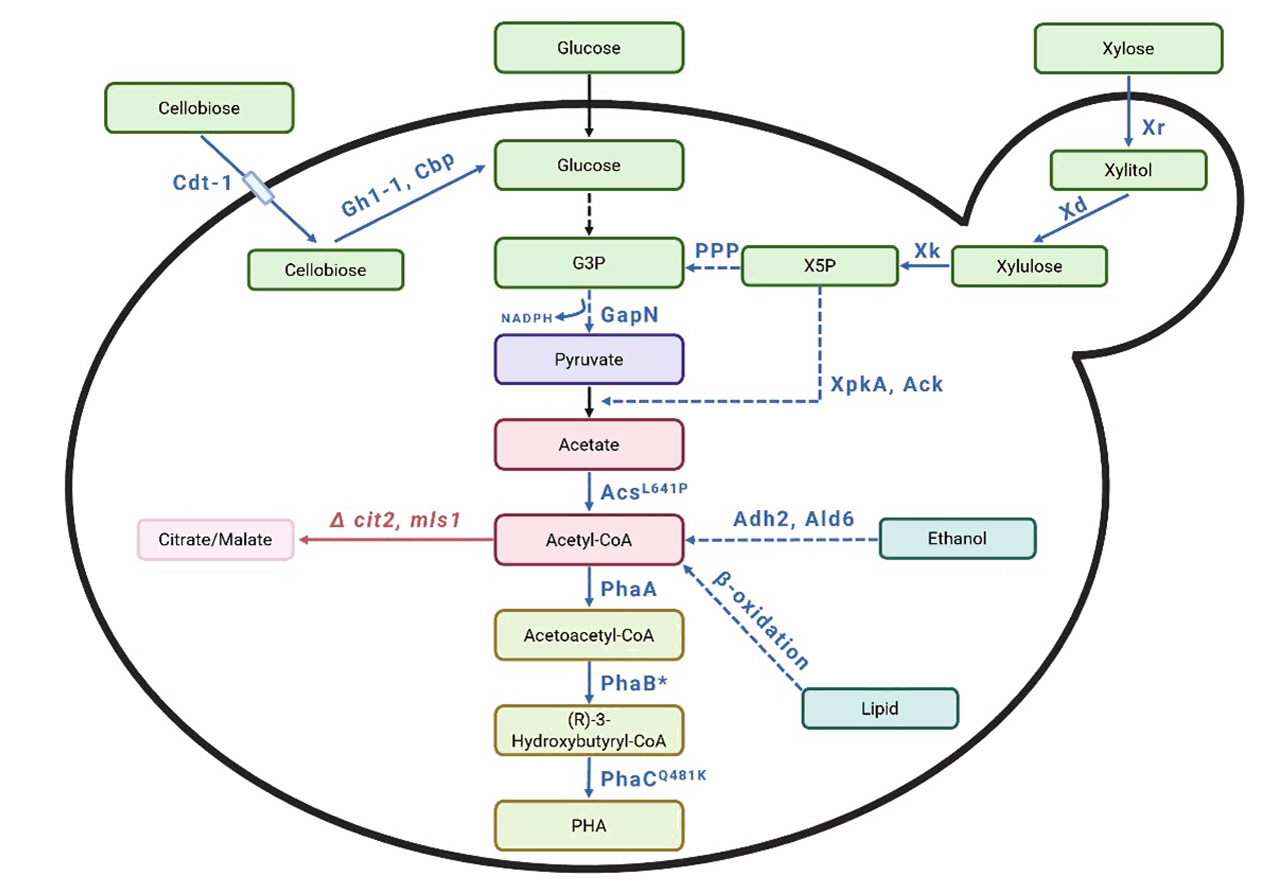
| S. cerevisiae | Glucose | Expressed XpkA and Ack from Aspergillus nidulans to increase acetyl-CoA levels. Incorporated an acetyl-CoA synthase variant (AcsL641P) from Salmonella enterica. Integrated GapN (NADP⁺-dependent glyceraldehyde-3-phosphate dehydrogenase) from Streptococcus mutans into the genome via the Li-Ac method. Codon optimized all genes for efficient expression | Codon optimization DNA 2.0 and GenScript for synthesizing genes, PTEF1-PPGK1 bidirectional promoter | Shake flask, anaerobic | PHB 180 | 0.03 mg/g glucose | Kocharin et al. (2013) |
| S. cerevisiae | Glucose | Synthesized phaA, phaB, and phaC based on R. eutropha H16. Introduced them into S. cerevisiae in plasmid form using the Li-Ac method. Used the strain described by Chen et al. (2012) to increase acetyl-CoA levels. Deleted cit2 and mls1, overexpressed ALD6 and ADH2. Codon optimized all genes for efficient expression | Codon optimization, PTEF1-PPGK1 bidirectional promoter, DNA 2.0 for synthesize genes | Shake flask, aerobic | PHB 43.11 | 0.13 mg/g glucose | Kocharin et al. (2012) |
| S. cerevisiae | Xylose | Introduced XR and XD genes to enable xylose utilization. PCR-amplified phaA, phaB, and phaC genes from C. necator. Assembled these genes into a single plasmid using In-Fusion cloning. Introduced the plasmid into S. cerevisiae via the Li-Ac method. Codon optimized all genes for efficient expression | In-fusion cloning, Codon optimization, overlap extension PCR | Shake flask, aerobic | PHB 45 | 1.17 mg/g xylose | Sandström et al. (2015) |
| S. cerevisiae | Xylose | Achieved PHB production under anaerobic conditions using xylose as the carbon source. Replaced PhaB from C. necator with NADH-dependent PhaB from Allochromatium vinosum. Introduced the gene into S. cerevisiae in plasmid form via the Li-Ac method | Codon optimization, heterologous gene expression, promoter and terminator pairing | Bioreactor, anaerobic | PHB 360 | 7 mg/g xylose | Muñoz de Las Heras et al. (2016) |
| S. cerevisiae | Xylose high nitrogen supply | Investigated whether nitrogen limitation in yeast influences PHB production. Introduced Xr and Xd from Scheffersomyces stipitis to enable xylose utilization. Replaced PhaB from C. necator with NADH-dependent PhaB from A. vinosum. Assembled these genes into a single plasmid using In-Fusion cloning. Introduced the plasmid into S. cerevisiae via the Li-Ac method. Codon optimized and PCR-amplified all genes | In-fusion cloning, Codon optimization, heterologous gene expression, promoter and terminator pairing | Shake flask, anaerobic | PHB 730 | 13.8 mg/g xylose | Portugal-Nunes et al. (2017) |
| S. cerevisiae | Cellobiose | Integrated phaA, phaB1, and phaC1 into the S. cerevisiae genome using Gibson assembly, Li-Ac method, and CRISPR-Cas9. Inserted Gh1-1 from Neurospora crassa and Cbp from Ruminococcus flavefaciens for cellobiose metabolism using the Modular Cloning (MoClo) approach. Codon optimized all genes for efficient expression | Gibson Assembly, Modular cloningCRISPR-Cas9 mediated genome integration, promoter and terminator pairing | Bioreactor | PHB - | 49.4 mg/g cellobiose | Ylinen et al. (2022) |
| Y. lipolytica | Tridecanoate | Directed carbon flux in Y. lipolytica toward either the β-oxidation pathway or the PHA biosynthesis pathway based on the genotype of POX1-6 genes encoding acyl-CoA oxidase isoenzymes. Downregulated the R-3-hydroxyacyl-CoA dehydrogenase multifunctional enzyme domain in the fatty acid β-oxidation pathway via site-directed mutagenesis, generating the MFE-AG16SBC variant. Deleted an acyltransferase gene and overexpressed enoyl-CoA hydratase 2 to further redirect fatty acids from lipid synthesis to the β-oxidation pathway | site-directed mutagenesis, loxP-Cre recombinase, pGEMT easy cloning | Shake flask | PHA - | 7.3% DCW | Haddouche et al. (2011) |
| Y. lipolytica | Tridecanoate | Directed carbon flux in Y. lipolytica toward either the β-oxidation pathway or the PHA biosynthesis pathway based on the genotype of POX1-6 genes encoding acyl-CoA oxidase isoenzymes. Introduced PhaC from Pseudomonas aeruginosa and modified its C-terminal region to include the 34-amino acid sequence from Brassica napus glyoxysomal isocitrate lyase (icl) for peroxisome targeting | Site-directed mutagenesis, STADEN package for sequence analysis | Shake flask | PHA - | 17.64 mg/g biomass | Haddouche et al. (2010) |
| Y. lipolytica | Methyl laurate | Generated various PhaC variants through site-directed mutagenesis. Compared their production yields of mcl-PHA with different chain lengths | Site-directed mutagenesis | - | PHA - | 28% DCW | Rigouin et al. (2019) |
| Y. lipolytica | Triolein | Directed carbon flux in Y. lipolytica toward either the β-oxidation pathway or the PHA biosynthesis pathway based on the genotype of POX1-6 genes encoding acyl-CoA oxidase isoenzymes. Heterologously expressed PhaC1 from Pseudomonas aeruginosa PAO1 with a peroxisomal signal. Introduced the gene into S. cerevisiae in plasmid form via the Li-Ac method. Codon optimized all genes for efficient expression | Codon optimization | Shake flask | PHA 1110 | 5.0% DCW | Gao et al. (2015) |
- Agarwal P, Soni R, Kaur P, Madan A, Mishra R, et al. 2022. Cyanobacteria as a promising alternative for sustainable environment: Synthesis of biofuel and biodegradable plastics. Front Microbiol. 13: 939347.ArticlePubMedPMC
- Ali SS, Elsamahy T, Al-Tohamy R, Zhu D, Mahmoud YAG, et al. 2021. Plastic wastes biodegradation: Mechanisms, challenges and future prospects. Sci Total Environ. 780: 146590.ArticlePubMed
- Alonso-Gutierrez J, Chan R, Batth TS, Adams PD, Keasling JD, et al. 2013. Metabolic engineering of Escherichia coli for limonene and perillyl alcohol production. Metab Eng. 19: 33–41. ArticlePubMed
- Amaradio MN, Jansen G, Costanza J, Patanè A, Branduardi P, et al. 2023. L-lactate production in engineered Saccharomyces cerevisiae using a multistage multiobjective automated design framework. Biotechnol Bioeng. 120(7): 1929–1952. ArticlePubMed
- Averesch NJH, Martínez VS, Nielsen LK, Krömer JO. 2018. Toward synthetic biology strategies for adipic acid production: An in silico tool for combined thermodynamics and stoichiometric analysis of metabolic networks. ACS Synth Biol. 7(2): 490–509. ArticlePubMed
- Baek SH, Kwon EY, Bae SJ, Cho BR, Kim SY, et al. 2017. Improvement of D-lactic acid production in Saccharomyces cerevisiae under acidic conditions by evolutionary and rational metabolic engineering. Biotechnol J. 12(10): 1700015.Article
- Baek SH, Kwon EY, Kim YH, Hahn JS. 2016. Metabolic engineering and adaptive evolution for efficient production of D-lactic acid in Saccharomyces cerevisiae. Appl Microbiol Biotechnol. 100(6): 2737–2748. ArticlePubMedPDF
- Balasubramanian VK, Muthuramalingam JB, Chen YP, Chou JY. 2023. Recent trends in lactic acid-producing microorganisms through microbial fermentation for the synthesis of polylactic acid. Arch Microbiol. 206(1): 31.ArticlePubMedPDF
- Beardslee T, Picataggio S. 2012. Bio-based adipic acid from renewable oils. Lipid Technol. 24(10): 223–225. ArticleLink
- Brown B, Immethun C, Wilkins M, Saha R. 2022. Biotechnical applications of phasins: Small proteins with large potential. Renew Sustain Energy Rev. 158: 112129.Article
- Cachera P, Kurt NC, Ropke A, Strucko T, Mortensen UH, et al. 2024. Genome-wide host-pathway interactions affecting cis-cis-muconic acid production in yeast. Metab Eng. 83: 75–85. ArticlePubMed
- Carruthers DN, Lee TS. 2022. Translating advances in microbial bioproduction to sustainable biotechnology. Front Bioeng Biotechnol. 10: 968437.ArticlePubMedPMC
- Casalini T, Rossi F, Castrovinci A, Perale G. 2019. A perspective on polylactic acid-based polymers use for nanoparticles synthesis and applications. Front Bioeng Biotechnol. 7: 259.ArticlePubMedPMC
- Chek MF, Hiroe A, Hakoshima T, Sudesh K, Taguchi S. 2019. PHA synthase (PhaC): Interpreting the functions of bioplastic-producing enzyme from a structural perspective. Appl Microbiol Biotechnol. 103(3): 1131–1141. ArticlePubMedPDF
- Chen GQ. 2010. Introduction of bacterial plastics PHA, PLA, PBS, PE, PTT, and PPP. In Chen GQ. (ed.), Plastics from Bacteria: Natural Functions and Applications, Microbiology Monographs, Vol. 14, pp. 1–16, Springer.
- Chen Y, Banerjee D, Mukhopadhyay A, Petzold CJ. 2020. Systems and synthetic biology tools for advanced bioproduction hosts. Curr Opin Biotechnol. 64: 101–109. ArticlePubMed
- Chen YJ, Tsai PC, Hsu CH, Lee CY. 2014. Critical residues of class II PHA synthase for expanding the substrate specificity and enhancing the biosynthesis of polyhydroxyalkanoate. Enzyme Microb Technol. 56: 60–66. ArticlePubMed
- Choi SS, Lee HN, Park EH, Lee SJ, Kim ES. 2020. Recent advances in microbial production of cis, cis-muconic acid. Biomolecules. 10(9): 1238.ArticlePubMedPMC
- Choi SS, Seo SY, Park SO, Lee HN, Song JS, et al. 2019. Cell factory design and culture process optimization for dehydroshikimate biosynthesis in Escherichia coli. Front Bioeng Biotechnol. 7: 241.ArticlePubMedPMC
- Choi B, Tafur Rangel A, Kerkhoven EJ, Nygard Y. 2024. Engineering of Saccharomyces cerevisiae for enhanced metabolic robustness and L-lactic acid production from lignocellulosic biomass. Metab Eng. 84: 23–33. ArticlePubMed
- Dasgupta A, Chowdhury N, De RK. 2020. Metabolic pathway engineering: Perspectives and applications. Comput Methods Programs Biomed. 192: 105436.ArticlePubMed
- De Souza FM, Gupta RK. 2024. Bacteria for bioplastics: Progress, applications, and challenges. ACS Omega. 9(8): 8666–8686. ArticlePubMedPMCLink
- Elvers D, Song CH, Steinbüchel A, Leker J. 2016. Technology trends in biodegradable polymers: Evidence from patent analysis. Polym Rev. 56(4): 584–606. Article
- Emadian SM, Onay TT, Demirel B. 2017. Biodegradation of bioplastics in natural environments. Waste Manag. 59: 526–536. ArticlePubMed
- European-Bioplastics. 2023. Bioplastics market development update 2023. Available from: https://www.european-bioplastics.org/bioplastics-market-development-update-2023-2/
- Feehan R, Montezano D, Slusky JSG. 2021. Machine learning for enzyme engineering, selection and design. Protein Eng Des Sel. 34: gzab019.ArticlePubMedPMC
- Ferreira FV, Cividanes LS, Gouveia RF, Lona LMF. 2019. An overview on properties and applications of poly(butylene adipate-co-terephthalate)-PBAT based composites. Polym Eng Sci. 59(s2): E7–E15. ArticleLink
- Gao C, Qi Q, Madzak C, Lin CSK. 2015. Exploring medium-chain-length polyhydroxyalkanoates production in the engineered yeast Yarrowia lipolytica. J Ind Microbiol Biotechnol. 42(9): 1255–1262. ArticlePubMedPDF
- Garner KL. 2021. Principles of synthetic biology. Essays Biochem. 65(5): 791–811. ArticlePubMedPMCPDF
- Gohil N, Panchasara H, Patel S, Ramírez-García R, Singh V. 2017. Book review: Recent advances in yeast metabolic engineering. Front Bioeng Biotechnol. 5: 71.ArticlePMC
- Guettler MV, Jain MK, Rumler D. 1996. Method for making succinic acid, bacterial variants for use in the process, and methods for obtaining variants. Available from: https://patents.google.com/patent/US5573931A/en
- Gurdo N, Volke DC, McCloskey D, Nikel PI. 2023. Automating the design-build-test-learn cycle towards next-generation bacterial cell factories. New Biotechnol. 74: 1–15. ArticlePubMed
- Haddouche R, Delessert S, Sabirova J, Neuvéglise C, Poirier Y, et al. 2010. Roles of multiple acyl-CoA oxidases in the routing of carbon flow towards beta-oxidation and polyhydroxyalkanoate biosynthesis in Yarrowia lipolytica. FEMS Yeast Res. 10(7): 917–927. ArticlePubMed
- Haddouche R, Poirier Y, Delessert S, Sabirova J, Pagot Y, et al. 2011. Engineering polyhydroxyalkanoate content and monomer composition in the oleaginous yeast Yarrowia lipolytica by modifying the β-oxidation multifunctional protein. Appl Microbiol Biotechnol. 91(5): 1327–1340. ArticlePubMedPDF
- Hazer B, Steinbüchel A. 2007. Increased diversification of polyhydroxyalkanoates by modification reactions for industrial and medical applications. Appl Microbiol Biotechnol. 74(1): 1–12. ArticlePubMedPDF
- Henton DE, Gruber P, Lunt J, Randall J. 2005. Polylactic acid technology. In Natural fibers, biopolymers, and biocomposites, pp. 559–607. CRC Press.
- Hong KK, Nielsen J. 2012. Metabolic engineering of Saccharomyces cerevisiae: A key cell factory platform for future biorefineries. Cell Mol Life Sci. 69(16): 2671–2690. ArticlePubMedPMCPDF
- Huang S, Xue Y, Yu B, Wang L, Zhou C, et al. 2021. A review of the recent developments in the bioproduction of polylactic acid and its precursors optically pure lactic acids. Molecules. 26(21): 6446.ArticlePubMedPMC
- Huo G, Zhu Y, Liu Q, Tao R, Diao N, et al. 2017. Metabolic engineering of an E. coli ndh knockout strain for PHB production from mixed glucose-xylose feedstock. J Chem Technol Biotechnol. 92(10): 2739–2745. Article
- Jang BK, Ju YB, Jeong DY, Jung SK, Kim CK, et al. 2021. L-Lactic acid production using engineered Saccharomyces cerevisiae with improved organic acid tolerance. J Fungi (Basel). 7(11): 928.ArticlePubMedPMC
- Jayakumar A, Radoor S, Siengchin S, Shin GH, Kim JT. 2023. Recent progress of bioplastics in their properties, standards, certifications and regulations: A review. Sci Total Environ. 878: 163156.ArticlePubMed
- Ju JH, Oh BR, Heo SY, Lee YU, Shon JH, et al. 2020. Production of adipic acid by short- and long-chain fatty acid acyl-CoA oxidase engineered in yeast Candida tropicalis. Bioprocess Biosyst Eng. 43(1): 33–43. ArticlePubMedPDF
- Jung YK, Kim TY, Park SJ, Lee SY. 2010. Metabolic engineering of Escherichia coli for the production of polylactic acid and its copolymers. Biotechnol Bioeng. 105(1): 161–171. ArticlePubMed
- Kayitesi E, Onojakpor O, Moyo SM. 2023. Highlighting the impact of lactic-acid-bacteria-derived flavours or aromas on sensory perception of African fermented cereals. Fermentation. 9(2): 111.Article
- Kildegaard KR, Tramontin LRR, Chekina K, Li M, Goedecke TJ, et al. 2019. CRISPR/Cas9-RNA interference system for combinatorial metabolic engineering of Saccharomyces cerevisiae. Yeast. 36(5): 237–247. ArticlePubMedPMCLink
- Kim GB, Kim WJ, Kim HU, Lee SY. 2020. Machine learning applications in systems metabolic engineering. Curr. Opin. Biotechnol. 64: 1–9. ArticlePubMed
- Kocharin K, Chen Y, Siewers V, Nielsen J. 2012. Engineering of acetyl-CoA metabolism for the improved production of polyhydroxybutyrate in Saccharomyces cerevisiae. AMB Express. 2: 52.ArticlePubMedPMCPDF
- Kocharin K, Siewers V, Nielsen J. 2013. Improved polyhydroxybutyrate production by Saccharomyces cerevisiae through the use of the phosphoketolase pathway. Biotechnol Bioeng. 110(8): 2216–2224. ArticlePubMed
- Kruyer NS, Peralta-Yahya P. 2017. Metabolic engineering strategies to bio-adipic acid production. Curr Opin Biotechnol. 45: 136–143. ArticlePubMed
- Lackner M, Mukherjee A, Koller M. 2023. What are “Bioplastics”? Defining renewability, biosynthesis, biodegradability, and biocompatibility. Polymers. 15(24): 4695.ArticlePubMedPMC
- Lajus S, Dusséaux S, Verbeke J, Rigouin C, Guo Z, et al. 2020. Engineering the yeast Yarrowia lipolytica for production of polylactic acid homopolymer. Front Bioeng Biotechnol. 8: 954.ArticlePubMedPMC
- Larroude M, Rossignol T, Nicaud JM, Ledesma-Amaro R. 2018. Synthetic biology tools for engineering Yarrowia lipolytica. Biotechnol Adv. 36(8): 2150–2164. ArticlePubMedPMC
- Leaf TA, Peterson MS, Stoup SK, Somers D, Srienc F. 1996. Saccharomyces cerevisiae expressing bacterial polyhydroxybutyrate synthase produces poly-3-hydroxybutyrate. Microbiology. 142(5): 1169–1180. ArticlePubMed
- Leavitt JM, Wagner JM, Tu CC, Tong A, Liu Y, et al. 2017. Biosensor-enabled directed evolution to improve muconic acid production in Saccharomyces cerevisiae. Biotechnol J. 12(10): 1600687.ArticlePubMedLink
- Ledesma-Amaro R, Nicaud JM. 2016. Yarrowia lipolytica as a biotechnological chassis to produce usual and unusual fatty acids. Prog Lipid Res. 61: 40–50. ArticlePubMed
- Lee JY, Kang CD, Lee SH, Park YK, Cho KM. 2015. Engineering cellular redox balance in Saccharomyces cerevisiae for improved production of L-lactic acid. Biotechnol Bioeng. 112(4): 751–758. ArticlePubMed
- Lee SM, Lee JY, Hahn JS, Baek SH. 2024. Engineering of Yarrowia lipolytica as a platform strain for producing adipic acid from renewable resource. Bioresour Technol. 391: 129920.ArticlePubMed
- Lee JW, Na D, Park JM, Lee J, Choi S, et al. 2012. Systems metabolic engineering of microorganisms for natural and non-natural chemicals. Nat. Chem. Biol. 8(6): 536–546. ArticlePubMedPDF
- Lian J, Mishra S, Zhao H. 2018. Recent advances in metabolic engineering of Saccharomyces cerevisiae: new tools and their applications. Metab. Eng. 50: 85–108. ArticlePubMed
- Liao X, Ma H, Tang YJ. 2022. Artificial intelligence: a solution to involution of design-build-test-learn cycle. Curr Opin Biotechnol. 75: 102712.ArticlePubMed
- Liu T, Peng B, Huang S, Geng A. 2020. Recombinant xylose-fermenting yeast construction for the co-production of ethanol and cis,cis-muconic acid from lignocellulosic biomass. Bioresour Technol Rep. 9: 100395.Article
- Liu H, Wei L, Ba L, Yuan Q, Liu Y. 2022. Biopolymer production in microbiology by application of metabolic engineering. Polym Bull. 79(8): 5773–5794. ArticlePDF
- Marshall S, Payne K, Leys D. 2017. The UbiX-UbiD system: The biosynthesis and use of prenylated flavin (prFMN). Arch Biochem Biophys. 632: 209–221. ArticlePubMed
- Masutani K, Kimura Y. 2018. Present situation and future perspectives of poly (lactic acid). In Di Lorenzo ML, Androsch R. (eds.), Synthesis, Structure and Properties of Poly(lactic acid), pp. 1–25. Springer International Publishing.
- Matsumoto K, Takase K, Aoki E, Doi Y, Taguchi S. 2005. Synergistic effects of Glu130Asp substitution in the type II polyhydroxyalkanoate (PHA) synthase: enhancement of PHA production and alteration of polymer molecular weight. Biomacromolecules. 6(1): 99–104. ArticlePubMed
- McKenna R, Nielsen DR. 2011. Styrene biosynthesis from glucose by engineered E. coli. Metab Eng. 13(5): 544–554. ArticlePubMed
- Mehmood A, Raina N, Phakeenuya V, Wonganu B, Cheenkachorn K. 2023. The current status and market trend of polylactic acid as biopolymer: Awareness and needs for sustainable development. Mater Today Proc. 72(6): 3049–3055. Article
- Mitsui R, Yamada R, Matsumoto T, Yoshihara S, Tokumoto H, et al. 2020. Construction of lactic acid-tolerant Saccharomyces cerevisiae by using CRISPR-Cas-mediated genome evolution for efficient D-lactic acid production. Appl Microbiol Biotechnol. 104(21): 9147–9158. ArticlePubMedPDF
- Moshood TD, Nawanir G, Mahmud F, Mohamad F, Ahmad MH, et al. 2022. Sustainability of biodegradable plastics: New problem or solution to solve the global plastic pollution? CRGSC. 5: 100273.Article
- Muñoz de Las Heras A, Portugal-Nunes DJ, Rizza N, Sandström AG, Gorwa-Grauslund MF. 2016. Anaerobic poly-3-D-hydroxybutyrate production from xylose in recombinant Saccharomyces cerevisiae using a NADH-dependent acetoacetyl-CoA reductase. Microb Cell Fact. 15: 197.ArticlePubMedPMC
- Murakami S, Shimamoto T, Nagano H, Tsuruno M, Okuhara H, et al. 2015. Producing human ceramide-NS by metabolic engineering using yeast Saccharomyces cerevisiae. Sci Rep. 5(1): 16319.ArticlePubMedPMCPDF
- Na D, Kim TY, Lee SY. 2010. Construction and optimization of synthetic pathways in metabolic engineering. Curr Opin Microbiol. 13(3): 363–370. ArticlePubMed
- Narancic T, Cerrone F, Beagan N, O'Connor KE. 2020. Recent advances in bioplastics: Application and biodegradation. Polymers. 12(4): 920.ArticlePubMedPMC
- Narancic T, Verstichel S, Reddy Chaganti S, Morales-Gamez L, Kenny ST, et al. 2018. Biodegradable plastic blends create new possibilities for end-of-life management of plastics but they are not a panacea for plastic pollution. Environ Sci Technol. 52(18): 10441–10452. ArticlePubMed
- Nicolai T, Deparis Q, Foulquie-Moreno MR, Thevelein JM. 2021. In-situ muconic acid extraction reveals sugar consumption bottleneck in a xylose-utilizing Saccharomyces cerevisiae strain. Microb Cell Fact. 20(1): 114.ArticlePubMedPMC
- Ning Y, Liu H, Zhang R, Jin Y, Yu Y, et al. 2022. Research progress on the construction of artificial pathways for the biosynthesis of adipic acid by engineered microbes. Fermentation. 8(8): 393.Article
- Niu W, Draths KM, Frost JW. 2002. Benzene-free synthesis of adipic acid. Biotechnol Prog. 18(2): 201–211. ArticlePubMed
- O’Brien CE, Oliveira-Pacheco J, Ó Cinnéide E, Haase MAB, Hittinger CT, et al. 2021. Population genomics of the pathogenic yeast Candida tropicalis identifies hybrid isolates in environmental samples. PLoS Pathog. 17(3): e1009138. ArticlePubMedPMC
- Ojo AO, De Smidt O. 2023. Lactic acid: A comprehensive review of production to purification. Processes. 11(3): 688.Article
- Osawa F, Fujii T, Nishida T, Tada N, Ohnishi T, et al. 2009. Efficient production of L-lactic acid by Crabtree-negative yeast Candida boidinii. Yeast. 26(9): 485–496. ArticlePubMed
- Park HJ, Bae JH, Ko HJ, Lee SH, Sung BH, et al. 2018. Low-pH production of D-lactic acid using newly isolated acid-tolerant yeast Pichia kudriavzevii NG7. Biotechnol Bioeng. 115(9): 2232–2242. ArticlePubMed
- Park YK, Ledesma-Amaro R. 2023. What makes Yarrowia lipolytica well suited for industry? Trends Biotechnol. 41(2): 242–254. ArticlePubMed
- Perera IC, Abeywickrama TD, Rahman FA. 2023. Role of genetically engineered yeast in plastic degradation. In Daverey A, Dutta K, Joshi S, Gea T. (eds.), Advances in yeast biotechnology for biofuels and sustainability, pp. 567–584, Elsevier.
- Polen T, Spelberg M, Bott M. 2013. Toward biotechnological production of adipic acid and precursors from biorenewables. J Biotechnol. 167(2): 75–84. ArticlePubMed
- Pollet E, Avérous L. 2011. Production, chemistry and properties of polyhydroxyalkanoates. In Plackett D. (ed.), Biopolymers-New Materials for Sustainable Films and Coatings, pp. 65–86.
- Portugal-Nunes DJ, Pawar SS, Lidén G, Gorwa-Grauslund MF. 2017. Effect of nitrogen availability on the poly-3-D-hydroxybutyrate accumulation by engineered Saccharomyces cerevisiae. AMB Express. 7: 1–12. ArticlePubMedPMCPDF
- Pyne ME, Bagley JA, Narcross L, Kevvai K, Exley K, et al. 2023. Screening non-conventional yeasts for acid tolerance and engineering Pichia occidentalis for production of muconic acid. Nat Commun. 14(1): 5294.ArticlePubMedPMCPDF
- Pyne ME, Narcross L, Melgar M, Kevvai K, Mookerjee S, et al. 2018. An engineered Aro1 protein degradation approach for increased cis, cis-muconic acid biosynthesis in Saccharomyces cerevisiae. Appl Environ Microbiol. 84(17): e01095–01018. ArticlePubMedPMCLink
- Raab AM, Gebhardt G, Bolotina N, Weuster-Botz D, Lang C. 2010. Metabolic engineering of Saccharomyces cerevisiae for the biotechnological production of succinic acid. Metab Eng. 12(6): 518–525. ArticlePubMed
- Raj K, Partow S, Correia K, Khusnutdinova AN, Yakunin AF, et al. 2018. Biocatalytic production of adipic acid from glucose using engineered Saccharomyces cerevisiae. Metab Eng Commun. 6: 28–32. ArticlePubMedPMC
- Rawls RL. 2000. 'Synthetic biology' makes its debut. Chem Eng News. 78(17): 49–53. Article
- Rigouin C, Lajus S, Ocando C, Borsenberger V, Nicaud JM, et al. 2019. Production and characterization of two medium-chain-length polyhydroxyalkanoates by engineered strains of Yarrowia lipolytica. Microb Cell Fact. 18(1): 99.ArticlePubMedPMC
- Rodrigues A, Maia D, Fernandes F. 2015. Production of lactic acid from glycerol by applying an alkaline hydrothermal process using homogeneous catalysts and high glycerol concentration. Braz J Chem Eng. 32(3): 749–755. Article
- Runguphan W, Keasling JD. 2014. Metabolic engineering of Saccharomyces cerevisiae for production of fatty acid-derived biofuels and chemicals. Metab Eng. 21: 103–113. ArticlePubMed
- Saharan BS, Grewal A, Kumar P. 2014. Biotechnological production of polyhydroxyalkanoates: A review on trends and latest developments. Chin J Biol. 2014: 1–18. ArticlePDF
- Saitoh S, Ishida N, Onishi T, Tokuhiro K, Nagamori E, et al. 2005. Genetically engineered wine yeast produces a high concentration of L-lactic acid of extremely high optical purity. Appl Environ Microbiol. 71(5): 2789–2792. ArticlePubMedPMCLink
- Sandström AG, Muñoz de las Heras A, Portugal-Nunes DJ, Gorwa-Grauslund MF. 2015. Engineering of Saccharomyces cerevisiae for the production of poly-3-D-hydroxybutyrate from xylose. AMB Express. 5: 1–9. ArticlePubMedPMC
- Shi M, Li M, Yang A, Miao X, Yang L, et al. 2022. Class I polyhydroxyalkanoate (PHA) synthase increased polylactic acid production in engineered Escherichia coli. Front Bioeng Biotechnol. 10: 919969.ArticlePubMedPMC
- Skoog E, Shin JH, Saez-Jimenez V, Mapelli V, Olsson L. 2018. Biobased adipic acid - The challenge of developing the production host. Biotechnol Adv. 36(8): 2248–2263. ArticlePubMed
- Song JY, Park JS, Kang CD, Cho HY, Yang D, et al. 2016. Introduction of a bacterial acetyl-CoA synthesis pathway improves lactic acid production in Saccharomyces cerevisiae. Metab Eng. 35: 38–45. ArticlePubMed
- Suástegui M, Yu Ng C, Chowdhury A, Sun W, Cao M, et al. 2017. Multilevel engineering of the upstream module of aromatic amino acid biosynthesis in Saccharomyces cerevisiae for high production of polymer and drug precursors. Metab Eng. 42: 134–144. ArticlePubMed
- Subramanian B, Basak S, Thirumurugan R, Saleena LM. 2024. Metabolic flux analysis and metabolic engineering for polyhydroxybutyrate (PHB) production. Polym Bull. 81: 10589–10608. ArticlePDF
- Tejayadi S, Cheryan M. 1995. Lactic acid from cheese whey permeate. Productivity and economics of a continuous membrane bioreactor. Appl Microbiol Biotechnol. 43: 242–248. ArticlePDF
- Tönjes S, Uitterhaegen E, Palmans I, Ibach B, De Winter K, et al. 2024. Metabolic engineering and process intensification for muconic acid production using Saccharomyces cerevisiae. Int J Mol Sci. 25(19): 10245.ArticlePubMedPMC
- Tran PHN, Jung JH, Ko JK, Gong G, Um Y, et al. 2023. Co-production of ethanol and polyhydroxybutyrate from lignocellulosic biomass using an engineered Saccharomyces cerevisiae. Renew Energy. 212: 601–611. Article
- Tsuge Y, Kato N, Yamamoto S, Suda M, Jojima T, et al. 2019. Metabolic engineering of Corynebacterium glutamicum for hyperproduction of polymer-grade L- and D-lactic acid. Appl Microbiol Biotechnol. 103(8): 3381–3391. ArticlePubMedPDF
- Wang F, Lee SY. 1997. Poly (3-hydroxybutyrate) production with high productivity and high polymer content by a fed-batch culture of Alcaligenes latus under nitrogen limitation. Appl Environ Microbiol. 63(9): 3703–3706. ArticlePubMedPMCLink
- Wang G, Ozmerih S, Guerreiro R, Meireles AC, Carolas A, et al. 2020. Improvement of cis,cis-muconic acid production in Saccharomyces cerevisiae through biosensor-aided genome engineering. ACS Synth Biol. 9(3): 634–646. ArticlePubMedPMCLink
- Wang G, Tavares A, Schmitz S, Franca L, Almeida H, et al. 2022. An integrated yeast-based process for cis,cis-muconic acid production. Biotechnol Bioeng. 119(2): 376–387. ArticlePubMedPMCLink
- Watcharawipas A, Sae-Tang K, Sansatchanon K, Sudying P, Boonchoo K, et al. 2021. Systematic engineering of Saccharomyces cerevisiae for D-lactic acid production with near theoretical yield. FEMS Yeast Res. 21(4): foab024.ArticlePubMedPDF
- Wu S, Chen W, Lu S, Zhang H, Yin L. 2022. Metabolic engineering of shikimic acid biosynthesis pathway for the production of shikimic acid and its branched products in microorganisms: Advances and prospects. Molecules. 27(15): 4779.ArticlePubMedPMC
- Wu X, Ren Y, Chen S, Cai P, Zhou YJ. 2025. Production of L-lactic acid from methanol by engineered yeast Pichia pastoris. Bioresour Technol. 415: 131730.ArticlePubMed
- Xu W. 2023. A study on the synthesis, modification and current market status of PBAT. E3S Web Conf. 385: 04007.Article
- Xu N, Gao H, Wang Y, Liu C, Hu L, et al. 2025. Recent advances in bio-based production of organic acids by genetically engineered yeasts. Biochem Eng J. 215: 109587.Article
- Xue Z, Sharpe PL, Hong SP, Yadav NS, Xie D, et al. 2013. Production of omega-3 eicosapentaenoic acid by metabolic engineering of Yarrowia lipolytica. Nat Biotechnol. 31(8): 734–740. ArticlePubMedPDF
- Yamada R, Ogura K, Kimoto Y, Ogino H. 2019. Toward the construction of a technology platform for chemicals production from methanol: D-lactic acid production from methanol by an engineered yeast Pichia pastoris. World J Microbiol Biotechnol. 35(2): 37.ArticlePubMedPDF
- Yamamoto Y, Yamada R, Matsumoto T, Ogino H. 2023. Construction of a machine-learning model to predict the optimal gene expression level for efficient production of D-lactic acid in yeast. World J Microbiol Biotechnol. 39(3): 69.ArticlePubMedPDF
- Yang H, Lin X, Zhong X, Cao M, Yuan J, et al. 2024. Current status and advances in the green synthesis of muconic acid. Crit Rev Biotechnol. 9: 1–18. ArticlePubMed
- Ylinen A, De Ruijter JC, Jouhten P, Penttilä M. 2022. PHB production from cellobiose with Saccharomyces cerevisiae. Microb Cell Fact. 21(1): 124.ArticlePubMedPMCPDF
- Ylinen A, Maaheimo H, Anghelescu-Hakala A, Penttilä M, Salusjärvi L, et al. 2021. Production of D-lactic acid containing polyhydroxyalkanoate polymers in yeast Saccharomyces cerevisiae. J Ind Microbiol Biotechnol. 48(5-6): kuab028.ArticlePubMedPMCPDF
- Zhang B, Li R, Yu L, Wu C, Liu Z, et al. 2023a. L-lactic acid production via sustainable neutralizer-free route by engineering acid-tolerant yeast Pichia kudriavzevii. J Agric Food Chem. 71(29): 11131–11140. ArticlePubMedLink
- Zhang X, Liu Y, Wang J, Zhao Y, Deng Y. 2020. Biosynthesis of adipic acid in metabolically engineered Saccharomyces cerevisiae. J Microbiol. 58(12): 1065–1075. ArticlePubMedPDF
- Zhang FL, Zhang L, Zeng DW, Liao S, Fan Y, et al. 2023b. Engineering yeast cell factories to produce biodegradable plastics and their monomers: current status and prospects. Biotechnol Adv. 68: 108222.ArticlePubMed
- Zhao M, Huang D, Zhang X, Koffas MAG, Zhou J, et al. 2018. Metabolic engineering of Escherichia coli for producing adipic acid through the reverse adipate-degradation pathway. Metab Eng. 47: 254–262. ArticlePubMed
- Zhao J, Li Q, Sun T, Zhu X, Xu H, et al. 2013. Engineering central metabolic modules of Escherichia coli for improving β-carotene production. Metab Eng. 17: 42–50. ArticlePubMed
- Zhao X, Wang Y, Chen X, Yu X, Li W, et al. 2023. Sustainable bioplastics derived from renewable natural resources for food packaging. Matter. 6(1): 97–127. Article
- Zheng X, Cairns TC, Ni X, Zhang L, Zhai H, et al. 2022. Comprehensively dissecting the hub regulation of PkaC on high-productivity and pellet macromorphology in citric acid producing Aspergillus niger. Microb Biotechnol. 15(6): 1867–1882. ArticlePubMedPMCLink
- Zhou K. 2022. Engineering microbes to synthesize functionalized biopolymers. J Mater Chem B. 10(37): 7132–7135. ArticlePubMed
- Zhu P, Luo R, Li Y, Chen X. 2022. Metabolic engineering and adaptive evolution for efficient production of L-lactic acid in Saccharomyces cerevisiae. Microbiol Spectr. 10(6): e0227722. ArticlePubMed
References
Supplementary Information
References
Citations

- Reprogramming of Saccharomyces cerevisiae for sustainable cis, cis-muconic acid production from lignocellulosic biomass
Huong-Giang Le, Ja-Kyong Ko, Sun-Mi Lee
Biotechnology and Bioprocess Engineering.2026;[Epub] CrossRef - Enzymatic and microbial routes to bioplastics: The green chemistry frontier of biopolymers
Giovanni Gallo, Emma Piccoli, Luca Bombardi, Martina Aulitto, Salvatore Fusco
FEBS Open Bio.2026; 16(4): 709. CrossRef - From organic wastes to value: yeast-based bioconversion of waste-derived feedstocks into valuable compounds
Ticiana Fernandes, Maria João Sousa, Ricardo Franco-Duarte
Food Bioscience.2026; 79: 108871. CrossRef - Advancing microbial engineering through synthetic biology
Ki Jun Jeong
Journal of Microbiology.2025; 63(3): e2503100. CrossRef - Biorefinery-based production of biodegradable bioplastics: advances and challenges in circular bioeconomy
Ariane Fátima Murawski de Mello, Clara Matte Borges Machado, Lucia Carolina Ramos Neyra, Diego Yamir Ocán-Torres, Rafael Novaes Barros, Mariana Camargo Medeiros, Carlos Ricardo Soccol, Luciana Porto de Souza Vandenberghe
npj Materials Sustainability.2025;[Epub] CrossRef

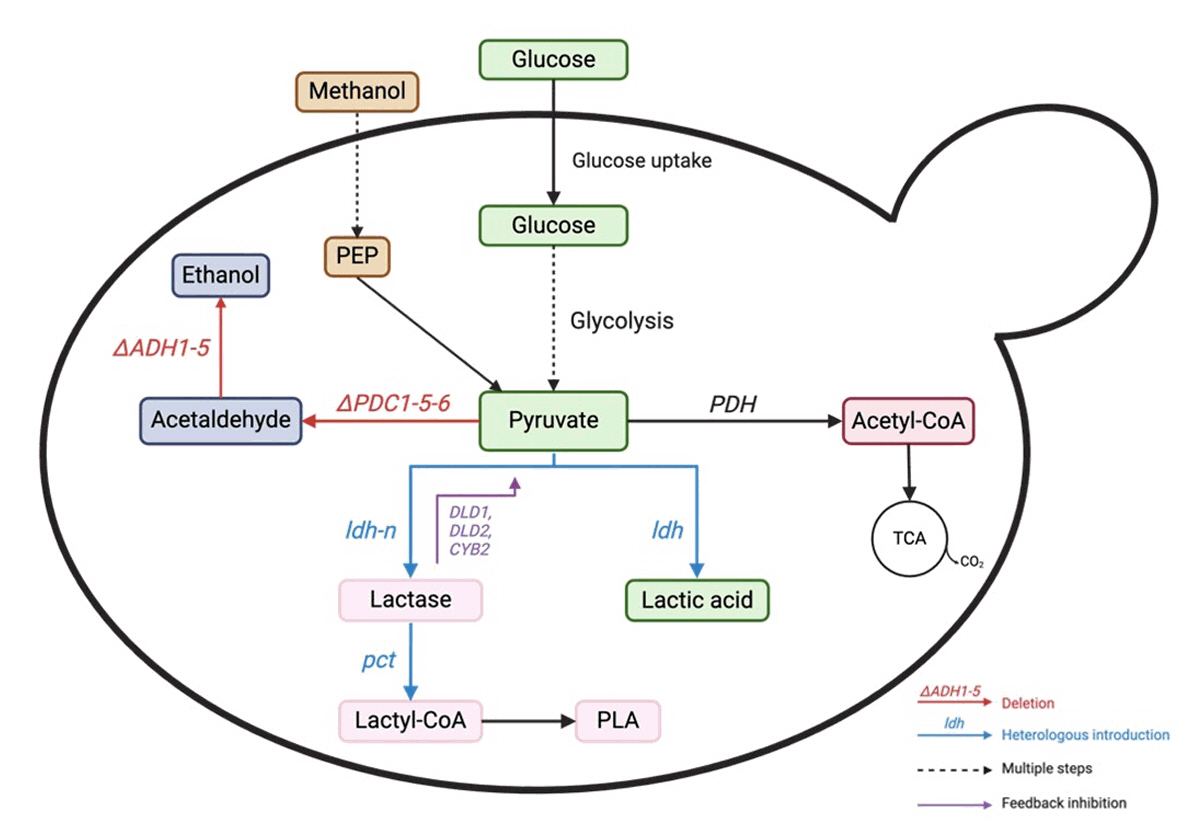
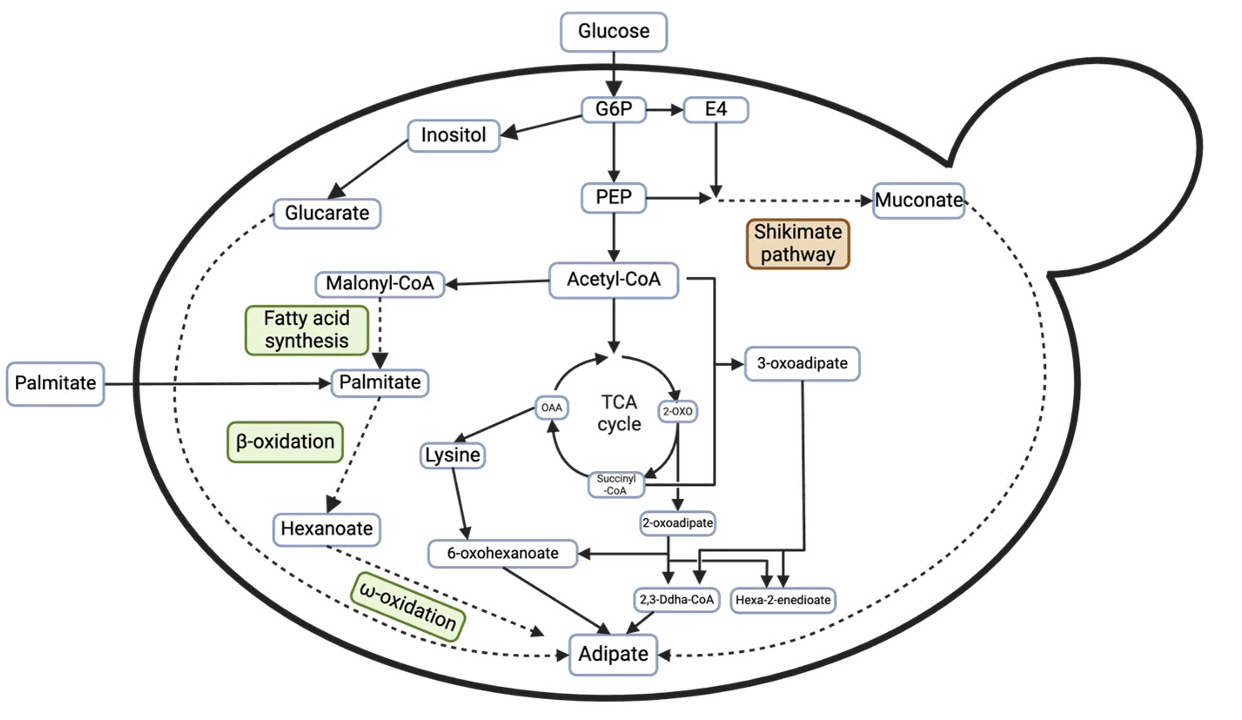
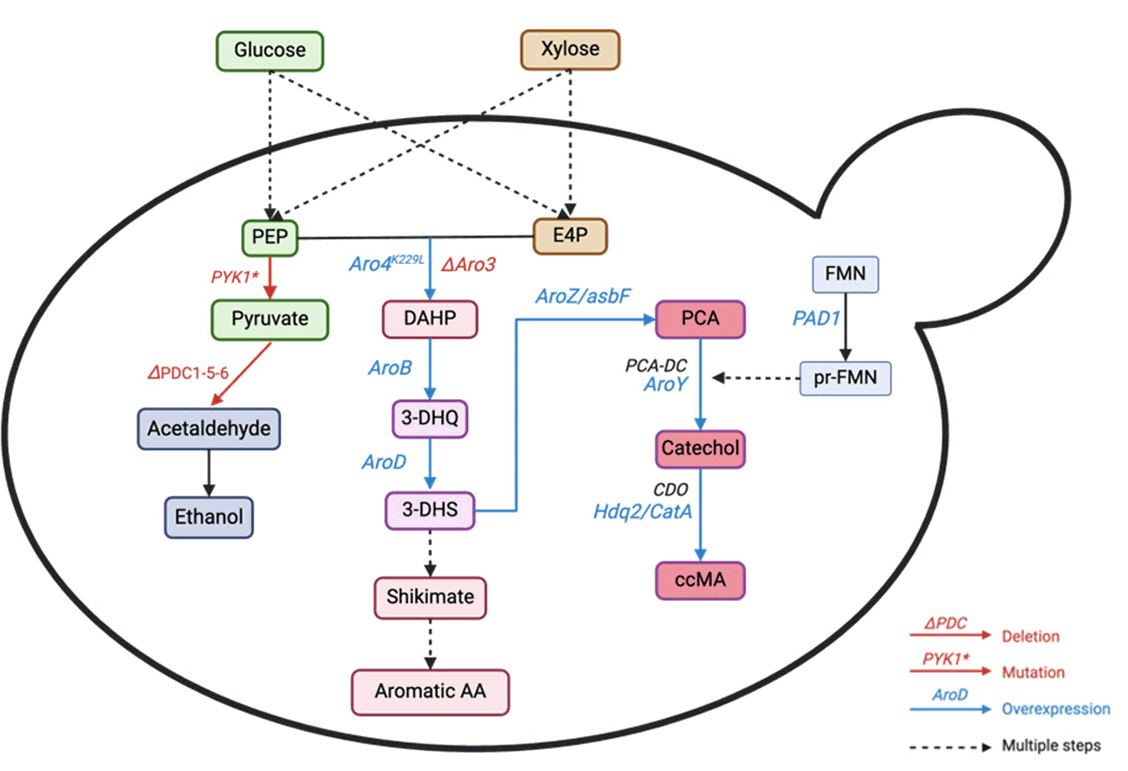
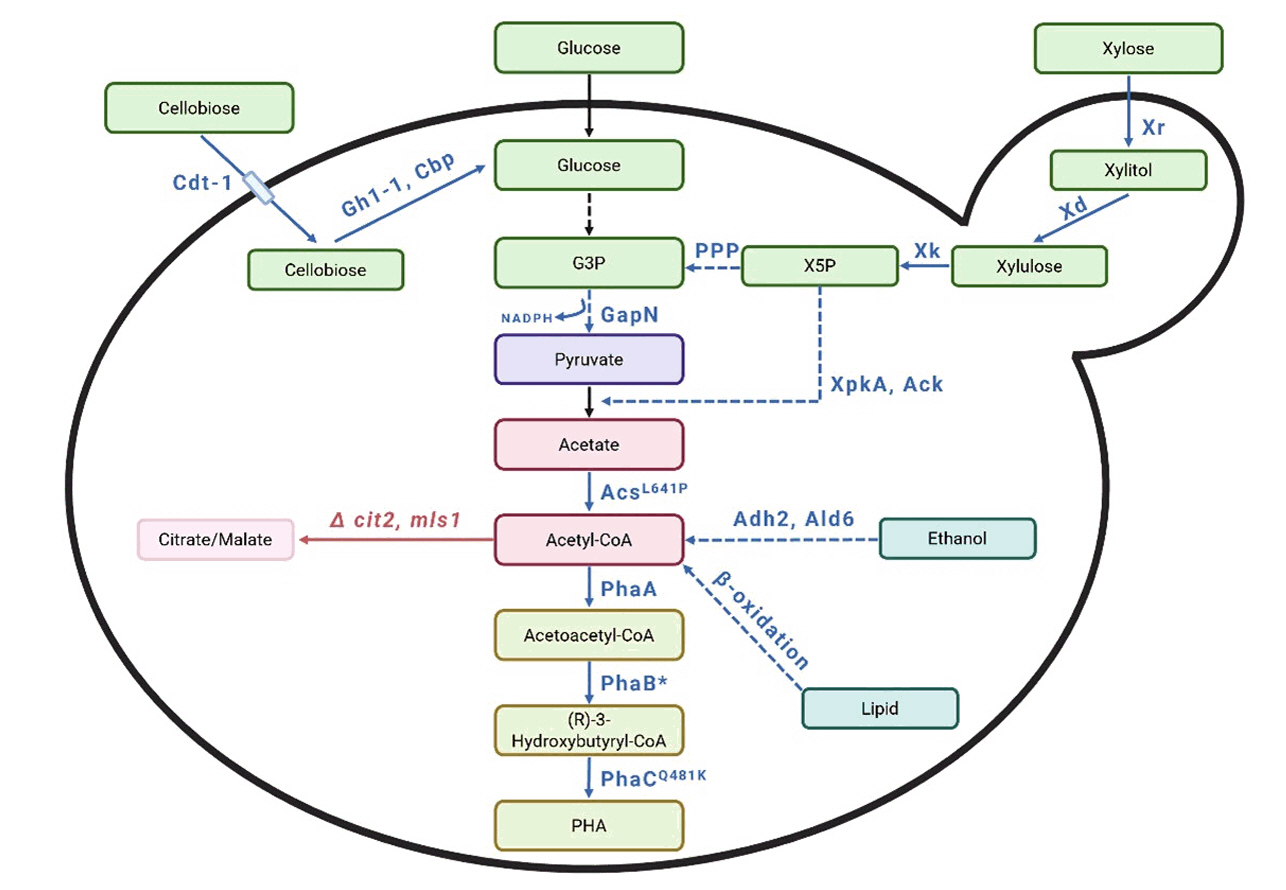
Fig. 1.
Fig. 2.
Fig. 3.
Fig. 4.
Fig. 5.
Fig. 6.
Fig. 7.
| Strains | Substrate | Engineering Strategies | Tools | Culture condition | Titer (g/L) | Productivity (g/L/h) | References |
|---|---|---|---|---|---|---|---|
| Saccharomyces cerevisiae SP1130 | Glucose | Express heterologous lactate dehydrogenase (LDH) genes, attenuating several key pathway genes including glycerol-3-phosphate dehydrogenase 1 (GPD1) and cytochrome-c oxidoreductase 2 (CYB2). Knocking out the pyruvate decarboxylase 1 (PDC1) and alcohol dehydrogenase 1 (ADH1) to disrupt the ethanol production pathway, redirect metabolic flux towards lactic acid production. Introduce an alternative pathway to produce acetyl-CoA using the acetylating acetaldehyde dehydrogenase (A-ALD) gene from E. coli to compensate for the attenuated ADH1 | Heterologous gene expression, Introduction of a synthetic pathway | Fed-batch fermentation | 142 | 3.55 | |
| S. cerevisiae T165R | Cane juice | Integrate six copies of L-LDH into the genome of host cell | Multi-integration of a heterologous gene | 1-L jar fermenter | L-LA 122 | - | |
| S. cerevisiae S.c-NO.2-100 | Glucose | Rewire the biosynthetic pathway for LA production by screening and introducing L-LDH, reducing ethanol accumulation by deleting PDC1, ADH1, introducing bacterial acetyl-CoA synthesis pathway. Strain evolution for acid tolerance by adaptive evolution. Delete NADH-consuming enzymes (NDE1/2) to enhance the cofactor availability. Overexpress bacterial transporters JEN1 and remodel the cell membrane by deleting SAM2 for efficient LA export | Heterologous gene expression, Introduction of heterologous transporter, ALE | 5-L batch bioreactor | L-LA 121.5 | 1.69 | |
| S. cerevisiae BK01 | Acetate-rich buckwheat husk hydrolysates | Adaptive laboratory evolution (ALE) on 8% lactic acid on S. cerevisiae SR8LDH strain which was previously constructed by the introduction of LDH gene from Lactobacillus acidophilus ATCC4356 under the control of the PGK1 promoter. The mutant BK01 was characterized by genome sequencing and metabolomic profiling | ALE, metabolomic profiling | 20 mL fermentation | L-LA 119 | 1.24 | |
| S. cerevisiae ASc- d789M | Glucose | Screen D-LDH from bacteria, overexpress LpDLDH, delete glycerol production genes GPD1, GPD2, and lactate dehydrogenase gene DLD1 to minimize by-products, downregulation ADH1 using the L-methionine repressible promoter to minimize the impact on growth. Created an intraspecific hybrid strain by mating the engineered strain with a weak acid-tolerant wild-type strain BCC39850 to improve acid tolerance and D-LA production | Gene knockout by CRISPR/Cas9, CRISPR/Cas12a, yeast mating | Fed-batch fermentation | D-LA 23.41 ± 1.65 | 0.81 | |
| S. cerevisiae | Glucose | Genome evolution applied a method combining GMES and CRISPR-δ integration (GMES/CRISPR) to integrate 13 bacterial genes including HXT7, HXK2, PGI1, PFK1, PFK2, FBA1, TPI1, TDH3, PGK1, GPM1, ENO2, and PYK2, and d-lactate dehydrogenase (d-LDH) into genomic DNA | Genome evolution by GMES and CRISPR-δ integration, overlap extension PCR, transcriptome analysis, real-time PCR | Semi-neutralizing fermentation | D-LA 52.2 | 2.17 | |
| S. cerevisiae JHY5330 | Glucose | Introduce D-lactate dehydrogenase gene (ldhA) from bacteria, reduce D-LA consumption by deleting DLD1 encoding D-lactate dehydrogenase and JEN1 encoding monocarboxylate transporter, eliminate ethanol production gene PDC1 and ADH1, glycerol production genes GPD1 and GPD2, improve strain LA tolerance by ALE and overexpression of HAA1 encoding a transcriptional activator involved in weak acid stress response | ALE, gene knockout, PCR-mediated method using the Cre/loxP system, quantitative reverse transcription PCR (qRT-PCR) | Fed-batch fermentation | D-LA 112 | 2.2 | |
| S. cerevisiae JHY5610 | Glucose | Employed ALE for LA-tolerance of host strain, coupled with introduction of LA production gene Lm. ldhA, and deletion of ethanol and glycerol production genes (ADH, GPD) and D-LA degradation genes to redirect metabolic flux toward LA production | ALE, gene knockout, whole genome sequencing | Fed-batch fermentation | D-LA 82.6 | 1.5 | |
| S. cerevisiae | Glucose | Rewire metabolic fluxes toward the production of L-lactic acid by deleting PDC1, CYB2, GPD1, and replacing LDH. Re-engineer the intracellular redox balance by deleting NDE1 and NDE2 encoding isoenzymes of the external NADH dehydrogenase | Gene knockout, re-engineer intracellular redox | Fed-batch | L-LA 117 | - | |
| Pichia pastoris | Methanol | Screen LDH genes, co-utilize LDH cofactors, knockout DLD1, DLD2 and CYB2 to block LA consumption pathway, and construct a coordinated dual pathway in cytoplasm and mitochondria | Site-directed mutation, CRISPR-Cas9, fluorescence microscopy analysis | Fed-batch fermentation | L-LA 4.2 | 0.0126 | |
| Pichia kudriavzevii NG7 | Glucose | Replace the pyruvate decarboxylase 1 gene (PDC1) with the d-lactate dehydrogenase gene (d-LDH) derived from Lactobacillus plantarum, improve strain tolerance by ALE | ALE, whole genome sequencing | Bioreactor fed-batch | D-LA 154 | 4.16 | |
| Candida boidinii | Glucose | Disrupted PDC1 to reduce ethanol production, introduce and express Bovine L-LDH gene under the PDC1 promoter and optimize fermentation condition | Heterologous gene expression | Batch fermentation | L-LA 85.9 | 1.79 | |
| S. cerevisiae | Glucose | Poly (D-lactic acid) (PDLA) and copolymer P (LA-3HB) were produced by express PHA synthase PhaC1437Ps6-19, propionyl-CoA transferase Pct540Cp, acetyl-CoA acetyltransferase PhaA, and acetoacetyl-CoA reductase PhaB1 | Gibson Assembly, modular cloning method (MoClo), GC-MS, NMR analysis | Erlenmeyer baffled flasks fermentation | PDLA 0.73% (CDW) | - | |
| Yarrowia lipolytica | Glucose, racemic lactic acid | Disrupt lactic acid consumption pathway, introduce heterologous pathway for PDLA production containing propionyl-CoA transferase (pct) from Clostridium propionicum to convert lactic acid into lactyl-CoA, and an evolved polyhydroxyalkanoic acid PHA synthase polymerizing lactyl-CoA into PDLA | Site-directed mutagenesis, UHPLC-FTMS | Erlenmeyer baffled flasks fermentation | PDLA 26 mg/g-DCW | - |
| Strains | Substrate | Engineering Strategies | Tools | Culture condition | Titer | References |
|---|---|---|---|---|---|---|
| E. coli BL21 (DE3) | Glycerol | Reconstruct five-step reverse adipate-degradation pathway (RADP) from Thermobifida fusca in E. coli, then overexpress 5-Carboxy-2-pentenoyl-CoA reductase (Tfu_1647) - the gene coding for the rate-limiting step in the RADP, delete succinate-CoA ligase gene (sucD) to eliminate competitive | Introduction of a synthetic pathway, Gibson assembly, CRISPR/Cas9, gene overexpression, SDS-PAGE | Fed-batch fermentation in 5-L bioreactor | 68.0 g/L | |
| Candida spp. | Fatty acid | Rewire the ω- then the β-oxidation pathway | - | Two-stage fed-batch process | 50 g/L | |
| Candida tropicalis | C12 methyl laurate | Create a mutant strain of C. tropicali by deleting AOX genes encoding acyl-CoA oxidases to produce AA via ω- and the β-oxidation pathway | Gene deletion | Bioreactor, fed-batch | 12.1 g/L | |
| S. cerevisiae | Glucose | Use the reverse adipate degradation pathway (RADP) from Thermobifida fusca, |
Introduction of a synthetic pathway, Gibson assembly, overlap extension PCR | Fed-batch bioreactor | 10.09 mg/L | |
| S. cerevisiae | Glucose | Rewire the cis, cis-muconic acid pathway by expressing enoate reductases (ERs) from Bacillus coagulans to convert ccMA into AA | Introduction of a synthetic pathway, Gibson assembly | Three stage fermentations | 2.59 mg/L, ccMA > 284 mg/L | |
| Y. lipolytica | Fatty acid methyl esters | Regulate the β-oxidation pathway by disrupting acyl-CoA oxidases (POX1 and POX3) and enhance ω-oxidation through overexpression of ALK5, CPR1, and FAO1, two-phase bioprocess to enhance AA production | Gene deletion, two-phase bioprocess | 20 ml in 250 ml baffled flask | 1176.2 mg/L |
| Strains | Substrate | Engineering Strategies | Tools | Culture condition | Titer (g/L) | Productivity (g/L/h) | References |
|---|---|---|---|---|---|---|---|
| S. cerevisiae | Glucose | Employed ccMA-biosensor, coupled with GFP expression to screen UV-mutagenesis libraries for ccMA producing yeast, then overexpress genes encoding for PCA decarboxylase and AROM protein and restore URA3 prototrophy. | UV-mutagenesis, fluorescence activated cell sorting (FACS), heterologous gene expression | Fed-batch | 20.8 | 0.139 | |
| S. cerevisiae | Glucose | Rewiring the shikimate pathway flux and enhancing the phosphoenolpyruvate supply by mutating PYK1 (A336S) to limit PEP-to-pyruvate conversion and overexpressing PYC1 and PCK1 to redirect pyruvate to PEP, overexpress DAHP synthase (ScARO4K229L), 3-dehydroquinate synthase (EcaroB), and 3-dehydroquinase | Cas9-assisted approach, Gibson assembly, USER cloning, EasyClone method | 2L fermenter | 22.5 | 0.19 | |
| S. cerevisiae | Glucose, xylose, mixture | Eliminated feedback inhibition in the shikimate pathway, insert heterologous pathway for MA production using 3-dehydroshikimate (DHS), protocatechuic acid decarboxylase (PCAD) and oxygen-consuming catechol 1,2-dioxygenase (CDO), delete PDC to eliminate ethanol production, minimized PCA production by enhancing PCAD overexpression and production of its co-factor | Introduction of a synthetic pathway, genome editing using CRISPR/Cas9 | 300 ml shake flasks with 40 mL YP medium | 4.5 | - | |
| S. cerevisiae | Glucose and xylose | Eliminates C2 dependency and ethanol production by ALE and deleting PDC1, PDC5, and PDC6, improving MA tolerance for host strain, promoting growth and production of target product by internal deletion of MTH1, MTH1ΔM41-T78, overexpressing PAD1 as well as reduces side production of intermediate protocatechuic acid (PCA) | Introduction of a synthetic pathway, enzyme engineering, ALE | Fed-batch | 9.3 | 0.100 | |
| S. cerevisiae | Glucose | Combine adaptive laboratory evolution (ALE) and rational metabolic engineering, improve flux by truncation of ARO1 and overexpress an endogenous aromatic decarboxylase | ALE, Gibson assembly | Fed-batch bioreactor | 2.1 | - | |
| S. cerevisiae | Glucose | Employ the computational tool YEASTRACT for predicting novel transcriptional repressors and OptForce strain-design for identifying non-intuitive pathway interventions, strating from glycolytic and pentose phosphate pathway pathway, follow by engineer ARO2, ARO3, ARO4, and the pentafunctional ARO, delete ric1 to increase transcription of ARO2, ARO3, ARO4 | Computational tool for predicting engineering target, Introduction of a synthetic pathway Gibson assembly | - | 0.32 | - | |
| S. cerevisiae | Glucose, xylose (supplement 1 g/L catechol) | Overexpress xylose isomerase gene from Bacteroides valgutus and pentose phosphate pathway genes from S. cerevisiae, then imported a three step ccMA production pathway from E. coli with AroZ-Neu, AroY-Com, and CatA-Cup genes, further overexpression of gene Aro1 (with a stop codon of AroE) and feedback-resistant Aro4opt mutant gene, ALE to improve xylose fermentation and ccMA | Introduction of a synthetic pathway, Enzyme engineering, ALE | Batch, shake flask | 0.424 (1.286 with catechol supplemented) | - | |
| S. cerevisiae | Glucose, supplement amino acids | Balance MA pathway performance with aromatic amino acid prototrophy by destabilizing Aro1, delete ARO4 and ARO3, introduce AroZ, AroY, Hqd2 CatA, Pad1. | CRISPR-mediated homology-directed repair (HDR) | Fed-batch bioreactor | 5.1 | - | |
| S. cerevisiae | Glucose | Construct cassettes using USER cloning-ligation-PCR, express a three-step heterologous pathway using KpAroY.B, KpAroY.Ciso, CaCatA, engineering TKL1, ZWF1, Aro1AroEΔ, Aro4K229L | Introduction of a synthetic pathway, CRISPR/Cas9, in vivo recombination USER assembly and RNA interference | Batch, shake flask | 0.8 | - | |
| Pichia occidentalis | Glucose | Screen host strain with high tolerance to MA, develop genome editing toolkit to engineer the non-conventional yeast and introduce the heterologous pathway for ccMA production | CRISPR-Cas9, antibiotic marker recycling, high-throughput screening | Fed-batch | 38.8 | 0.511 |
| Strains | Substrate | Engineering Strategies | Tools | Culture condition | Titer (mg/L) | Yield (mg/g) | References |
|---|---|---|---|---|---|---|---|
| E. coli | Glucose, xylose | Knocked out ndh gene. Introduced arabinose/xylose transport proteins from Bacillus subtilis (xylA, xylB, and araE). Incorporated the PHB biosynthesis pathway from C. necator | Modification of ribosome binding sites | Fed-batch culture | PHB 21010 | - | |
| S. cerevisiae | Silver grass, glucose, xylose | Constructed a polycistronic PHB cassette with phaA, phaB, and phaC using 2A peptide sequences. Introduced Cas9-expressing plasmid, gRNA, and polycistronic PHB cassette into S. cerevisiae via electroporation. Integrated the polycistronic PHB cassette into the genome | CRISPR-Cas9 mediated genome editing, 2A peptide sequence integration | Serum bottle, microaerobic | PHB 40.3 | 0.93 mg/g silver grass | |
| S. cerevisiae | Glucose | Expressed XpkA and Ack from Aspergillus nidulans to increase acetyl-CoA levels. Incorporated an acetyl-CoA synthase variant (AcsL641P) from Salmonella enterica. Integrated GapN (NADP⁺-dependent glyceraldehyde-3-phosphate dehydrogenase) from Streptococcus mutans into the genome via the Li-Ac method. Codon optimized all genes for efficient expression | Codon optimization DNA 2.0 and GenScript for synthesizing genes, PTEF1-PPGK1 bidirectional promoter | Shake flask, anaerobic | PHB 180 | 0.03 mg/g glucose | |
| S. cerevisiae | Glucose | Synthesized phaA, phaB, and phaC based on R. eutropha H16. Introduced them into S. cerevisiae in plasmid form using the Li-Ac method. Used the strain described by Chen et al. (2012) to increase acetyl-CoA levels. Deleted cit2 and mls1, overexpressed ALD6 and ADH2. Codon optimized all genes for efficient expression | Codon optimization, PTEF1-PPGK1 bidirectional promoter, DNA 2.0 for synthesize genes | Shake flask, aerobic | PHB 43.11 | 0.13 mg/g glucose | |
| S. cerevisiae | Xylose | Introduced XR and XD genes to enable xylose utilization. PCR-amplified phaA, phaB, and phaC genes from C. necator. Assembled these genes into a single plasmid using In-Fusion cloning. Introduced the plasmid into S. cerevisiae via the Li-Ac method. Codon optimized all genes for efficient expression | In-fusion cloning, Codon optimization, overlap extension PCR | Shake flask, aerobic | PHB 45 | 1.17 mg/g xylose | |
| S. cerevisiae | Xylose | Achieved PHB production under anaerobic conditions using xylose as the carbon source. Replaced PhaB from C. necator with NADH-dependent PhaB from Allochromatium vinosum. Introduced the gene into S. cerevisiae in plasmid form via the Li-Ac method | Codon optimization, heterologous gene expression, promoter and terminator pairing | Bioreactor, anaerobic | PHB 360 | 7 mg/g xylose | |
| S. cerevisiae | Xylose high nitrogen supply | Investigated whether nitrogen limitation in yeast influences PHB production. Introduced Xr and Xd from Scheffersomyces stipitis to enable xylose utilization. Replaced PhaB from C. necator with NADH-dependent PhaB from A. vinosum. Assembled these genes into a single plasmid using In-Fusion cloning. Introduced the plasmid into S. cerevisiae via the Li-Ac method. Codon optimized and PCR-amplified all genes | In-fusion cloning, Codon optimization, heterologous gene expression, promoter and terminator pairing | Shake flask, anaerobic | PHB 730 | 13.8 mg/g xylose | |
| S. cerevisiae | Cellobiose | Integrated phaA, phaB1, and phaC1 into the S. cerevisiae genome using Gibson assembly, Li-Ac method, and CRISPR-Cas9. Inserted Gh1-1 from Neurospora crassa and Cbp from Ruminococcus flavefaciens for cellobiose metabolism using the Modular Cloning (MoClo) approach. Codon optimized all genes for efficient expression | Gibson Assembly, Modular cloningCRISPR-Cas9 mediated genome integration, promoter and terminator pairing | Bioreactor | PHB - | 49.4 mg/g cellobiose | |
| Y. lipolytica | Tridecanoate | Directed carbon flux in Y. lipolytica toward either the β-oxidation pathway or the PHA biosynthesis pathway based on the genotype of POX1-6 genes encoding acyl-CoA oxidase isoenzymes. Downregulated the R-3-hydroxyacyl-CoA dehydrogenase multifunctional enzyme domain in the fatty acid β-oxidation pathway via site-directed mutagenesis, generating the MFE-AG16SBC variant. Deleted an acyltransferase gene and overexpressed enoyl-CoA hydratase 2 to further redirect fatty acids from lipid synthesis to the β-oxidation pathway | site-directed mutagenesis, loxP-Cre recombinase, pGEMT easy cloning | Shake flask | PHA - | 7.3% DCW | |
| Y. lipolytica | Tridecanoate | Directed carbon flux in Y. lipolytica toward either the β-oxidation pathway or the PHA biosynthesis pathway based on the genotype of POX1-6 genes encoding acyl-CoA oxidase isoenzymes. Introduced PhaC from Pseudomonas aeruginosa and modified its C-terminal region to include the 34-amino acid sequence from Brassica napus glyoxysomal isocitrate lyase (icl) for peroxisome targeting | Site-directed mutagenesis, STADEN package for sequence analysis | Shake flask | PHA - | 17.64 mg/g biomass | |
| Y. lipolytica | Methyl laurate | Generated various PhaC variants through site-directed mutagenesis. Compared their production yields of mcl-PHA with different chain lengths | Site-directed mutagenesis | - | PHA - | 28% DCW | |
| Y. lipolytica | Triolein | Directed carbon flux in Y. lipolytica toward either the β-oxidation pathway or the PHA biosynthesis pathway based on the genotype of POX1-6 genes encoding acyl-CoA oxidase isoenzymes. Heterologously expressed PhaC1 from Pseudomonas aeruginosa PAO1 with a peroxisomal signal. Introduced the gene into S. cerevisiae in plasmid form via the Li-Ac method. Codon optimized all genes for efficient expression | Codon optimization | Shake flask | PHA 1110 | 5.0% DCW |
Table 1.
Table 2.
Table 3.
Table 4.
TOP
 MSK
MSK
 ePub Link
ePub Link Cite this Article
Cite this Article








