- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Volume 63(3); 2025 > Review
-
Review
Synthetic biology strategies for sustainable bioplastic production by yeasts - Huong-Giang Le1,2, Yongjae Lee3, Sun-Mi Lee3,*
-
Journal of Microbiology 2025;63(3):e2501022.
DOI: https://doi.org/10.71150/jm.2501022
Published online: March 28, 2025
1Division of Energy and Environment Technology, University of Science and Technology (UST), Daejeon 34113, Republic of Korea
2Clean Energy Research Center, Korea Institute of Science and Technology (KIST), Seoul 02792, Republic of Korea
3Division of Environmental Science and Ecological Engineering, Korea University, Seoul 02841, Republic of Korea
- *Correspondence Sun-Mi Lee sunmilee@korea.ac.kr
• Received: January 17, 2025 • Revised: February 27, 2025 • Accepted: February 28, 2025
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Supplementary Information
References
Citations
Citations to this article as recorded by 

- Reprogramming of Saccharomyces cerevisiae for sustainable cis, cis-muconic acid production from lignocellulosic biomass
Huong-Giang Le, Ja-Kyong Ko, Sun-Mi Lee
Biotechnology and Bioprocess Engineering.2026;[Epub] CrossRef - Enzymatic and microbial routes to bioplastics: The green chemistry frontier of biopolymers
Giovanni Gallo, Emma Piccoli, Luca Bombardi, Martina Aulitto, Salvatore Fusco
FEBS Open Bio.2026; 16(4): 709. CrossRef - From organic wastes to value: yeast-based bioconversion of waste-derived feedstocks into valuable compounds
Ticiana Fernandes, Maria João Sousa, Ricardo Franco-Duarte
Food Bioscience.2026; 79: 108871. CrossRef - Advancing microbial engineering through synthetic biology
Ki Jun Jeong
Journal of Microbiology.2025; 63(3): e2503100. CrossRef - Biorefinery-based production of biodegradable bioplastics: advances and challenges in circular bioeconomy
Ariane Fátima Murawski de Mello, Clara Matte Borges Machado, Lucia Carolina Ramos Neyra, Diego Yamir Ocán-Torres, Rafael Novaes Barros, Mariana Camargo Medeiros, Carlos Ricardo Soccol, Luciana Porto de Souza Vandenberghe
npj Materials Sustainability.2025;[Epub] CrossRef
Synthetic biology strategies for sustainable bioplastic production by yeasts
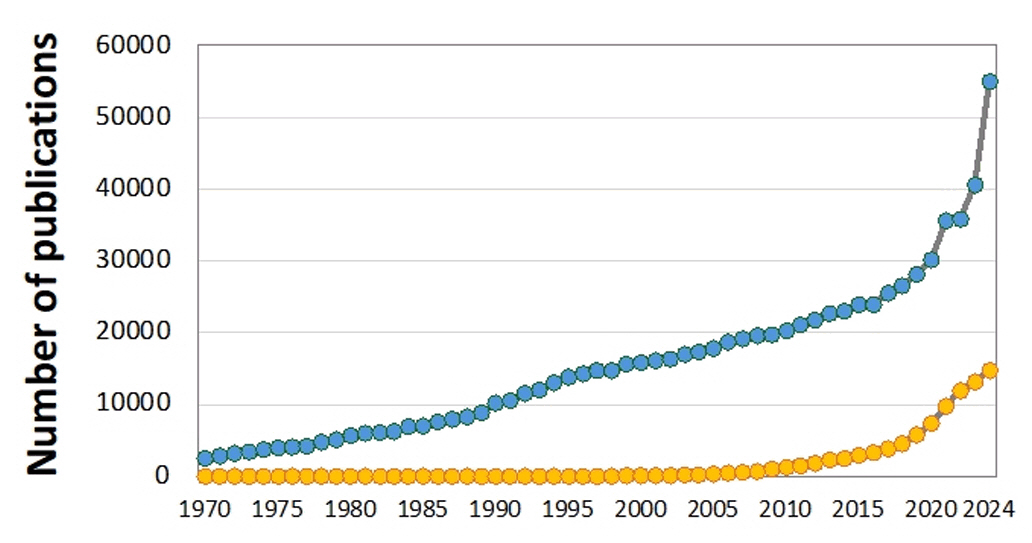

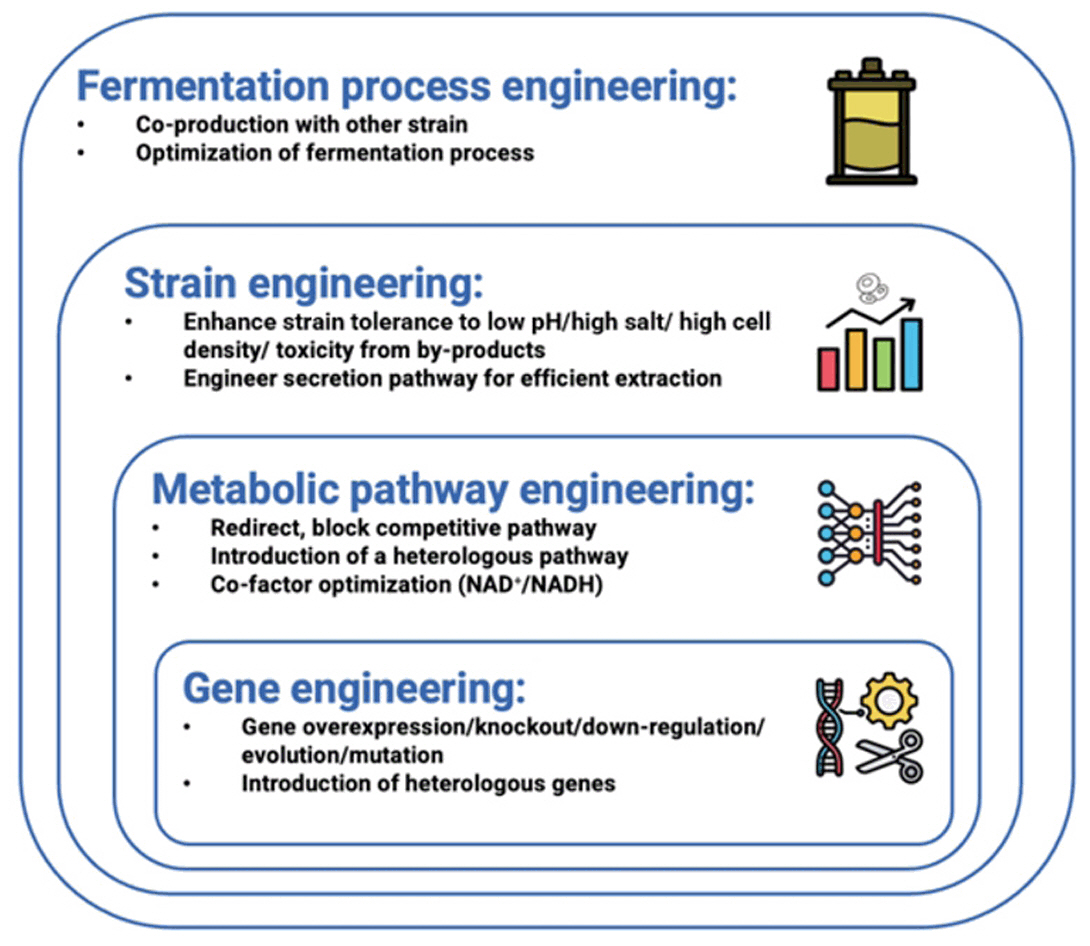
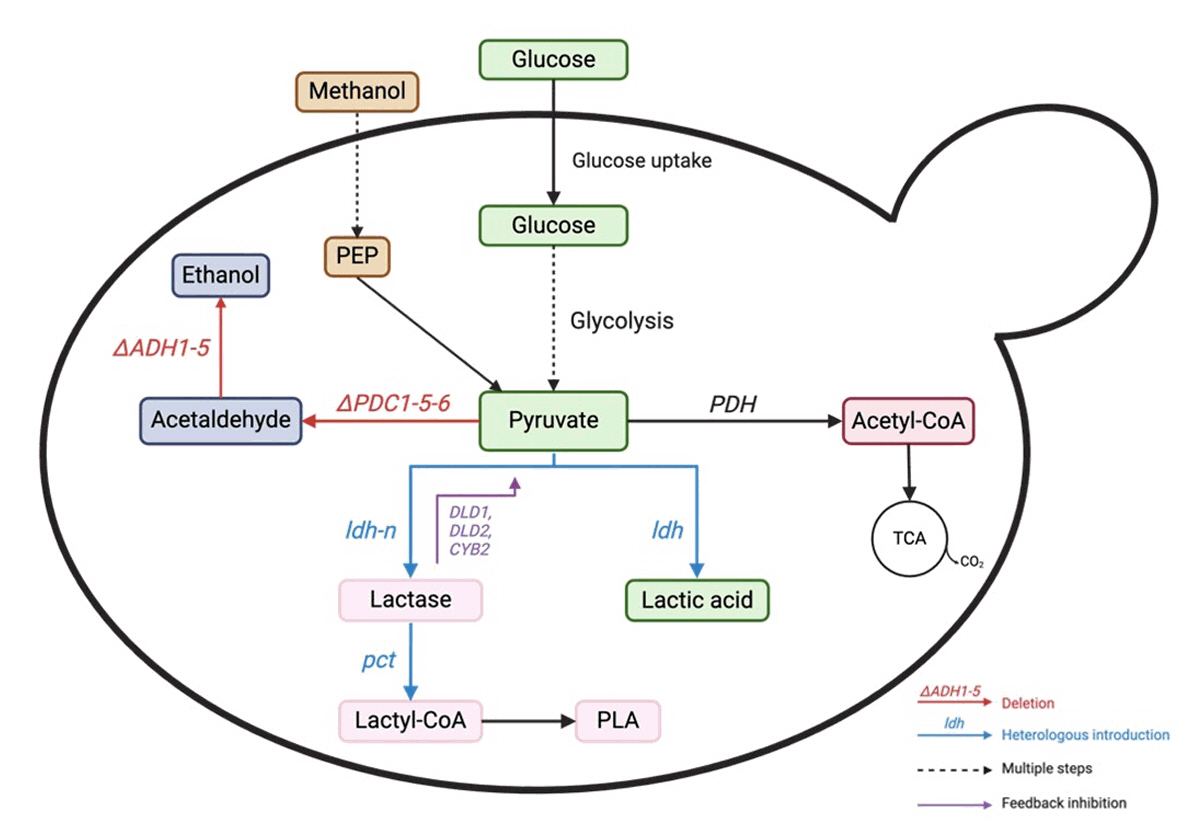
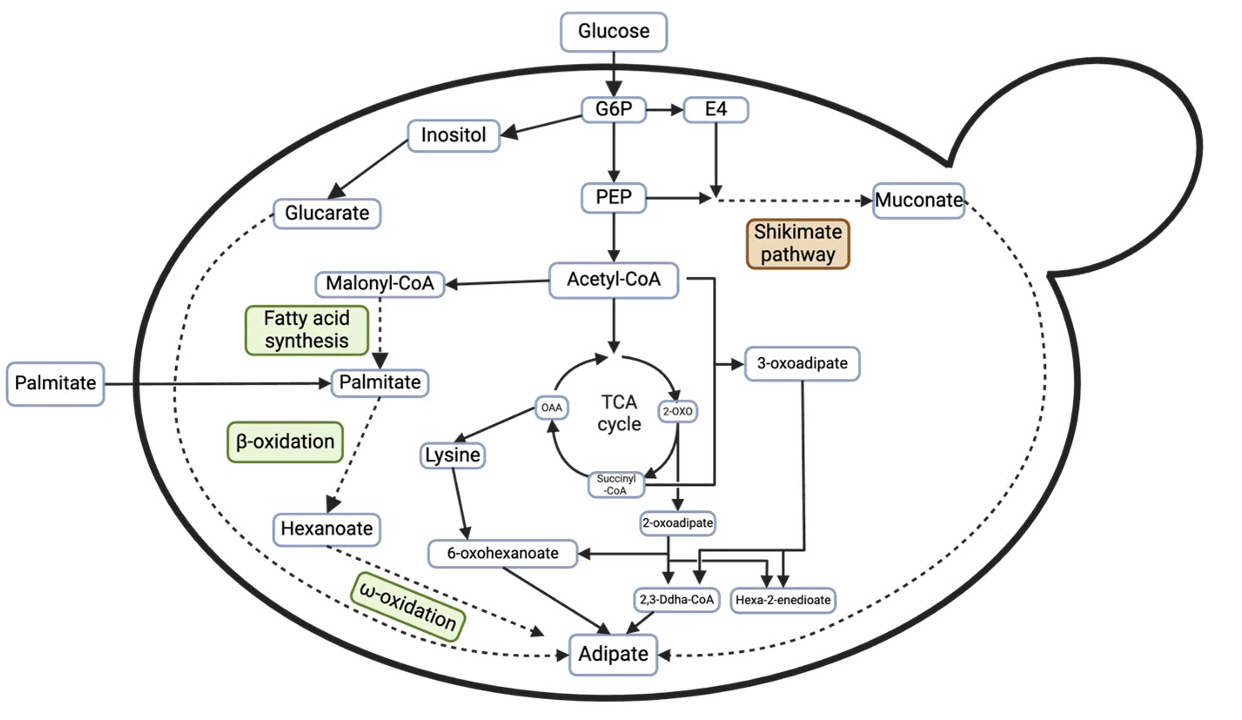
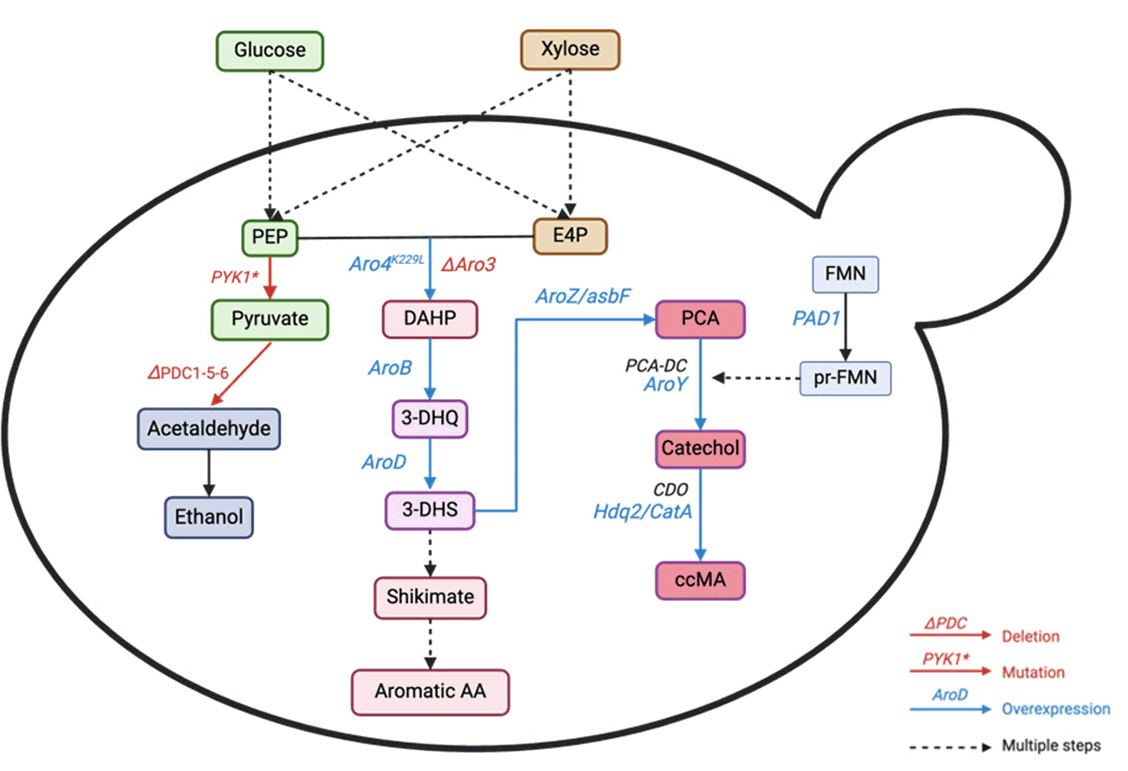
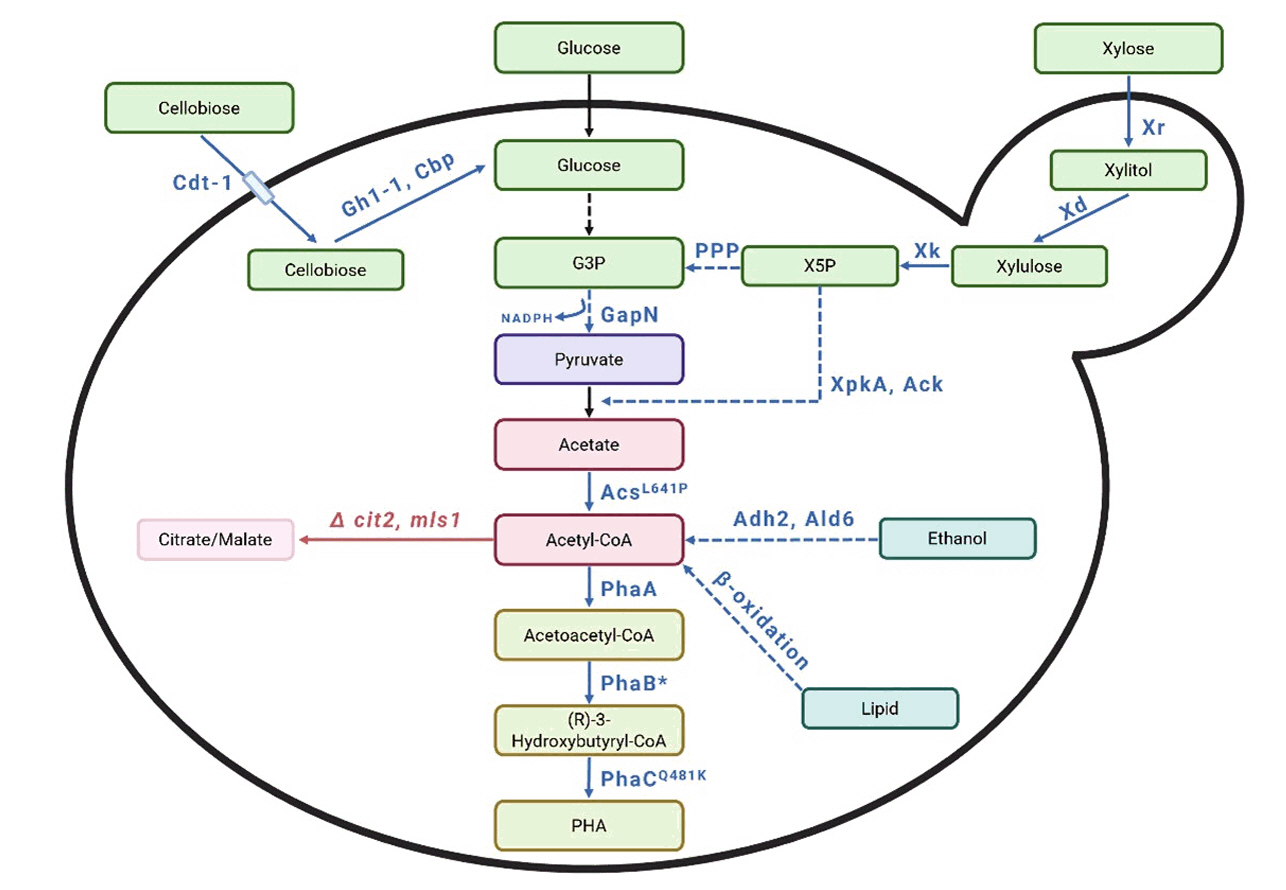
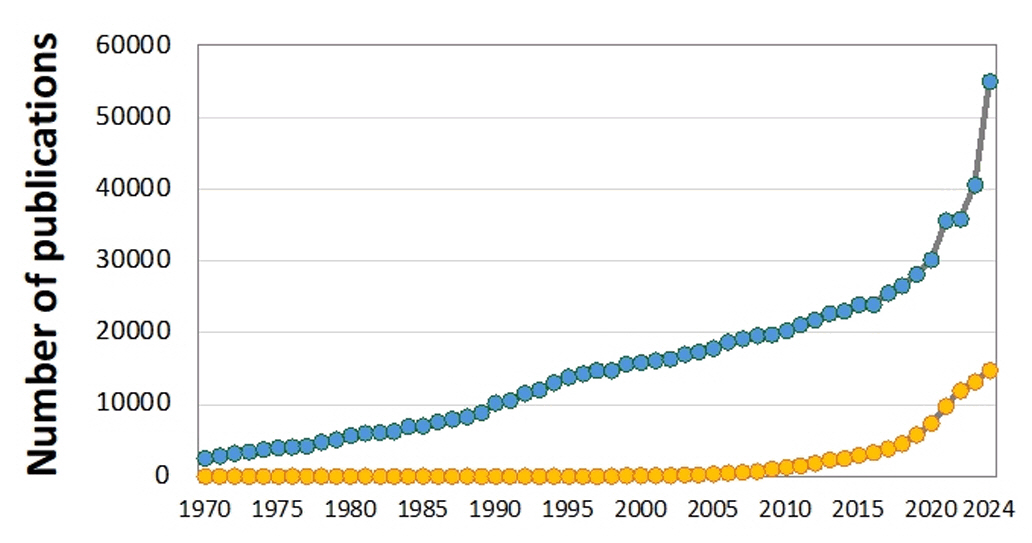
Fig. 1. Number of research publications on bioplastics (blue) and biodegradable (yellow) plastics between 1970 and 2024. Data from the Google Scholar search engine.
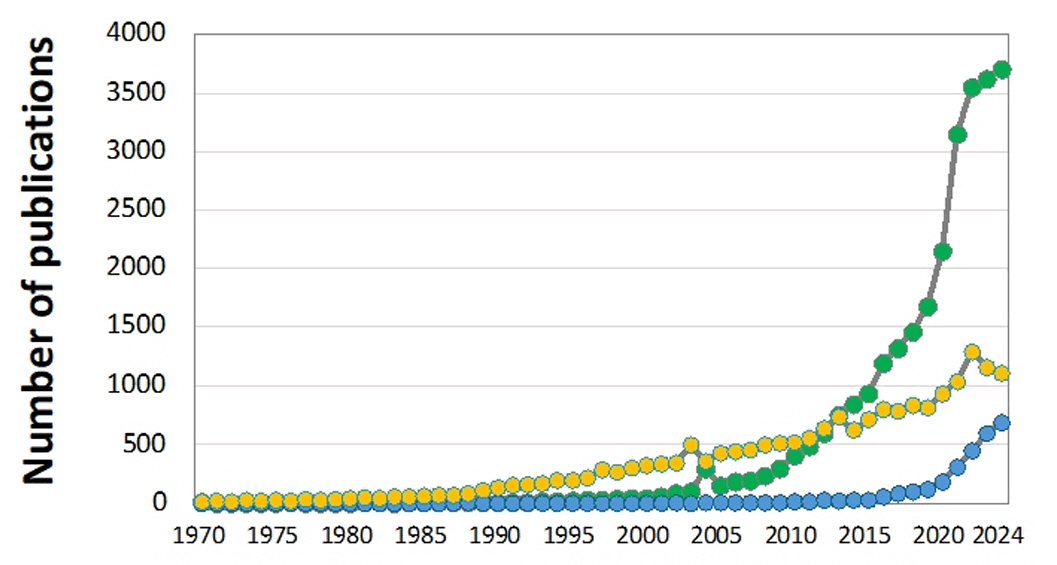
Fig. 2. Number of research publications on PLA (green), PBAT (blue), and PHAs (yellow) between 1970 and 2024. Data from the Google Scholar search engine.
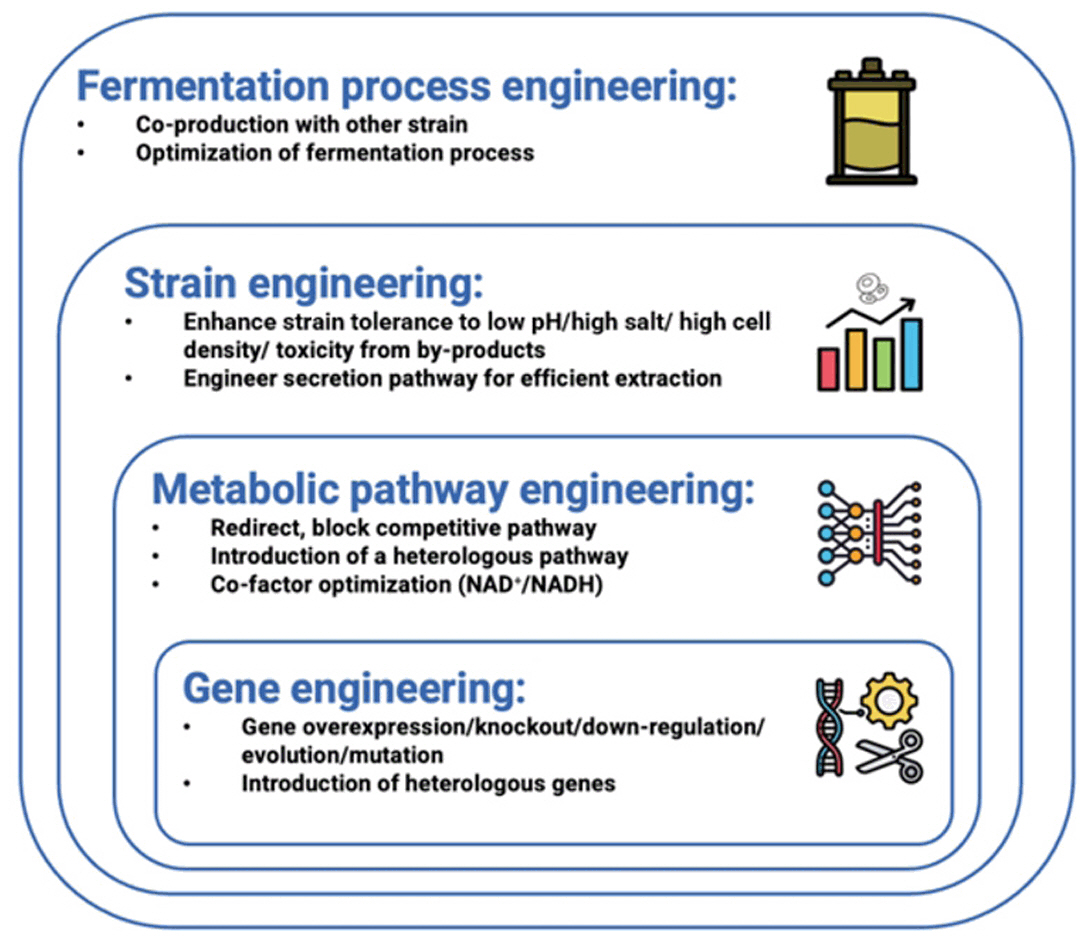
Fig. 3. Engineering strategies for yeast chassis in the production of biodegradable plastics.
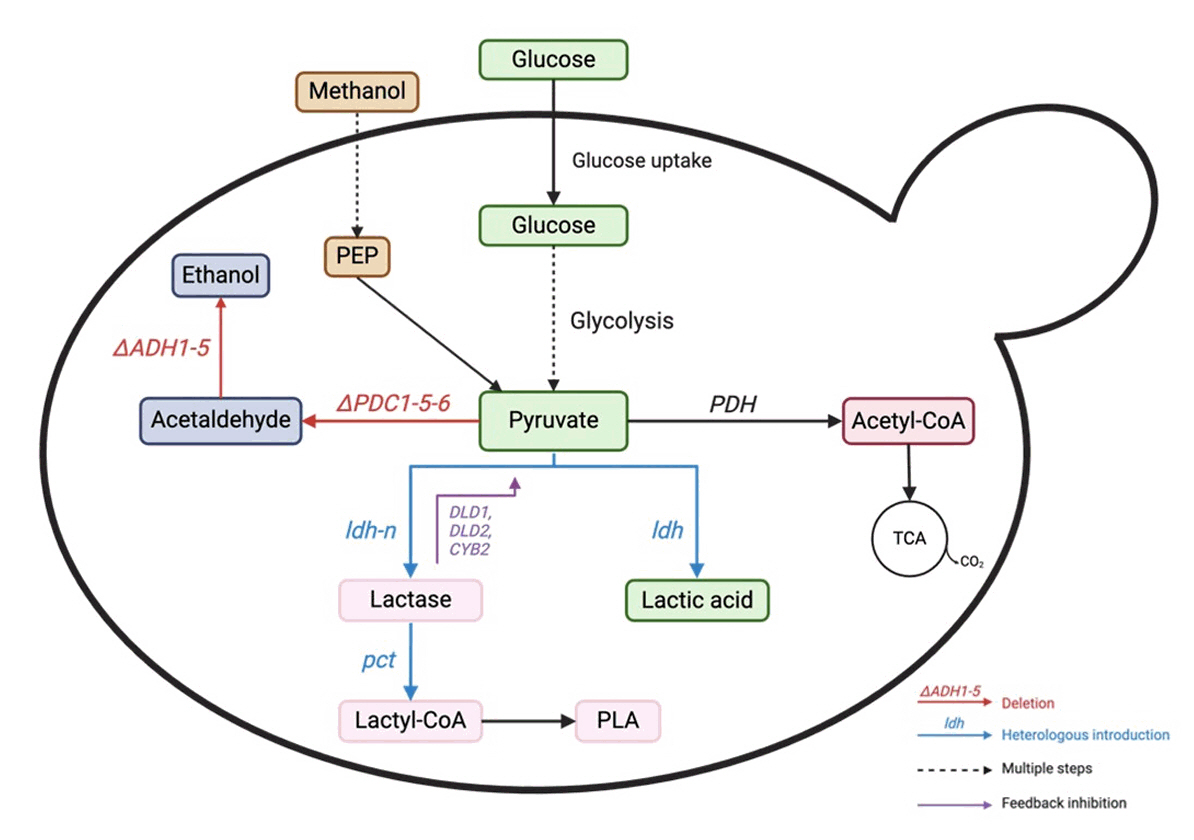
Fig. 4. Lactic acid production pathway from glucose in yeast. PDC: pyruvate decarboxylase, ADH: alcohol dehydrogenase, LDH: lactate dehydrogenase, PDH: pyruvate dehydrogenase. The red color represents gene deletion deactivation. The blue color represents the introduction of heterologous genes/gene overexpression.
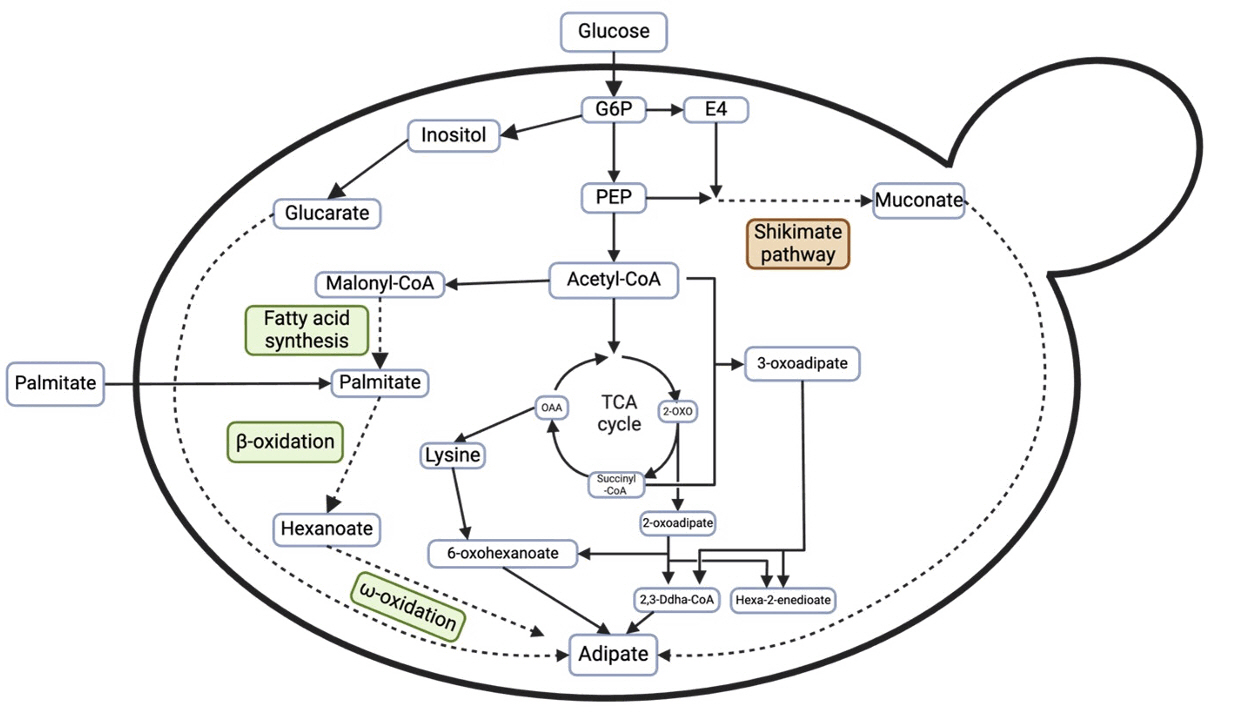
Fig. 5. Overview of metabolic pathway to adipic acid production (Averesch et al., 2018). G6P: Glucose-6-Phosphate, E4: Erythrose-4-Phosphate, PEP: Phosphoenolpyruvate, OAA: Oxaloacetate, 2-OXO: 2-Oxoglutarate.
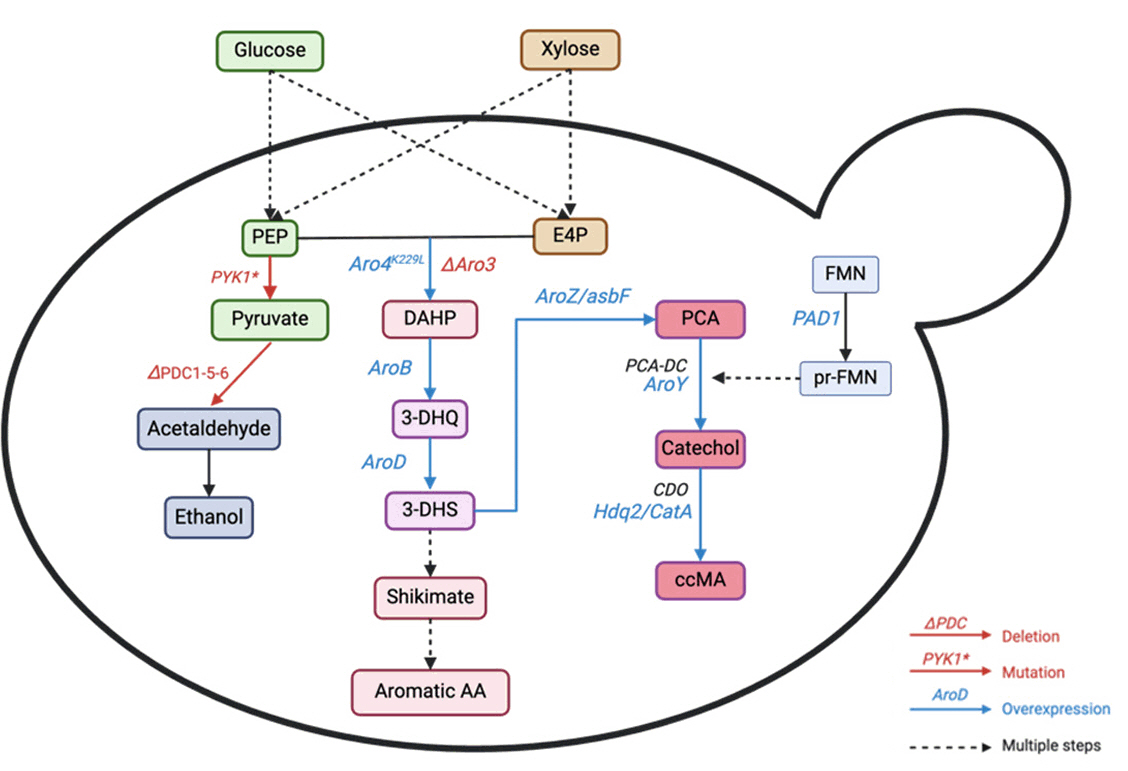
Fig. 6. Schematic representation of the metabolic pathway to produce muconic acid. PEP: Phosphoenolpyruvate, DAHP: 3-deoxyarabinoheptulosonate 7-phosphate, 3-DHQ: 3-dehydrogenate, 3-DHS: 3-DHS: 3-dehydroshikimate, PCA: protocatechuic acid, PCA-DC: PCA decarboxylase, CDO: catechol 1,2-dioxygenase, FMN, flavin mononucleotide, prFMN, prenylated FMN. The red color represents gene deletion deactivation. The blue color represents the introduction of heterologous genes/gene overexpression.
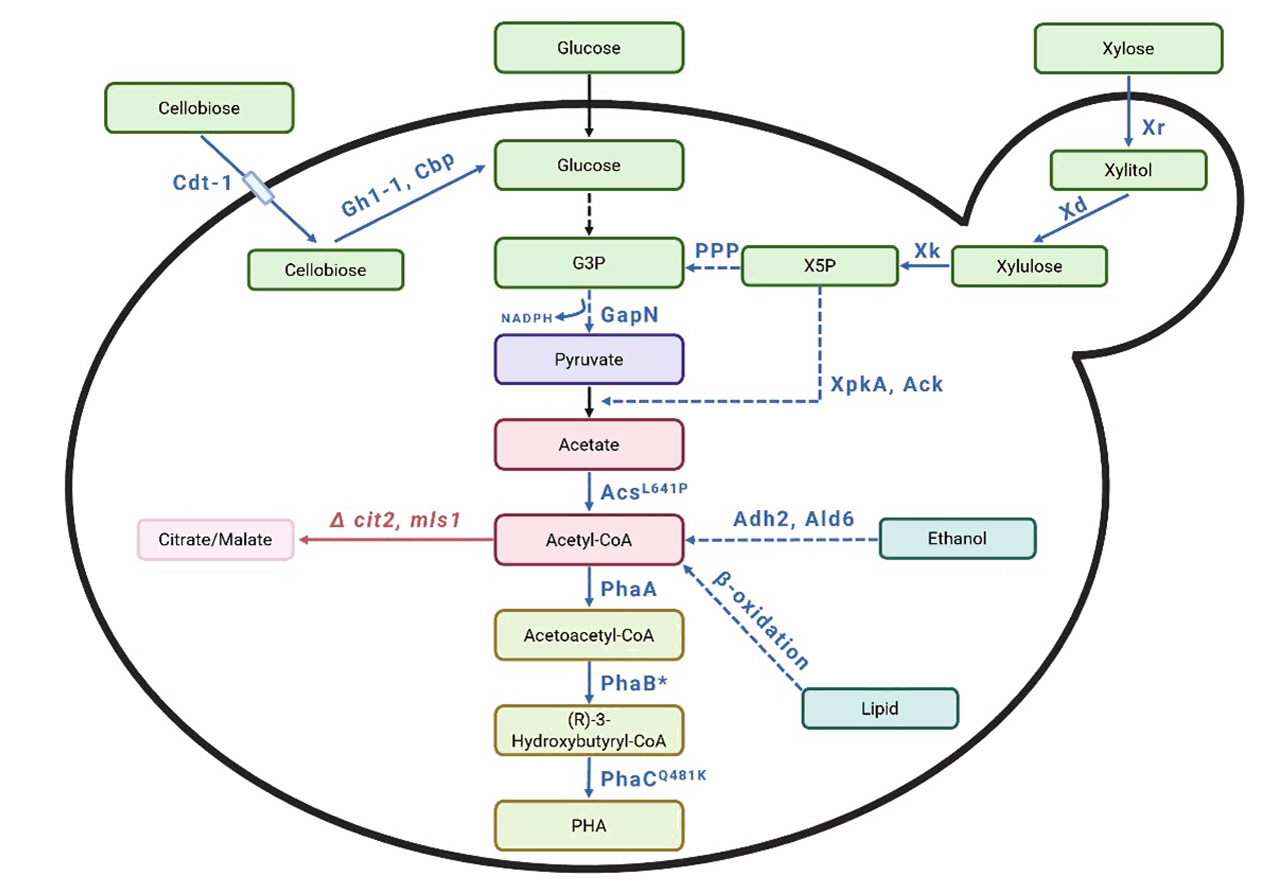
Fig. 7. Schematic representation of the metabolic pathway to produce polyhydroxyalkanoates from various substrates. G3P: Glyceraldehyde-3-phospahate, X5P: Xylulose-5-phosphate, PPP: Pentose phosphate pathway, Cdt-1: cellodextrin transporter, Gh1-1: β-glucosidase, Cbp: cellobiose phosphorylase, Xr: xylose reductase, Xd: xylitol dehydrogenase, Xk: xylulokinase, XpkA: xylulose-5-phosphate phosphoketolase, Ack: acetate kinase, GapN: NADP+-dependent glyceraldehyde-3-phosphate dehydrogenase, Adh2: alcohol dehydrogenase, Ald6: acetaldehyde dehydrogenase, cit2: citrate synthase, mls1: malate synthase PhaA: acetyl-CoA acetyltransferase, PhaB: acetoacetyl-CoA reductase, PhaC: PHA synthase. The red color represents gene deletion or deactivation. The blue color represents the introduction of heterologous genes or gene overexpression. A solid arrow indicates that the reaction occurs in a single step, while a dashed arrow indicates that it occurs over multiple steps.
Fig. 1.
Fig. 2.
Fig. 3.
Fig. 4.
Fig. 5.
Fig. 6.
Fig. 7.
Synthetic biology strategies for sustainable bioplastic production by yeasts
| Strains | Substrate | Engineering Strategies | Tools | Culture condition | Titer (g/L) | Productivity (g/L/h) | References |
|---|---|---|---|---|---|---|---|
| Saccharomyces cerevisiae SP1130 | Glucose | Express heterologous lactate dehydrogenase (LDH) genes, attenuating several key pathway genes including glycerol-3-phosphate dehydrogenase 1 (GPD1) and cytochrome-c oxidoreductase 2 (CYB2). Knocking out the pyruvate decarboxylase 1 (PDC1) and alcohol dehydrogenase 1 (ADH1) to disrupt the ethanol production pathway, redirect metabolic flux towards lactic acid production. Introduce an alternative pathway to produce acetyl-CoA using the acetylating acetaldehyde dehydrogenase (A-ALD) gene from E. coli to compensate for the attenuated ADH1 | Heterologous gene expression, Introduction of a synthetic pathway | Fed-batch fermentation | 142 | 3.55 | |
| S. cerevisiae T165R | Cane juice | Integrate six copies of L-LDH into the genome of host cell | Multi-integration of a heterologous gene | 1-L jar fermenter | L-LA 122 | - | |
| S. cerevisiae S.c-NO.2-100 | Glucose | Rewire the biosynthetic pathway for LA production by screening and introducing L-LDH, reducing ethanol accumulation by deleting PDC1, ADH1, introducing bacterial acetyl-CoA synthesis pathway. Strain evolution for acid tolerance by adaptive evolution. Delete NADH-consuming enzymes (NDE1/2) to enhance the cofactor availability. Overexpress bacterial transporters JEN1 and remodel the cell membrane by deleting SAM2 for efficient LA export | Heterologous gene expression, Introduction of heterologous transporter, ALE | 5-L batch bioreactor | L-LA 121.5 | 1.69 | |
| S. cerevisiae BK01 | Acetate-rich buckwheat husk hydrolysates | Adaptive laboratory evolution (ALE) on 8% lactic acid on S. cerevisiae SR8LDH strain which was previously constructed by the introduction of LDH gene from Lactobacillus acidophilus ATCC4356 under the control of the PGK1 promoter. The mutant BK01 was characterized by genome sequencing and metabolomic profiling | ALE, metabolomic profiling | 20 mL fermentation | L-LA 119 | 1.24 | |
| S. cerevisiae ASc- d789M | Glucose | Screen D-LDH from bacteria, overexpress LpDLDH, delete glycerol production genes GPD1, GPD2, and lactate dehydrogenase gene DLD1 to minimize by-products, downregulation ADH1 using the L-methionine repressible promoter to minimize the impact on growth. Created an intraspecific hybrid strain by mating the engineered strain with a weak acid-tolerant wild-type strain BCC39850 to improve acid tolerance and D-LA production | Gene knockout by CRISPR/Cas9, CRISPR/Cas12a, yeast mating | Fed-batch fermentation | D-LA 23.41 ± 1.65 | 0.81 | |
| S. cerevisiae | Glucose | Genome evolution applied a method combining GMES and CRISPR-δ integration (GMES/CRISPR) to integrate 13 bacterial genes including HXT7, HXK2, PGI1, PFK1, PFK2, FBA1, TPI1, TDH3, PGK1, GPM1, ENO2, and PYK2, and d-lactate dehydrogenase (d-LDH) into genomic DNA | Genome evolution by GMES and CRISPR-δ integration, overlap extension PCR, transcriptome analysis, real-time PCR | Semi-neutralizing fermentation | D-LA 52.2 | 2.17 | |
| S. cerevisiae JHY5330 | Glucose | Introduce D-lactate dehydrogenase gene (ldhA) from bacteria, reduce D-LA consumption by deleting DLD1 encoding D-lactate dehydrogenase and JEN1 encoding monocarboxylate transporter, eliminate ethanol production gene PDC1 and ADH1, glycerol production genes GPD1 and GPD2, improve strain LA tolerance by ALE and overexpression of HAA1 encoding a transcriptional activator involved in weak acid stress response | ALE, gene knockout, PCR-mediated method using the Cre/loxP system, quantitative reverse transcription PCR (qRT-PCR) | Fed-batch fermentation | D-LA 112 | 2.2 | |
| S. cerevisiae JHY5610 | Glucose | Employed ALE for LA-tolerance of host strain, coupled with introduction of LA production gene Lm. ldhA, and deletion of ethanol and glycerol production genes (ADH, GPD) and D-LA degradation genes to redirect metabolic flux toward LA production | ALE, gene knockout, whole genome sequencing | Fed-batch fermentation | D-LA 82.6 | 1.5 | |
| S. cerevisiae | Glucose | Rewire metabolic fluxes toward the production of L-lactic acid by deleting PDC1, CYB2, GPD1, and replacing LDH. Re-engineer the intracellular redox balance by deleting NDE1 and NDE2 encoding isoenzymes of the external NADH dehydrogenase | Gene knockout, re-engineer intracellular redox | Fed-batch | L-LA 117 | - | |
| Pichia pastoris | Methanol | Screen LDH genes, co-utilize LDH cofactors, knockout DLD1, DLD2 and CYB2 to block LA consumption pathway, and construct a coordinated dual pathway in cytoplasm and mitochondria | Site-directed mutation, CRISPR-Cas9, fluorescence microscopy analysis | Fed-batch fermentation | L-LA 4.2 | 0.0126 | |
| Pichia kudriavzevii NG7 | Glucose | Replace the pyruvate decarboxylase 1 gene (PDC1) with the d-lactate dehydrogenase gene (d-LDH) derived from Lactobacillus plantarum, improve strain tolerance by ALE | ALE, whole genome sequencing | Bioreactor fed-batch | D-LA 154 | 4.16 | |
| Candida boidinii | Glucose | Disrupted PDC1 to reduce ethanol production, introduce and express Bovine L-LDH gene under the PDC1 promoter and optimize fermentation condition | Heterologous gene expression | Batch fermentation | L-LA 85.9 | 1.79 | |
| S. cerevisiae | Glucose | Poly (D-lactic acid) (PDLA) and copolymer P (LA-3HB) were produced by express PHA synthase PhaC1437Ps6-19, propionyl-CoA transferase Pct540Cp, acetyl-CoA acetyltransferase PhaA, and acetoacetyl-CoA reductase PhaB1 | Gibson Assembly, modular cloning method (MoClo), GC-MS, NMR analysis | Erlenmeyer baffled flasks fermentation | PDLA 0.73% (CDW) | - | |
| Yarrowia lipolytica | Glucose, racemic lactic acid | Disrupt lactic acid consumption pathway, introduce heterologous pathway for PDLA production containing propionyl-CoA transferase (pct) from Clostridium propionicum to convert lactic acid into lactyl-CoA, and an evolved polyhydroxyalkanoic acid PHA synthase polymerizing lactyl-CoA into PDLA | Site-directed mutagenesis, UHPLC-FTMS | Erlenmeyer baffled flasks fermentation | PDLA 26 mg/g-DCW | - |
| Strains | Substrate | Engineering Strategies | Tools | Culture condition | Titer | References |
|---|---|---|---|---|---|---|
| E. coli BL21 (DE3) | Glycerol | Reconstruct five-step reverse adipate-degradation pathway (RADP) from Thermobifida fusca in E. coli, then overexpress 5-Carboxy-2-pentenoyl-CoA reductase (Tfu_1647) - the gene coding for the rate-limiting step in the RADP, delete succinate-CoA ligase gene (sucD) to eliminate competitive | Introduction of a synthetic pathway, Gibson assembly, CRISPR/Cas9, gene overexpression, SDS-PAGE | Fed-batch fermentation in 5-L bioreactor | 68.0 g/L | |
| Candida spp. | Fatty acid | Rewire the ω- then the β-oxidation pathway | - | Two-stage fed-batch process | 50 g/L | |
| Candida tropicalis | C12 methyl laurate | Create a mutant strain of C. tropicali by deleting AOX genes encoding acyl-CoA oxidases to produce AA via ω- and the β-oxidation pathway | Gene deletion | Bioreactor, fed-batch | 12.1 g/L | |
| S. cerevisiae | Glucose | Use the reverse adipate degradation pathway (RADP) from Thermobifida fusca, |
Introduction of a synthetic pathway, Gibson assembly, overlap extension PCR | Fed-batch bioreactor | 10.09 mg/L | |
| S. cerevisiae | Glucose | Rewire the cis, cis-muconic acid pathway by expressing enoate reductases (ERs) from Bacillus coagulans to convert ccMA into AA | Introduction of a synthetic pathway, Gibson assembly | Three stage fermentations | 2.59 mg/L, ccMA > 284 mg/L | |
| Y. lipolytica | Fatty acid methyl esters | Regulate the β-oxidation pathway by disrupting acyl-CoA oxidases (POX1 and POX3) and enhance ω-oxidation through overexpression of ALK5, CPR1, and FAO1, two-phase bioprocess to enhance AA production | Gene deletion, two-phase bioprocess | 20 ml in 250 ml baffled flask | 1176.2 mg/L |
| Strains | Substrate | Engineering Strategies | Tools | Culture condition | Titer (g/L) | Productivity (g/L/h) | References |
|---|---|---|---|---|---|---|---|
| S. cerevisiae | Glucose | Employed ccMA-biosensor, coupled with GFP expression to screen UV-mutagenesis libraries for ccMA producing yeast, then overexpress genes encoding for PCA decarboxylase and AROM protein and restore URA3 prototrophy. | UV-mutagenesis, fluorescence activated cell sorting (FACS), heterologous gene expression | Fed-batch | 20.8 | 0.139 | |
| S. cerevisiae | Glucose | Rewiring the shikimate pathway flux and enhancing the phosphoenolpyruvate supply by mutating PYK1 (A336S) to limit PEP-to-pyruvate conversion and overexpressing PYC1 and PCK1 to redirect pyruvate to PEP, overexpress DAHP synthase (ScARO4K229L), 3-dehydroquinate synthase (EcaroB), and 3-dehydroquinase | Cas9-assisted approach, Gibson assembly, USER cloning, EasyClone method | 2L fermenter | 22.5 | 0.19 | |
| S. cerevisiae | Glucose, xylose, mixture | Eliminated feedback inhibition in the shikimate pathway, insert heterologous pathway for MA production using 3-dehydroshikimate (DHS), protocatechuic acid decarboxylase (PCAD) and oxygen-consuming catechol 1,2-dioxygenase (CDO), delete PDC to eliminate ethanol production, minimized PCA production by enhancing PCAD overexpression and production of its co-factor | Introduction of a synthetic pathway, genome editing using CRISPR/Cas9 | 300 ml shake flasks with 40 mL YP medium | 4.5 | - | |
| S. cerevisiae | Glucose and xylose | Eliminates C2 dependency and ethanol production by ALE and deleting PDC1, PDC5, and PDC6, improving MA tolerance for host strain, promoting growth and production of target product by internal deletion of MTH1, MTH1ΔM41-T78, overexpressing PAD1 as well as reduces side production of intermediate protocatechuic acid (PCA) | Introduction of a synthetic pathway, enzyme engineering, ALE | Fed-batch | 9.3 | 0.100 | |
| S. cerevisiae | Glucose | Combine adaptive laboratory evolution (ALE) and rational metabolic engineering, improve flux by truncation of ARO1 and overexpress an endogenous aromatic decarboxylase | ALE, Gibson assembly | Fed-batch bioreactor | 2.1 | - | |
| S. cerevisiae | Glucose | Employ the computational tool YEASTRACT for predicting novel transcriptional repressors and OptForce strain-design for identifying non-intuitive pathway interventions, strating from glycolytic and pentose phosphate pathway pathway, follow by engineer ARO2, ARO3, ARO4, and the pentafunctional ARO, delete ric1 to increase transcription of ARO2, ARO3, ARO4 | Computational tool for predicting engineering target, Introduction of a synthetic pathway Gibson assembly | - | 0.32 | - | |
| S. cerevisiae | Glucose, xylose (supplement 1 g/L catechol) | Overexpress xylose isomerase gene from Bacteroides valgutus and pentose phosphate pathway genes from S. cerevisiae, then imported a three step ccMA production pathway from E. coli with AroZ-Neu, AroY-Com, and CatA-Cup genes, further overexpression of gene Aro1 (with a stop codon of AroE) and feedback-resistant Aro4opt mutant gene, ALE to improve xylose fermentation and ccMA | Introduction of a synthetic pathway, Enzyme engineering, ALE | Batch, shake flask | 0.424 (1.286 with catechol supplemented) | - | |
| S. cerevisiae | Glucose, supplement amino acids | Balance MA pathway performance with aromatic amino acid prototrophy by destabilizing Aro1, delete ARO4 and ARO3, introduce AroZ, AroY, Hqd2 CatA, Pad1. | CRISPR-mediated homology-directed repair (HDR) | Fed-batch bioreactor | 5.1 | - | |
| S. cerevisiae | Glucose | Construct cassettes using USER cloning-ligation-PCR, express a three-step heterologous pathway using KpAroY.B, KpAroY.Ciso, CaCatA, engineering TKL1, ZWF1, Aro1AroEΔ, Aro4K229L | Introduction of a synthetic pathway, CRISPR/Cas9, in vivo recombination USER assembly and RNA interference | Batch, shake flask | 0.8 | - | |
| Pichia occidentalis | Glucose | Screen host strain with high tolerance to MA, develop genome editing toolkit to engineer the non-conventional yeast and introduce the heterologous pathway for ccMA production | CRISPR-Cas9, antibiotic marker recycling, high-throughput screening | Fed-batch | 38.8 | 0.511 |
| Strains | Substrate | Engineering Strategies | Tools | Culture condition | Titer (mg/L) | Yield (mg/g) | References |
|---|---|---|---|---|---|---|---|
| E. coli | Glucose, xylose | Knocked out ndh gene. Introduced arabinose/xylose transport proteins from Bacillus subtilis (xylA, xylB, and araE). Incorporated the PHB biosynthesis pathway from C. necator | Modification of ribosome binding sites | Fed-batch culture | PHB 21010 | - | |
| S. cerevisiae | Silver grass, glucose, xylose | Constructed a polycistronic PHB cassette with phaA, phaB, and phaC using 2A peptide sequences. Introduced Cas9-expressing plasmid, gRNA, and polycistronic PHB cassette into S. cerevisiae via electroporation. Integrated the polycistronic PHB cassette into the genome | CRISPR-Cas9 mediated genome editing, 2A peptide sequence integration | Serum bottle, microaerobic | PHB 40.3 | 0.93 mg/g silver grass | |
| S. cerevisiae | Glucose | Expressed XpkA and Ack from Aspergillus nidulans to increase acetyl-CoA levels. Incorporated an acetyl-CoA synthase variant (AcsL641P) from Salmonella enterica. Integrated GapN (NADP⁺-dependent glyceraldehyde-3-phosphate dehydrogenase) from Streptococcus mutans into the genome via the Li-Ac method. Codon optimized all genes for efficient expression | Codon optimization DNA 2.0 and GenScript for synthesizing genes, PTEF1-PPGK1 bidirectional promoter | Shake flask, anaerobic | PHB 180 | 0.03 mg/g glucose | |
| S. cerevisiae | Glucose | Synthesized phaA, phaB, and phaC based on R. eutropha H16. Introduced them into S. cerevisiae in plasmid form using the Li-Ac method. Used the strain described by Chen et al. (2012) to increase acetyl-CoA levels. Deleted cit2 and mls1, overexpressed ALD6 and ADH2. Codon optimized all genes for efficient expression | Codon optimization, PTEF1-PPGK1 bidirectional promoter, DNA 2.0 for synthesize genes | Shake flask, aerobic | PHB 43.11 | 0.13 mg/g glucose | |
| S. cerevisiae | Xylose | Introduced XR and XD genes to enable xylose utilization. PCR-amplified phaA, phaB, and phaC genes from C. necator. Assembled these genes into a single plasmid using In-Fusion cloning. Introduced the plasmid into S. cerevisiae via the Li-Ac method. Codon optimized all genes for efficient expression | In-fusion cloning, Codon optimization, overlap extension PCR | Shake flask, aerobic | PHB 45 | 1.17 mg/g xylose | |
| S. cerevisiae | Xylose | Achieved PHB production under anaerobic conditions using xylose as the carbon source. Replaced PhaB from C. necator with NADH-dependent PhaB from Allochromatium vinosum. Introduced the gene into S. cerevisiae in plasmid form via the Li-Ac method | Codon optimization, heterologous gene expression, promoter and terminator pairing | Bioreactor, anaerobic | PHB 360 | 7 mg/g xylose | |
| S. cerevisiae | Xylose high nitrogen supply | Investigated whether nitrogen limitation in yeast influences PHB production. Introduced Xr and Xd from Scheffersomyces stipitis to enable xylose utilization. Replaced PhaB from C. necator with NADH-dependent PhaB from A. vinosum. Assembled these genes into a single plasmid using In-Fusion cloning. Introduced the plasmid into S. cerevisiae via the Li-Ac method. Codon optimized and PCR-amplified all genes | In-fusion cloning, Codon optimization, heterologous gene expression, promoter and terminator pairing | Shake flask, anaerobic | PHB 730 | 13.8 mg/g xylose | |
| S. cerevisiae | Cellobiose | Integrated phaA, phaB1, and phaC1 into the S. cerevisiae genome using Gibson assembly, Li-Ac method, and CRISPR-Cas9. Inserted Gh1-1 from Neurospora crassa and Cbp from Ruminococcus flavefaciens for cellobiose metabolism using the Modular Cloning (MoClo) approach. Codon optimized all genes for efficient expression | Gibson Assembly, Modular cloningCRISPR-Cas9 mediated genome integration, promoter and terminator pairing | Bioreactor | PHB - | 49.4 mg/g cellobiose | |
| Y. lipolytica | Tridecanoate | Directed carbon flux in Y. lipolytica toward either the β-oxidation pathway or the PHA biosynthesis pathway based on the genotype of POX1-6 genes encoding acyl-CoA oxidase isoenzymes. Downregulated the R-3-hydroxyacyl-CoA dehydrogenase multifunctional enzyme domain in the fatty acid β-oxidation pathway via site-directed mutagenesis, generating the MFE-AG16SBC variant. Deleted an acyltransferase gene and overexpressed enoyl-CoA hydratase 2 to further redirect fatty acids from lipid synthesis to the β-oxidation pathway | site-directed mutagenesis, loxP-Cre recombinase, pGEMT easy cloning | Shake flask | PHA - | 7.3% DCW | |
| Y. lipolytica | Tridecanoate | Directed carbon flux in Y. lipolytica toward either the β-oxidation pathway or the PHA biosynthesis pathway based on the genotype of POX1-6 genes encoding acyl-CoA oxidase isoenzymes. Introduced PhaC from Pseudomonas aeruginosa and modified its C-terminal region to include the 34-amino acid sequence from Brassica napus glyoxysomal isocitrate lyase (icl) for peroxisome targeting | Site-directed mutagenesis, STADEN package for sequence analysis | Shake flask | PHA - | 17.64 mg/g biomass | |
| Y. lipolytica | Methyl laurate | Generated various PhaC variants through site-directed mutagenesis. Compared their production yields of mcl-PHA with different chain lengths | Site-directed mutagenesis | - | PHA - | 28% DCW | |
| Y. lipolytica | Triolein | Directed carbon flux in Y. lipolytica toward either the β-oxidation pathway or the PHA biosynthesis pathway based on the genotype of POX1-6 genes encoding acyl-CoA oxidase isoenzymes. Heterologously expressed PhaC1 from Pseudomonas aeruginosa PAO1 with a peroxisomal signal. Introduced the gene into S. cerevisiae in plasmid form via the Li-Ac method. Codon optimized all genes for efficient expression | Codon optimization | Shake flask | PHA 1110 | 5.0% DCW |
Table 1. Comparison of lactic acid production by engineered yeasts
Table 2. Comparison of adipic acid production by engineered yeast
Table 3. Comparison of cis,cis-muconic acid production by engineered yeast
Table 4. Comparison of PHA production by engineered yeast
Table 1.
Table 2.
Table 3.
Table 4.
TOP
 MSK
MSK
 ePub Link
ePub Link Cite this Article
Cite this Article








