- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Volume 63(3); 2025 > Review
-
Review
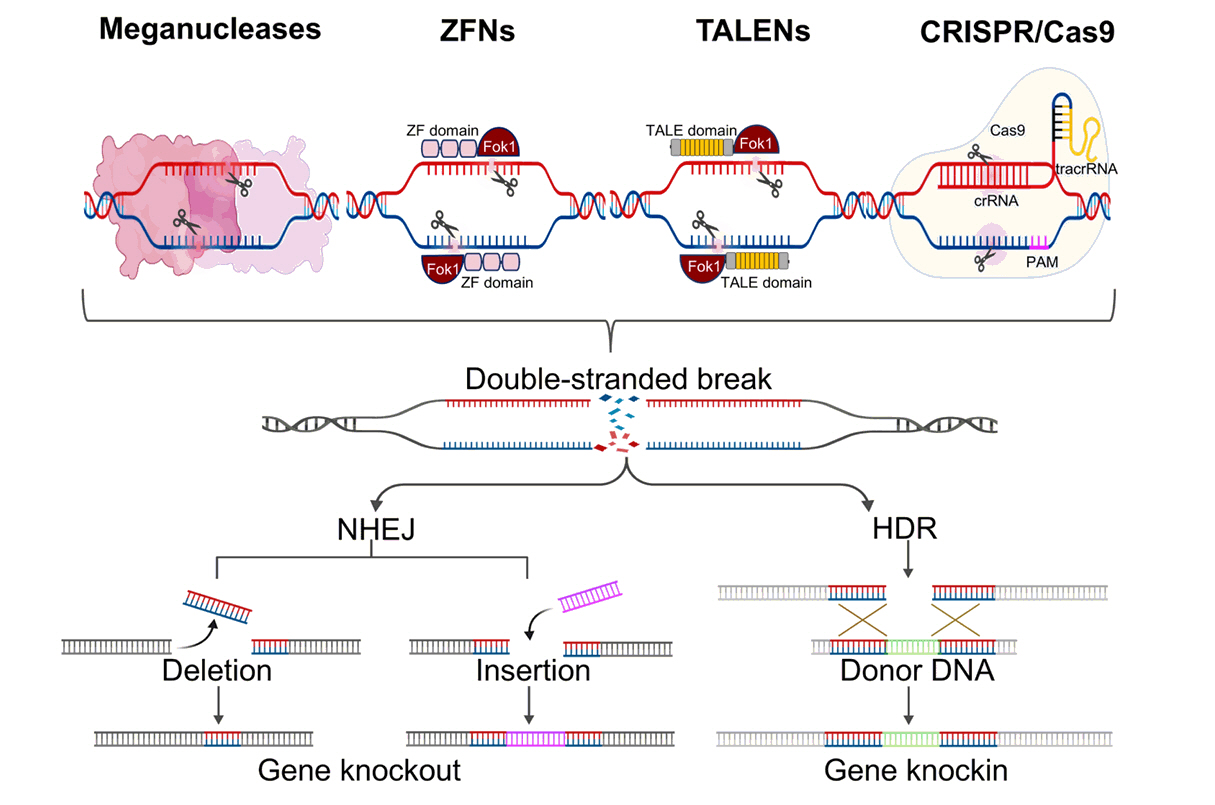
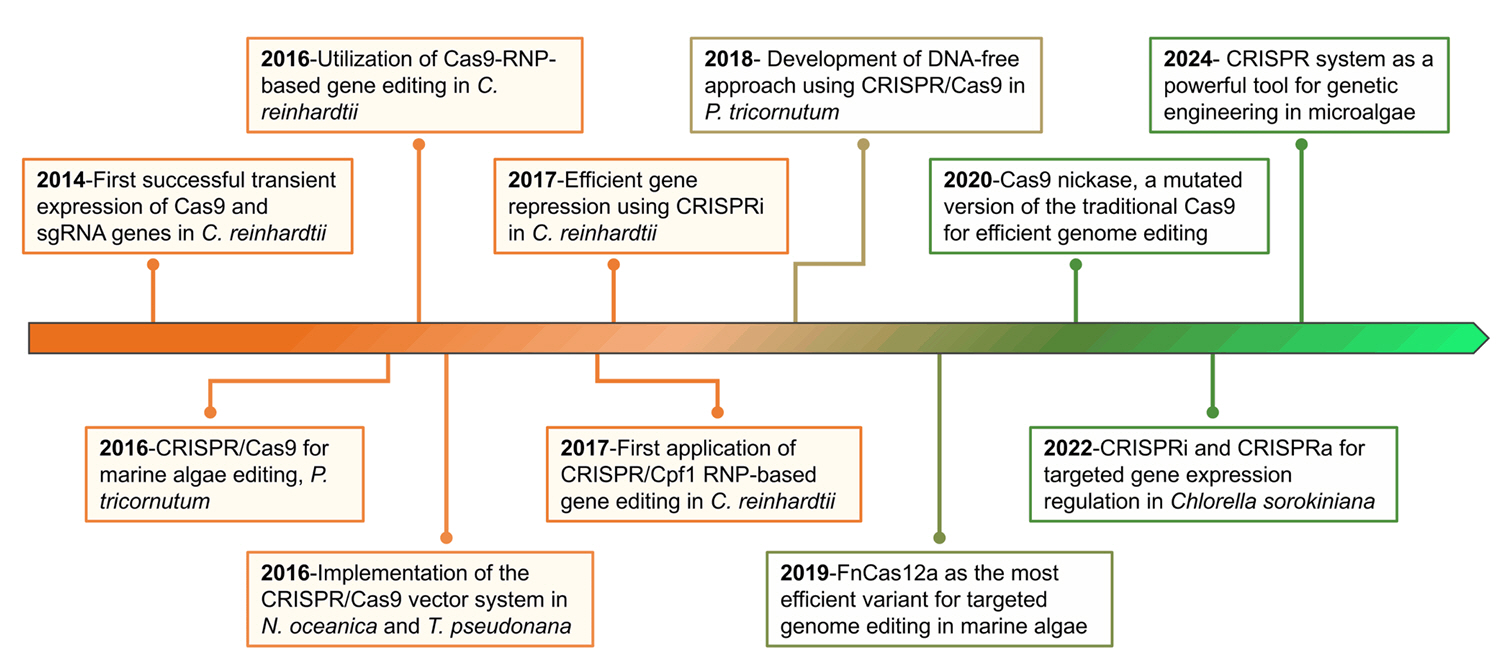
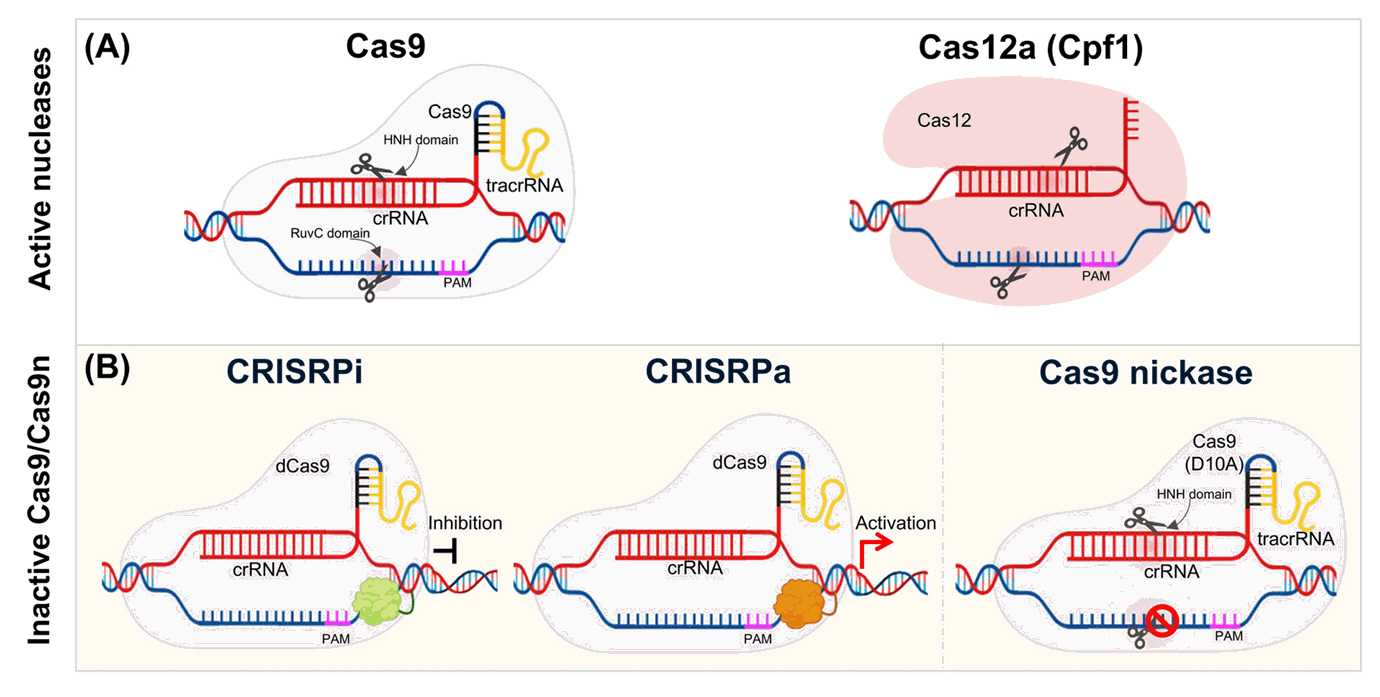
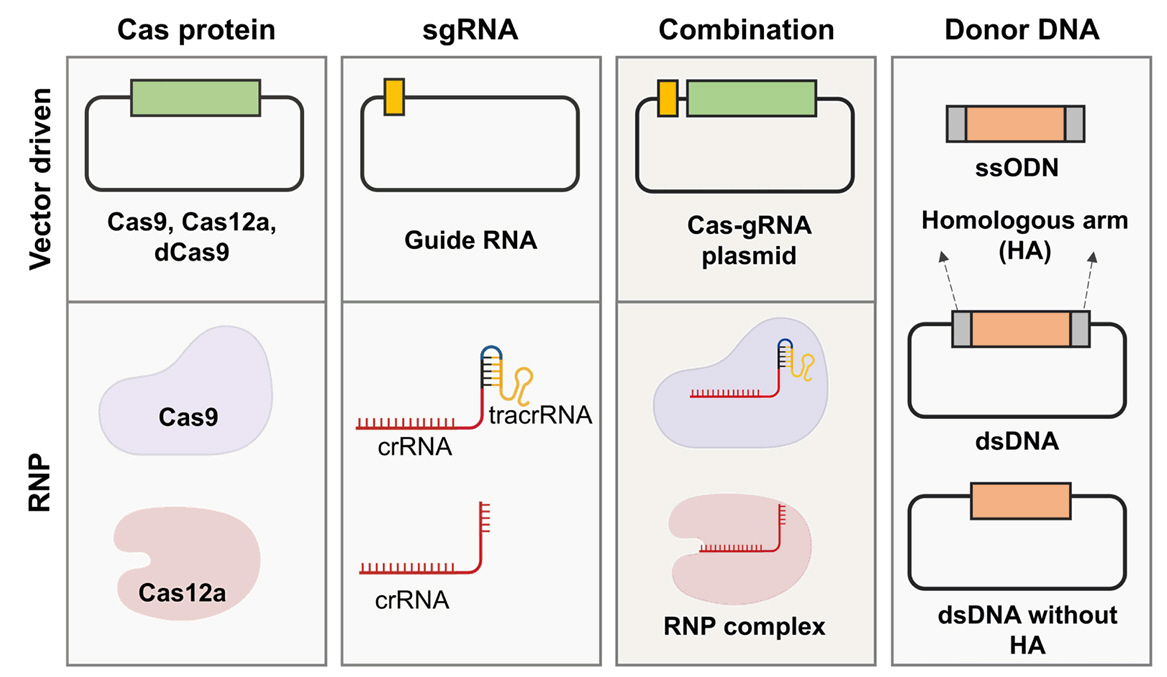
Progress and challenges in CRISPR/Cas applications in microalgae - Quynh-Giao Tran1,†, Trang Thi Le1,2,†, Dong-Yun Choi1, Dae-Hyun Cho1, Jin-Ho Yun1,2, Hong Il Choi1,2, Hee-Sik Kim1,2,*, Yong Jae Lee1,2,*
-
Journal of Microbiology 2025;63(3):e2501028.
DOI: https://doi.org/10.71150/jm.2501028
Published online: March 28, 2025
1Cell Factory Research Center, Korea Research Institute of Bioscience and Biotechnology (KRIBB), Daejeon 34141, Republic of Korea
2Department of Environmental Biotechnology, KRIBB School of Biotechnology, University of Science and Technology (UST), Daejeon 34113, Republic of Korea
- *Correspondence Hee-Sik Kim hkim@kribb.re.kr Yong Jae Lee leeyj@kribb.re.kr
- †These authors contributed equally to this work.
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Supplementary Information
References
Citations

- Active and targeted micro/nanoplastics remediation via engineered microalgae co-displaying polymer-binding peptides and plastic-degrading enzymes: A critical review and perspectives
Ling Wang, Mingjing Zhang, Jialin Wang, Chen Hu, Zhanyou Chi, Lei Li, Wenjun Luo, Chengze Li, Chenba Zhu
Algal Research.2026; 93: 104455. CrossRef - Advancing microbial engineering through synthetic biology
Ki Jun Jeong
Journal of Microbiology.2025; 63(3): e2503100. CrossRef - Progress and prospects in metabolic engineering approaches for isoprenoid biosynthesis in microalgae
Sonia Mohamadnia, Borja Valverde-Pérez, Omid Tavakoli, Irini Angelidaki
Biotechnology for Biofuels and Bioproducts.2025;[Epub] CrossRef - Beyond Biomass: Reimagining Microalgae as Living Environmental Nano-Factories
Thinesh Selvaratnam, Shaseevarajan Sivanantharajah, Kirusha Sriram
Environments.2025; 12(7): 221. CrossRef - Harnessing MicroRNAs and CRISPR to enhance biofuel production in microalgae
Dariga K. Kirbayeva, Altynay Y. Shayakhmetova, Bekzhan D. Kossalbayev, Assemgul K. Sadvakasova, Meruyert O. Bauenova
International Journal of Hydrogen Energy.2025; 157: 150399. CrossRef - Beyond Cutting: CRISPR-Driven Synthetic Biology Toolkit for Next-Generation Microalgal Metabolic Engineering
Limin Yang, Qian Lu
International Journal of Molecular Sciences.2025; 26(15): 7470. CrossRef - Mechanistic Role of Heavy Metals in Driving Antimicrobial Resistance: From Rhizosphere to Phyllosphere
Rahul Kumar, Tanja P. Vasić, Sanja P. Živković, Periyasamy Panneerselvam, Gustavo Santoyo, Sergio de los Santos Villalobos, Adeyemi Nurudeen Olatunbosun, Aditi Pandit, Leonard Koolman, Debasis Mitra, Pankaj Gautam
Applied Microbiology.2025; 5(3): 79. CrossRef - Strain Improvement Through Genetic Engineering and Synthetic Biology for the Creation of Microalgae with Enhanced Lipid Accumulation, Stress Tolerance, and Production of High-value
Alebachew Molla, Gedif Meseret
Science Frontiers.2025; 6(3): 80. CrossRef - The Role of Molecular Tools in Microalgal Strain Improvement: Current Status and Future Perspectives
Alebachew Molla, Gedif Meseret
Advances in Bioscience and Bioengineering.2025; 13(3): 51. CrossRef - CRISPR-Cas9 genome editing in microalgae for improved high-value products (HVP) production
Fazleen Haslinda Mohd Hatta, Nurin Nisa’ Ahmad Zamri, Norazlina Ahmad
Asia Pacific Journal of Molecular Biology and Biotechnology.2025; : 245. CrossRef - Advances in Algae-Based Bioplastics: From Strain Engineering and Fermentation to Commercialization and Sustainability
Nilay Kumar Sarker, Prasad Kaparaju
Fermentation.2025; 11(10): 574. CrossRef - Exploring the nutritional and bioactive potential of microalgal sulfated polysaccharides for functional food applications
Fajar Sofyantoro, Eka Sunarwidhi Prasedya, Fahrul Nurkolis, Andri Frediansyah
Food Science and Biotechnology.2025;[Epub] CrossRef - Harnessing microalgae for bioproducts: innovations in synthetic biology
Zheng Li, Yuhui Cheng, Chengcheng Li, Qianyi Wu, Yi Xin
World Journal of Microbiology and Biotechnology.2025;[Epub] CrossRef
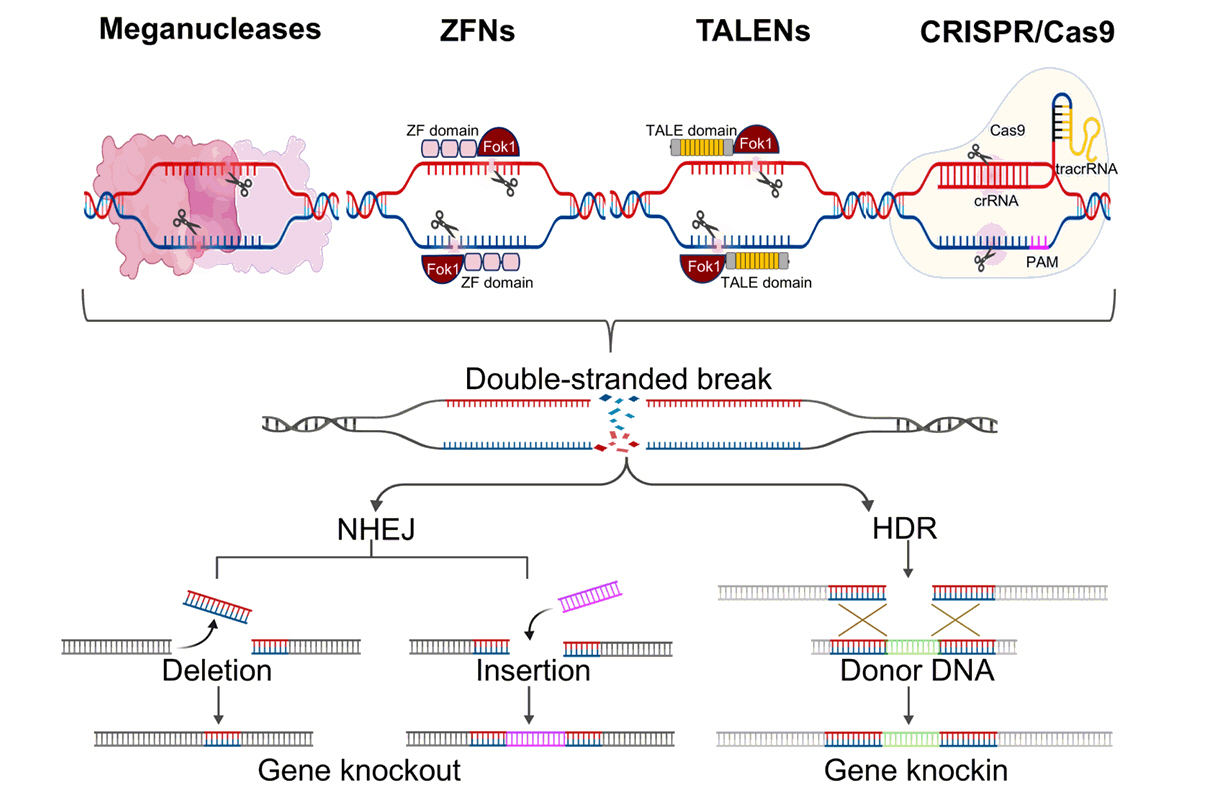
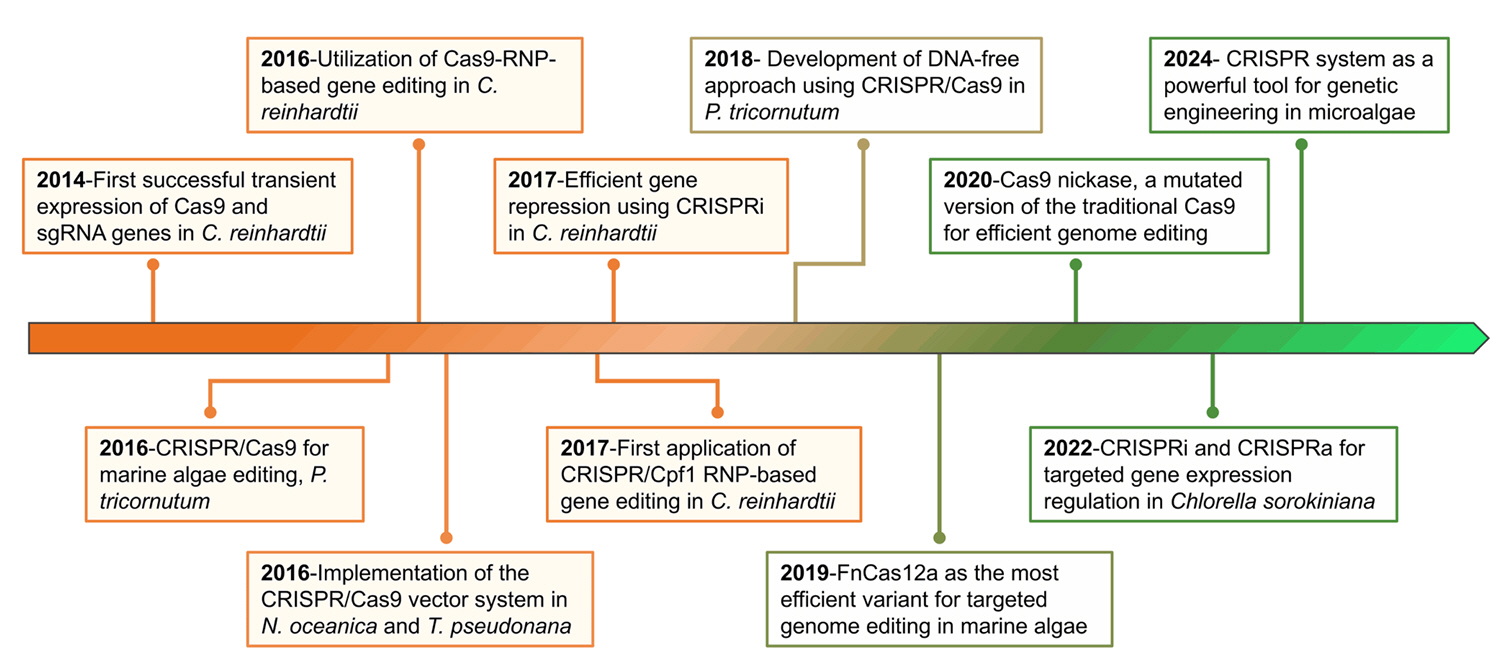
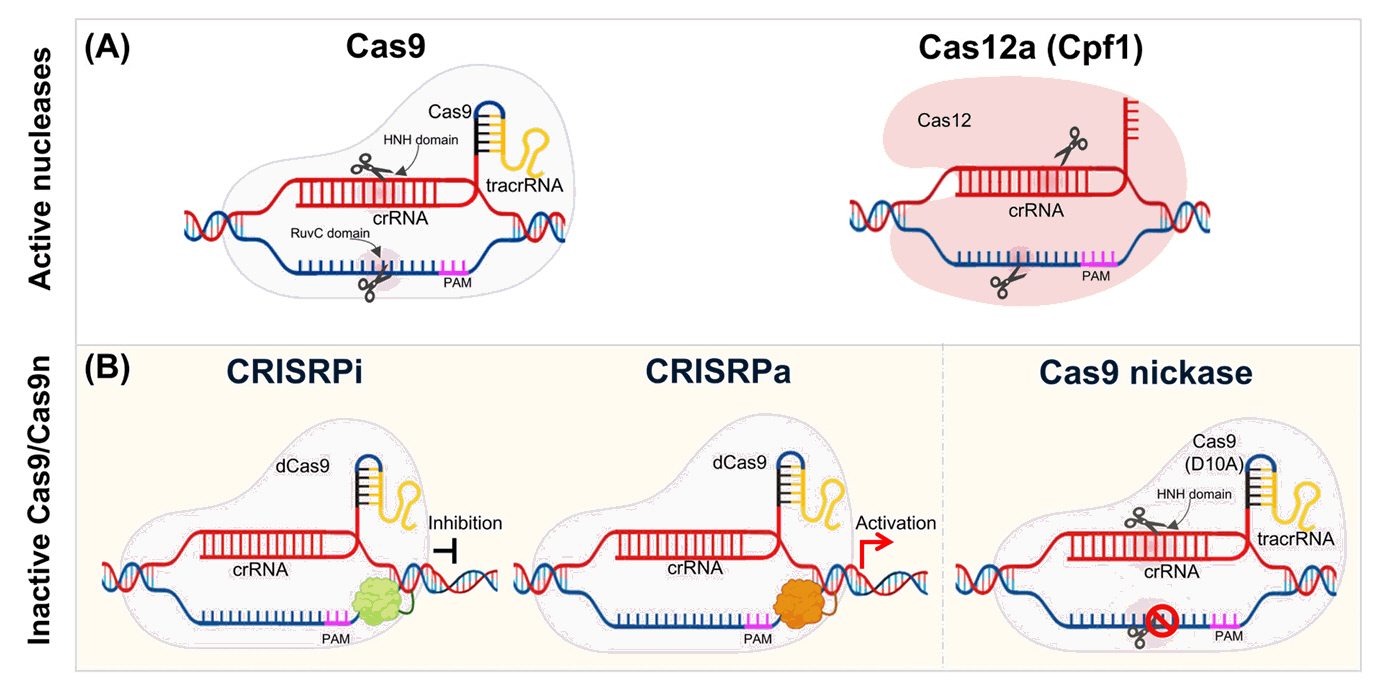
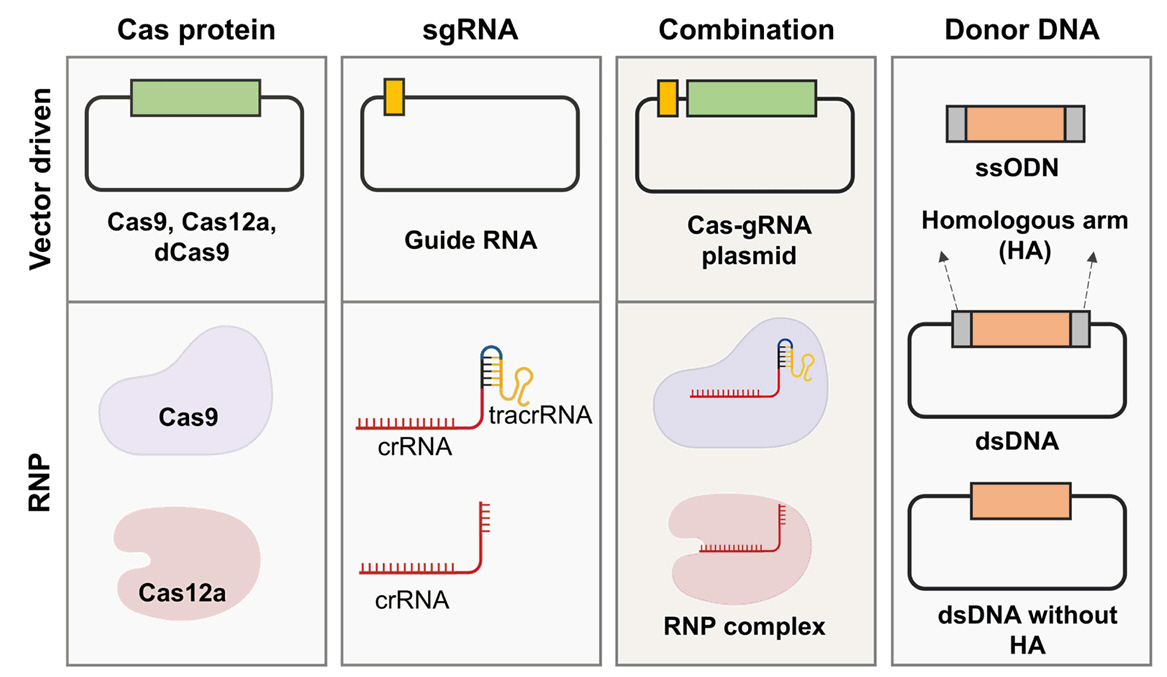
Fig. 1.
Fig. 2.
Fig. 3.
Fig. 4.
| Algal strain | CRISPR/Cas system | Strategy |
Targeted genes | Selection Marker | Products | Mutagenesis efficiency (%) |
Targeted mutagenesis frequency |
References | |
|---|---|---|---|---|---|---|---|---|---|
| Approaches | Delivery method | ||||||||
| Chlamydomonas reinhardtii | Cas9 | Vector driven | Electroporation | FKB12 | Rapamycin | First successful transient expression of Cas9 and sgRNA genes in C. reinhardtii | 0.0014 | 2 × 10-8 | |
| Cas9 | RNP | Electroporation | MAA7, CpSRP43, ChlM | Auxotrophic selection using 5-FI | Visible auxotrophic colonies with mutations targeted at the Cas9 cut sites | 40% (MAA7), 1.4% (CpSRP43), and 0.17% (ChlM) | 8.9 × 10-8 (MAA7), 3.3 × 10-8 (CpSRP43), and 5 × 10-8 (ChlM) | ||
| Cas9 | RNP | Electroporation | CpFTSY, ZEP | Based on the coloration of the cells | High zeaxanthin-producing mutants with improved photosynthetic productivity | 0.46 to 0.56% | Not mention | ||
| CRISPRi | Vector driven | Glass beads | PEPC1 | Hygromycin and paromomycin | A 74.4% increase in lipid content and a 94.2% enhancement in lipid productivity | 0.94 | Not mention | ||
| Cas9 | Vector driven/RNP | Electroporation | COP1/2, COP3, COP4, COP5, PHOT, UVR8, VGCC, MAT3, aCRY, PSY1 | Paromomycin | Protocols for the rapid isolation of non-selectable gene mutants | 5 to 15% | 2.5 × 10˗5 to 2.5 × 10-4 | ||
| Cas9 | Vector driven/RNP | Electroporation | FKB12, ALS, ARG | Rapamycin and zeocin | Gene-within-a-gene hybrid construct, composed of a Cas9 gene containing an artificial intron and an inserted sgRNA gene | Not mention | 3 × 10-8 | ||
| Cpf1 (Cas12a) | RNP | Electroporation | FKB12, CpFTSY, CpSRP43, PHT7 | Rapamycin and based on coloration and chlorophyll fluorescence | CRISPR/Cpf1-mediated DNA editing efficiencies increased 500-fold with the use of single-stranded oligodeoxynucleotides (ssODNs) | ∼10% | Not mention | ||
| Cas9 | RNP | Electroporation | PLA2 | Hygromycin | Lipid productivity in phospholipase A2 knockout mutants increased by 64.25%, reaching 80.92 g/L/d | Not mention | Not mention | ||
| Cas9 | RNP | Electroporation | ELT1 | Hygromycin | Total lipid accumulated up to 28% of dry biomass, with a 27.2% increase in C18:1 ratio | 0.1368 | Not mention | ||
| Cas9 | RNP | Electroporation | LCYE, ZEP | Hygromycin | Mutant with 60% higher zeaxanthin yield (5.24 mg/L) and content (7.28 mg/g) | Not mention | Not mention | ||
| Cas9 | RNP | Electroporation and glass beads | IFT81, FAP70, MOT17, CDPK13, CEP131 | Paromomycin | Development of CRISPR-based targeted insertional mutagenesis method (TIM) for C. reinhardtii | 40% to 95% | Not mention | ||
| Cas9 | RNP | Cell-penetrating peptide pVEC | Maa7, FKB12 | Auxotrophic selection using 5-FI (MAA7) and Rapamycin (FKB12) | Delivery of Cas9/sgRNA RNP into C. reinhardtii using cell-penetrating peptide pVEC | 8.41% to 46.56% | Not mention | ||
| Cas9 | RNP | Electroporation | PPX1, FTSY, WDTC1 | Paromomycin or oxyfluorfen | Generation of individual strains with precise mutations in multiple target genes | 1.3% (PPX1 and FTSY) and 0.8% (WDTC1) | Not mention | ||
| Cas9 | RNP | Electroporation | SPD1 | Paromomycin or hygromycin | Targeted knockout of CrSPD1 induces spermidine auxotroph, which could be used as a selectable marker in biotechnology | 10% to 66% | Not mention | ||
| Cas9 | RNP | Glass bead | NR | Hygromycin | Generation of mutants with a bacterial phytase gene cassette knocking into the NR gene | 0.1481 | Not mention | ||
| Cas9 | RNP | Electroporation | PSR1, CpFTSY | 30% starch | Generation of mutants with impaired extracellular phosphatase synthesis in response to Pi deprivation | 0.22 | Not mention | ||
| Cas9 | RNP | Electroporation | LCYE | Hygromycin | A 2.3-fold increase in astaxanthin accumulation in the ΔLCYE mutant | 0.0417 | Not mention | ||
| Nannochloropsis spp. | Cas9 | Vector driven | Electroporation | NR | Grow normally under NH4Cl but fail to grow under NaNO3 | Knockout of the NR gene | 0.1% to 1% | Not mention | |
| Cas9 | In vitro–synthesized gRNAs/Cas9-expressing line | Electroporation | Insertional mutagenesis of 18 gene of 20 transcription factors | Blasticidin or Hygromycin | Improvement in the total carbon-to-lipid ratio from 20% (wild type) to 40–55% (mutants) | 6.25 to 78% | Not mention | ||
| Cas9 | In vitro–synthesized gRNAs/reporter-free Cas9-expressing line | Electroporation | Aco1 | Blasticidin and Hygromycin | Mutants with doubled lipid productivity and ∼50% reduced photosynthetic antenna size | ∼50% | Not mention | ||
| Cas9 and Cas12a | RNP | Electroporation | NR | Zeocin | FnCas12a generated HDR-based mutants with up to 93% efficiency | 34–71% (for Cas9) and 3-93% (for Cas12a) | Not mention | ||
| Cas9 | In vitro–synthesized gRNAs/reporter-free Cas9- expressing line | Electroporation | Knock-in of FAD12 gene at the T1 hotspot | Zeocin | Improved production of polyunsaturated fatty acids (PUFAs) | 0.714 | Not mention | ||
| Cas12a | RNP | Electroporation | NR | Grow normally under NH4Cl but fail to grow under NaNO3 | Generation of markerless genome editing tool to knock out target genes | 0.1052 | Not mention | ||
| Cas9 | Vector driven | Electroporation | LER1 and LER2 | Hygromycin | Double deletion of both LER1 and LER2 (from chromosome 9), total ~214 kb | 0.285 | Not mention | ||
| dCas9 | Vector driven | Electroporation | g1248 | Zeocin | Growth and photosynthetic parameters (Fv/Fm) of the mutants increased by 23% and 12%, respectively, compared to the wild type under ambient CO2 levels | Not mention | Not mention | ||
| Cas9 | Vector driven | Bombardment | LSMT | Hygromycin | Mutation induced 18–20% reduction in fructose-1,6-bisphosphate aldolases, along with 9.7–13.8% increase in dry weight and enhanced growth | 0.1 | 2 × 10-8 | ||
| Phaeodactylum tricornutum | Cas9 | Vector driven | Bombardment | CpSRP54 | Zeocin | Generation of stable targeted gene mutations in marine algae | 0.31 | Not mention | |
| Cas9 | Vector-driven and RNP | Bombardment | PtUMPS, PtAPT, PtAureo1a | Nourseothricin, 2-FA, adenine, and uracil | A single-step generation of triple knockout strains | 65% to 100% | Not mention | ||
| Cas9 | Vector driven | Bacterial conjugation | Phatr3_J46193 | Phleomycin | Bacterial conjugation-mediated Cas9 delivery to minimize genome exposure to nuclease activity | Not mention | 2 × 10-5 | ||
| Cas9 | Vector driven | Electroporation | LACS | Zeocin | Reduced growth rate and altered molecular profiles of PC and TAGs | Not mention | Not mention | ||
| Cas9 | Vector driven | Conjugation of plasmids | CryP | Zeocin | Increased light-harvesting protein levels in CryP knockout mutants | Not mention | Not mention | ||
| Cas9 | Vector driven | Bombardment | PtTHIC and PtSSSP | Zeocin | Targeted mutation of the TPP aptamer in the THIC gene encoding HMP-P synthase does not affect thiamine biosynthesis in P. tricornutum | Not mention | Not mention | ||
| Cas9 | Vector driven | Bombardment | CpFTSY | Zeocin | Generation and characterization of CpFTSY mutants | Not mention | Not mention | ||
| Cas9 | Vector driven | Bombardment | FucT | Zeocin | Knockout of PtFucT1 affected PtGnTI activity in the complex, converting the N-glycan to a mannose-type N-glycan | 0.333 | Not mention | ||
| Cas9 | Vector driven | Electroporation | StLDP | Zeocin | StLDP functions as an LD scaffold protein in P. tricornutum, regulating LD numbers in the stldp mutant and complemented strains | 0.8125 | Not mention | ||
| Cas9 | Vector driven | Bacterial conjugation | ZEP 2, ZEP3 | No selective pressure | Generation of zep mutants as a platform for diatoxanthin production | Not mention | Not mention | ||
| Chlorella sp. | Cas9 | Vector driven | Electroporation | fad3 | Hygromycin | Mutants have 46% higher lipid accumulation | Not mention | Not mention | |
| Cas9 | Vector driven and RNP | Bombardment | NR, APT | NaNO2 and KClO3 | Generation of auxotrophic strains | Not mention | Not mention | ||
| CRISPRi and CRISPRa | Vector driven | Electroporation | Randomly mediate gene regulation | Hygromycin | Mutants with protein content of 60% to 65% (w/w) of dry cell weight | 0.5 | Not mention | ||
| Cas9 | Vector driven | Electroporation | GS | Hygromycin | Generation of mutants with enhanced biomass, protein, and lutein content | Not mention | Not mention | ||
| Cas9 | Vector driven | Electroporation | APT | Hygromycin | Combination of Alcalase treatment with PEG transformation for efficient gene editing in Chlorella | Not mention | 1 × 10-7 to 2 × 10-7 | ||
| Porphyridium purpureum | Cas9 | RNP | Bombardment | CHS1 | No selective pressure | Generation of chlorophyll synthase loss-of-function mutants with increased phycoerythrin levels | Not mention | Not mention | |
| Tetraselmis sp. | Cas9 | RNP | Bombardment | AGP | No selective pressure | Mutants had 2.7- and 3.1-fold increased lipid content (21.1% and 24.1% of DCW, respectively) | Not mention | ||
| Euglena gracilis | Cas9 | RNP | Electroporation | EgGSL2 | No selective pressure | Transgene-free targeted mutagenesis and ssODN-mediated gene knockin | 77.7 to 90.1% | Not mention | |
| LbCas12a | RNP | Electroporation | EgGSL2, EgcrtB | No selective pressure | High-efficiency genome editing system using direct delivery of LbCas12a RNP complexes | 77.2–94.5% | Not mention | ||
| Cas9 | RNP | Electroporation | Knockout of 16 carotenoid biosynthetic genes present in E. gracilis | No selective pressure | Mutants with different carotenoid compositions | Not mention | Not mention | ||
| Thalassiosira pseudonana | Cas9 | Vector driven | Bombardment | Urease | Nourseothricin | Significant reduction in growth rate and cell size compared to nitrate growth | 0.121 | 8 × 10-7 | |
| Cas9 | Vector driven | Bombardment | Sin1 | Nourseothricin | Mutants exhibit reduced biosilica content and morphological aberrations, affecting cell wall strength and stiffness | Not mention | Not mention | ||
| Cas9 nickase | Vector driven | Bombardment | TpθCA3 | Nourseothricin | Development of an efficient Cas9 nickase (D10A) system for highly specific indel introduction into target DNA | 0.39 | 3.4 × 10-7 | ||
| Components | Source/available toolkits | Company |
|---|---|---|
| Plasmids | Addgene (USA) | |
| SnapGene Plasmid Database (USA) | ||
| GenScript (USA) | ||
| IDT (USA) | ||
| Cas9 | ToolGen, Inc. (Korea) | |
| Macrogen, Inc. (Korea) | ||
| IDT (USA) | ||
| NEB (USA) | ||
| TaKaRa (Japan) | ||
| gRNA | MEGAshortscriptTM T7 Kit | Ambion (USA) |
| HiScribe T7 RNA Kit | NEB (USA) | |
| CUGA7 gRNA Synthesis Kit | Nippon Gene (Japan) | |
| EnGen® sgRNA Synthesis Kit | NEB (USA) | |
| Guide-itTM sgRNA In Vitro Transcription | TaKaRa (Japan) |
| Microalgal strains | Variants of Cas protein | Origin | Codon optimization | PAM | References |
|---|---|---|---|---|---|
| C. reinhardtii | SpCas9 | Streptococcus pyogenes | Yes | NGG | |
| C. reinhardtii | SaCas9 and SpCas9 | Staphylococcus aureus and S. pyogenes | Yes | NGG | |
| C. reinhardtii | SpCas9 | S. pyogenes | Yes | NGG | |
| C. reinhardtii | Dead SpCas9(dCas9) | S. pyogenes | Maize codon–optimized | NGG | |
| P. tricornutum | SpCas9 | S. pyogenes | Yes | NGG | |
| N. oceanica IMET1 | SpCas9 | S. pyogenes | Yes | NGG | |
| N. gaditana | SpCas9 | S. pyogenes | Yes | NGG | |
| N. salina | SpCas9 | S. pyogenes | Chlamydomonas-codon optimized | NGG | |
| N. oceanica | dCas9 | S. pyogenes | Yes | NGG | |
| T. pseudonana | SpCas9 | S. pyogenes | Human-codon optimized | NGG | |
| T. pseudonana | SpCas9 | S. pyogenes | Yes | NGG | |
| C. vulgaris | SpCas9 | S. pyogenes | Yes | NGG | |
| C. vulgaris | SpCas9 | S. pyogenes | Maize-codon optimized | NGG | |
| C. sorokiniana | dCas9 | S. pyogenes | Maize-codon optimized | NGG |
| Features | Microalgal strains | Host | References | |
|---|---|---|---|---|
| Cas9 | pCaMV 35S/tNOS | C. reinhardtii | Cauliflower mosaic virus (CaMV) | |
| C. vulgaris | ||||
| C. sorokiniana | ||||
| pHSP70A-RBCS2/RBCS2 3’UTR | C. reinhardtii | C. reinhardtii | ||
| pPsaD/tPsaD | C. reinhardtii | C. reinhardtii | ||
| pLHCF2/tLHCF1 | P. tricornutum | P. tricornutum | ||
| pVCP /tATUB | N. oceanica IMET1 | N. oceanica IMET1 | ||
| pRPL24/ tFRD | N. gaditana | N. gaditana | ||
| pRibi/tldsp | N. oceanica | N. oceanica | ||
| pTpFCP/tNAT | T. pseudonana | T. pseudonana | ||
| pNR (nitrate reductase promoter)/tNR (nitrate reductase terminator) | T. pseudonana | T. pseudonana | ||
| pFcpB/tFcpA | P. tricornutum | P. tricornutum | ||
| sgRNA | pU6/T6 | C. reinhardtii | Arabidopsis | |
| C. reinhardtii | C. reinhardtii | |||
| P. tricornutum | P. tricornutum | |||
| T. pseudonana | T. pseudonana | |||
| C. vulgaris | Arabidopsis | |||
| C. sorokiniana | Arabidopsis | |||
| pATPase/tfd | N. oceanica IMET1 | N. oceanica IMET1 | ||
| pPsaD/tPsaD | C. reinhardtii | C. reinhardtii | ||
| pRibi/tcs | N. oceanica | N. oceanica | ||
| Donor DNA | pβ-tub/trbcS2 (for HygR) | C. reinhardtii | C. reinhardtii | |
| pVCP/tVCP (for Sh bleR) | N. oceanica IMET1 | N. oceanica IMET1 | ||
| pβ-tub/tCOP1 (for aphVII) | C. reinhardtii | C. reinhardtii | ||
| pHSP70A/RBCS2 3’UTR (for bacterial phytase gene) | C. reinhardtii | C. reinhardtii | ||
| pTUB2/tCOP21 (for aphVIII) | C. reinhardtii | C. reinhardtii | ||
| Selection marker | pCaMV 35S/tNOS (for HygR) | C. reinhardtii | CaMV | |
| pFCP/pFCP (for Sh bleR) | P. tricornutum | P. tricornutum | ||
| pHSP70-RBCS2/RBCS2-3UTR (for aphVIII) | C. reinhardtii | C. reinhardtii | ||
| pPsaD/tPsaD (for Sh bleR) | C. reinhardtii | C. reinhardtii | ||
| pTCT/ tEIF3 (for blasticidin deaminase) | N. gaditana | N. gaditana | ||
| pEIF3/tFRD (for HygR) | N. gaditana | N. gaditana | ||
| pUEP/tUEP (for Sh bleR) | N. salina | N. salina |
Mutagenesis efficiency (%) was determined by calculating the proportion of mutants confirmed through genotyping methods, such as Sanger sequencing or polymerase chain reaction (PCR), against the total number of mutants subjected to genotyping. Targeted mutagenesis efficiency was defined by the number of mutants confirmed by genotyping in relation to the initial size of the mutant pool.
Table 1.
Table 2.
Table 3.
Table 4.
TOP
 MSK
MSK
 ePub Link
ePub Link Cite this Article
Cite this Article





