- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- For Contributors
- Policies
- E-Submission

- About
- Browse Articles
-
Special Issues
- Pioneering strategies for overcoming bacterial drug resistance (2026)
- Advancing microbial engineering through synthetic biology (2025)
- Host-associated microbiome (2024)
- Bacterial regulatory mechanisms for the control of complex cellular mechanisms (2023)
- Two years into COVID-19 pandemic: Where are we? (2022)
- Collections
- Policies
- For Contributors
Articles
- Page Path
- HOME > J. Microbiol > Volume 63(3); 2025 > Review
-
Review
Untranslated region engineering strategies for gene overexpression, fine-tuning, and dynamic regulation - Jun Ren†, So Hee Oh†, Dokyun Na*
-
Journal of Microbiology 2025;63(3):e2501033.
DOI: https://doi.org/10.71150/jm.2501033
Published online: March 28, 2025
Department of Biomedical Engineering, Chung-Ang University, Seoul 06974, Republic of Korea
- *Correspondence Dokyun Na blisszen@cau.ac.kr
- †These authors contributed equally to this work.
© The Microbiological Society of Korea
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
ABSTRACT
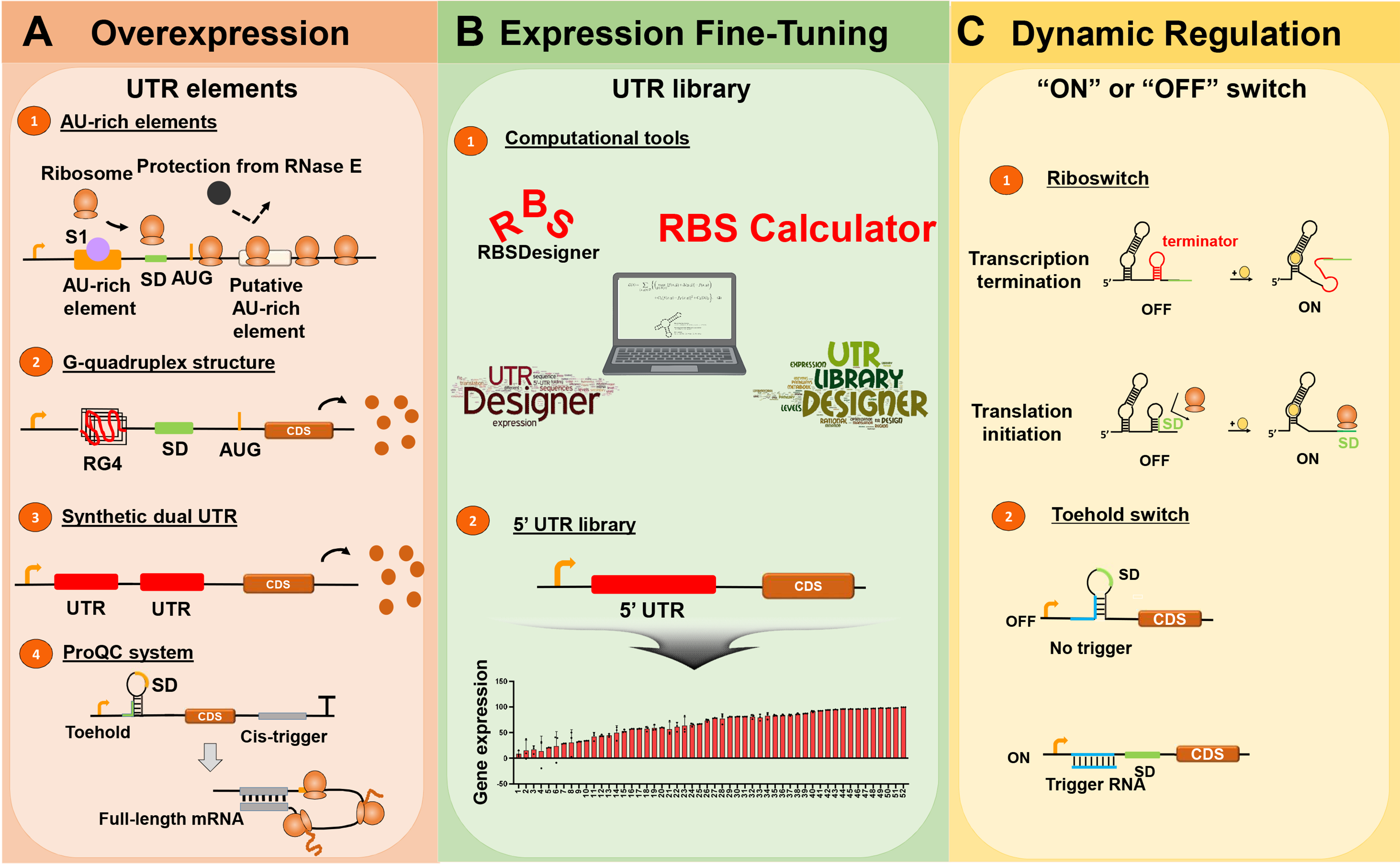
- Precise and tunable gene expression is crucial for various biotechnological applications, including protein overexpression, fine-tuned metabolic pathway engineering, and dynamic gene regulation. Untranslated regions (UTRs) of mRNAs have emerged as key regulatory elements that modulate transcription and translation. In this review, we explore recent advances in UTR engineering strategies for bacterial gene expression optimization. We discuss approaches for enhancing protein expression through AU-rich elements, RG4 structures, and synthetic dual UTRs, as well as ProQC systems that improve translation fidelity. Additionally, we examine strategies for fine-tuning gene expression using UTR libraries and synthetic terminators that balance metabolic flux. Finally, we highlight riboswitches and toehold switches, which enable dynamic gene regulation in response to environmental or metabolic cues. The integration of these UTR-based regulatory tools provides a versatile and modular framework for optimizing bacterial gene expression, enhancing metabolic engineering, and advancing synthetic biology applications.
Introduction
Overexpression strategies
Expression fine-tuning strategies
Dynamic gene regulation strategies
Conclusion
Conflict of Interest
The authors declare that they have no conflict of interest.
Funding
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (NRF-2023R1A2C1008156) and was also supported by the Chung-Ang University Research Grants in 2023.
| Expression | UTR elements | Description and applications | Ref |
|---|---|---|---|
| Protein over-expression | AU-rich elements | A well-known AU-rich element is originated from sodB gene. S1 protein stabilizes AU-rich mRNAs by recruiting ribosomes and preventing degradation. Knockdown of S1 reduced GFP mRNA by 34% and RFP mRNA by 61%. In its absence, Hfq protein compensates to protect mRNAs from degradation. | Lee et al. (2021b) |
| RG4 | The enhanced translation by an RG4 structure follows a physical barrier model, where bulky structures in the 5′ UTR guide ribosome movement toward the downstream start codon, increasing translation efficiency. | Lee et al. (2024) | |
| Key 5′ UTR features were optimized, including a 25–30 nt spacer between the stabilizer hairpin and RBS and 32% GC content in the spacer. Additional introduction of the RG4 RNA aptamer "Corn" into the 3′ UTR, significantly enhanced protein expression. Employing an optimal 5′ UTR increased recombinant luciferase protein expression by 1.8-fold. Similarly, modifying a gene with both the optimal 5′ UTR motif and the 3′ UTR motif (5×Corn) led to a 3.4-fold increase in recombinant Plasmodium falciparum lactate dehydrogenase (PfLDH) protein expression. | Sjekloća & Ferre-D’ Amaré (2019) | ||
| Synthetic dual UTRs | Two UTRs, enhancing transcription and translation, respectively, were identified from a library of <400,000 randomized UTRs in E. coli and concatenated. This synthetic dual UTR construct enhanced gene expression. The r31n47 dual UTR dramatically enhanced β-lactamase expression, compared with wild-type UTRs. For mCherry, transcript levels, fluorescence intensity, and half-life improved notably. Additionally, this dual UTR enhanced the solubility of both β-lactamase and mCherry proteins, further optimizing protein expression. | Lee et al. (2020) | |
| ProQC | The ProQC system was used to express vioB originated from C. violaceum, which converts L-tryptophan into violacein, a polyketide synthase product. This approach enhanced violacein production by 2.3-fold. Additionally, a synthetic fusion enzyme, CrtBI, was expressed using the ProQC system, while geranylgeranyl diphosphate synthase (crtE) was expressed, which resulted in a 2.2-fold increase in lycopene accumulation. | Yang et al. (2021b) | |
| Expression fine-optimization | RBS | A total of 41 synthetic 5′ UTRs were designed using the RBSCalculator, with translation initiation rates rationally controlled over a 100,000-fold range, enabling precise gene expression regulation. The 5′ UTR_33k_30 exhibited the highest mRNA increasing lacZ by 5-fold, txAbF by 6-fold, and msfGFP by 2.5-fold. | Chen et al. (2022a) |
| Optimal leader sequence was designed to minimize structure interference between SD and CDS. With the optimal leader sequence, RBS sequences were computationally designed using RBSDesigner to customize gene expression levels. Optimizing the expression levels of key metabolic enzymes encoded by cadA and PP3533 resulted in enhanced metabolite production. The expression-optimized strains produced 2.25 g/L of cadaverine (a 72% increase) and 2.59 g/L of L-proline (a 28% increase). | Yoo et al. (2020) | ||
| The shRBS library achieved a 10⁴-fold dynamic range in expression strength by modifying the spacer regions between the SD sequence and the start codon and in B. licheniformis. To assess the shRBS library for metabolic engineering, five shRBSs were selected to fine-tune leuS gene expression, promoting pulcherriminic acid production in B. licheniformis. Additionally, the rate-limiting enzyme YvmC was overexpressed. The production of pulcherriminic acid varied based on the strength of different shRBSs regulating leuS expression, achieving a two-fold range. The highest production of pulcherriminic acid was observed when U12-9 shRBS was utilized to leuS gene expression. | Rao et al. (2024) | ||
| A high-throughput screening method using 30–300 bp DNA fragments identified a novel 5′ UTR sequence from the B. subtilis genome, with the highest expression rate. Subsequently, a 5′ UTR library was constructed from the 5′ UTR by introducing mutations to diversify gene expression level. An artificial rib operon in B. subtilisi, in which the genes’ 5′ UTR were replaced with the high-efficient UTR. This artificial rib operon resulted in a 2.09-fold increase in riboflavin production. This enhancement was 4.7-fold higher compared with the operon with its original UTR sequences. | Liu et al. (2025) | ||
| A library of 90 promoter-UTRs was constructed based on RNA-Seq data analysis. The strongest promoter-UTR sequence, PNCgl1676-UTR, exhibited expression levels over five times higher than Psod-UTR, the most commonly used strong promoter-UTR in C. glutamicum. | Li et al. (2020) | ||
| A library of promoter–UTR sequences including RBS from various methanogens was constructed, which achieved a 140-fold dynamic range in expression strength in M. acetivorans. The strongest promoter-RBS was modified to generate six rationally designed high-expression 5′ -UTR variants. When evaluated using β-glucuronidase, they demonstrated a 140-fold range of expression strengths in M. acetivorans. | Zhu et al. (2024) | ||
| 5′ UTR libraries were designed using UTR Library Designer to diversify the expression levels of phlF and mcbR repressors. The expression levels of phlF and mcbR libraries were broadly diversified up to 18.57-fold and 15.14-fold, respectively. The most optimized strain using the libraries achieved a 2.82-fold increase in lycopene production compared with non-optimized strain. Additionally, a strain co-expressing phlF and mcbR with designed UTR sequences exhibited a 16.5-fold increase in 3-HP production compared with the parental strain. | Lee et al. (2021a) | ||
| 5′ UTR libraries of crtE, crtB, and crtI were generated using UTR Library Designer. For each gene, 16 different 5′ UTR sequences were designed. The highest lycopene titer was achieved by expressing crtE, crtB, and crtI, each using its optimally designed UTR sequence. | Yang et al. (2021b) | ||
| To fine-tune pckA expression, four 5′ UTR variants were generated using UTR Library Designer. The best UTR sequence, leading to fine-tuned pckA expression, resulted in a strain that exhibited a 49.8-fold increase in naringenin production compared with the non-optimized strain. | Kim et al. (2024) | ||
| Dynamic expression regulation | Riboswitch | Riboswitch is a regulatory element in 5’ UTR, controlling the expression of downstream coding sequence due to its structural change upon interaction with its ligand. | Jiang et al. (2024) |
| Two lysine riboswitches were utilized to activate and repress the expression of aspartate kinase III and homoserine dehydrogenase, respectively, in the lysine-producing strain C. glutamicum OW45. As a result, lysine production increased by 35% in strain QW48 (A263-lysC) and 43% in strain QW54 (R357-hom), compared with the parental strain QW45. | |||
| Toehold switches | Toehold switches are designed to detect specific RNA sequences, known as "triggers," with high sensitivity and specificity. Toehold switch was designed to detect coronavirus RNA, which acts like a trigger. The detection system implemented on a paper-cell free expression system could detect coronavirus. | Park & Lee (2021) |
| Expression | UTR elements | Advantages | Limitations | Ref |
|---|---|---|---|---|
| Protein Over-expression | AU-rich | Incorporating AU-rich elements into the 5′ UTR region can stabilize mRNA through S1 and Hfq proteins, thereby enhancing protein production. | The relatively long AU-rich element derived from the sodB gene, for example, may enhance RNase E accessibility, potentially reducing mRNA stability. | Lee et al. (2021b) |
| RG4 | RG4 structures act as internal ribosome entry sites, allowing translation initiation independent of a TSS. Furthermore, their incorporation into the 3′ UTR enhances mRNA stability and protects against RNase-mediated degradation. | The strong helical structure of RG4 complicates the incorporation of other elements like riboswitches into the 5' UTR, because ribonucleotides forming strong intramolecular interactions within the RG4 structure can also interact with other elements, disrupting their structures. | Lee et al. (2024) | |
| Synthetic dual UTRs | Dual UTRs elements enhance both transcription and translation with an optimal distance between two UTRs within the 5′ UTR region. | The concatenated UTRs may contain regulatory elements and thus the performance of optimized UTRs may vary under different environmental conditions or growth phases. | Lee et al. (2020) | |
| ProQC | The ProQC system ensures translation occurs only in the presence of full-length mRNAs by facilitating circularization through a toehold switch in the 5′ UTR and a cis-trigger sequence in the 3′ UTR. This mechanism enhances full-length translation, thereby improving protein quality in bacteria. | The incorporation of a toehold switch, forming a strong secondary structure, complicates its integration with other 5′ UTR elements. In addition, to further enhance full-length protein production, improving the circularization efficiency and stability of circular mRNA requires further investigation. | Yang et al. (2021b) | |
| Expression fine-optimization | RBS | A UTR library offers a diverse set of UTR sequences, including RBSs, that can be incorporated into the 5′ UTR region to fine-tune target gene expression by varying ribosome binding affinity. | UTR libraries are designed for static fine-tuning rather than dynamic regulation. | Hockenberry et al. (2018) |
| Dynamic expression regulation | Riboswitch | Riboswitches regulate gene expression by binding specific metabolites, switching expression "on" or "off," and controlling downstream gene activity. | Riboswitches may exhibit leaky expression in the OFF state or fail to achieve full activation in the ON state. | Olenginski et al. (2024) |
| Toehold switches | Toehold switches regulate gene expression by binding trigger RNAs, toggling expression "on" or "off," and controlling downstream gene activity. | The strong secondary structure of toehold switch complicates its integration with other 5′ UTR elements, and like riboswitches toehold switches often exhibit leaky expression in the OFF state or fail to achieve full activation in the ON state. Unintended interactions with endogenous RNAs can also lead to undesirable regulatory effects. | Chau et al. (2020) |
| Software | Description | Ref | Exemplar applications |
|---|---|---|---|
| RBSDesinger | RBSDesigner is a mathematical model accounting for the thermodynamic RBS folding (30 nt from SD) and the interaction of RBS with ribosomes. The thermodynamic parameters in the model were calculated using UNAFold software package. The model’s prediction performance was R2=0.77 – 0.87. | Na & Lee (2010) | The RBS of the ppc gene, optimized by RBSDesigner, enhanced both rapid cell growth and high 3-AP production in E. coli by facilitating the conversion of phosphoenolpyruvate to oxaloacetate, which is then converted to aspartate and ultimately to 3-AP (Song et al., 2015). |
| URL: http://ssbio.cau.ac.kr/web/?page_id=195 | |||
| RBSCalculator / Operon Calculator | RBSCalculator is a thermodynamic model composed of five energy parameters calculated from -35 to +35 nt of TIR by using NuPACK suite. Its prediction performance was R2=0.51 – 0.95 depending on the origin of UTR. | Salis (2011), Tian & Salis (2015) | Optimized expression of fabH and fabZ using RBS sequences designed by RBSCalculator enhanced the synthesis of fatty acyl-ACP from acetyl-CoA. Fatty acyl-ACP was then converted to fatty alcohol by the far gene, resulting in a high production titer of fatty alcohol (Chen et al., 2022b). |
| Operon Calculator is a biophysical model based on the model of RBSCalculator to predict the translation of bi- and tri-cistronic genes. Its performance is Pearson R2=0.57 – 0.91. | |||
| URL: https://salislab.net/software/predict_rbs_calculator, | |||
| http://salislab.net/software | |||
| UTRDesigner / UTR Library Designer | UTRDesigner is a thermodynamic model that accounts for several parameters, calculated from -10 to +35 nt of the TIR using NuPACK. Its performance is R² = 0.81. UTR Library Designer is a tool used to generate UTR sequences with diverse expression levels, utilizing UTRDesigner. | Seo et al. (2013, 2014) | For enhanced itaconic acid production, multiple genes (acs, gltA, and aceA) involved in acetate assimilation and the glyoxylate shunt pathway were overexpressed using optimized RBS sequences designed by UTRDesigner (Noh et al., 2018). |
| URL: https://sbi.postech.ac.kr/utr_designer/, https://sbi.postech.ac.kr/utr_library/ |
- Adamczyk PA, Reed JL. 2017. Escherichia coli as a model organism for systems metabolic engineering. Curr Opin Syst Biol. 6: 80–88. Article
- Al-Zeer MA, Dutkiewicz M, von Hacht A, Kreuzmann D, Rohrs V, et al. 2019. Alternatively spliced variants of the 5'-UTR of the ARPC2 mRNA regulate translation by an internal ribosome entry site (IRES) harboring a guanine-quadruplex motif. RNA Biol. 16(11): 1622–1632. ArticlePubMedPMC
- Arendt P, Pollier J, Callewaert N, Goossens A. 2016. Synthetic biology for production of natural and new-to-nature terpenoids in photosynthetic organisms. Plant J. 87(1): 16–37. ArticlePubMed
- Bahiri-Elitzur S, Tuller T. 2021. Codon-based indices for modeling gene expression and transcript evolution. Comput Struct Biotechnol J. 19: 2646–2663. ArticlePubMedPMC
- Baker KE, Mackie GA. 2003. Ectopic RNase E sites promote bypass of 5'-end-dependent mRNA decay in Escherichia coli. Mol Microbiol. 47(1): 75–88. ArticlePubMedLink
- Barrick JE, Breaker RR. 2007. The distributions, mechanisms, and structures of metabolite-binding riboswitches. Genome Biol. 8(11): R239.ArticlePubMedPMCPDF
- Bhattacharyya D, Diamond P, Basu S. 2015. An independently folding RNA G-quadruplex domain directly recruits the 40S ribosomal subunit. Biochemistry. 54(10): 1879–1885. ArticlePubMed
- Boussebayle A, Torka D, Ollivaud S, Braun J, Bofill-Bosch C, et al. 2019. Next-level riboswitch development—implementation of Capture-SELEX facilitates identification of a new synthetic riboswitch. Nucleic Acids Res. 47(9): 4883–4895. ArticlePubMedPMC
- Bu F, Lin X, Liao W, Lu Z, He Y, et al. 2024. Ribocentre-switch: a database of riboswitches. Nucleic Acids Res. 52(D1): D265–D272. ArticlePubMedPDF
- Buccitelli C, Selbach M. 2020. mRNAs, proteins and the emerging principles of gene expression control. Nat Rev Genet. 21(10): 630–644. ArticlePubMedPDF
- Cambray G, Guimaraes JC, Arkin AP. 2018. Evaluation of 244,000 synthetic sequences reveals design principles to optimize translation in Escherichia coli. Nat Biotechnol. 36(10): 1005–1015. ArticlePubMedPDF
- Cammas A, Dubrac A, Morel B, Lamaa A, Touriol C, et al. 2015. Stabilization of the G-quadruplex at the VEGF IRES represses cap-independent translation. RNA Biol. 12(3): 320–329. ArticlePubMed
- Ceres P, Trausch JJ, Batey RT. 2013. Engineering modular 'ON' RNA switches using biological components. Nucleic Acids Res. 41(22): 10449–10461. ArticlePubMedPMCPDF
- Chau THT, Mai DHA, Pham DN, Le HTQ, Lee EY. 2020. Developments of riboswitches and toehold switches for molecular detection—biosensing and molecular diagnostics. Int J Mol Sci. 21(9): 3192.ArticlePubMedPMC
- Chelkowska-Pauszek A, Kosinski JG, Marciniak K, Wysocka M, Bakowska-Zywicka K, et al. 2021. The role of RNA secondary structure in regulation of gene expression in bacteria. Int J Mol Sci. 22(15): 7845.ArticlePubMedPMC
- Chen F, Cocaign-Bousquet M, Girbal L, Nouaille S. 2022a. 5'UTR sequences influence protein levels in Escherichia coli by regulating translation initiation and mRNA stability. Front Microbiol. 13: 1088941.ArticlePubMed
- Chen RN, Liu YL, Zhong W, Hao XM, Mu TZ, et al. 2022b. Ribosome-binding sequences (RBS) engineering of key genes for high production of fatty alcohols. Biotechnol Bioprocess Eng. 27(4): 615–623. Article
- Choe D, Kim K, Kang M, Lee SG, Cho S, et al. 2022. Synthetic 3'-UTR valves for optimal metabolic flux control in Escherichia coli. Nucleic Acids Res. 50(7): 4171–4186. ArticlePubMedPMCPDF
- Delvillani F, Papiani G, Deho G, Briani F. 2011. S1 ribosomal protein and the interplay between translation and mRNA decay. Nucleic Acids Res. 39(17): 7702–7715. ArticlePubMedPMC
- Deng J, Wu Y, Zheng Z, Chen N, Luo X, et al. 2021. A synthetic promoter system for well-controlled protein expression with different carbon sources in Saccharomyces cerevisiae. Microb Cell Fact. 20(1): 202.ArticlePubMedPMCPDF
- Ding K, Dixit G, Parker BJ, Wen J. 2023. CRMnet: A deep learning model for predicting gene expression from large regulatory sequence datasets. Front Big Data. 6: 1113402.ArticlePubMedPMC
- Dong H, Nilsson L, Kurland CG. 1995. Gratuitous overexpression of genes in Escherichia coli leads to growth inhibition and ribosome destruction. J Bacteriol. 177(6): 1497–1504. ArticlePubMedPMCLink
- Du F, Liu YQ, Xu YS, Li ZJ, Wang YZ, et al. 2021. Regulating the T7 RNA polymerase expression in E. coli BL21 (DE3) to provide more host options for recombinant protein production. Microb Cell Fact. 20(1): 189.ArticlePubMedPMC
- Emory SA, Bouvet P, Belasco JG. 1992. A 5'-terminal stem loop structure can stabilize mRNA in Escherichia coli. Genes Dev. 6(1): 135–148. ArticlePubMed
- Goldman SR, Ebright RH, Nickels BE. 2009. Direct detection of abortive RNA transcripts in vivo. Science. 324(5929): 927–928. ArticlePubMedPMC
- Hockenberry AJ, Stern AJ, Amaral LAN, Jewett MC. 2018. Diversity of translation initiation mechanisms across bacterial species is driven by environmental conditions and growth demands. Mol Biol Evol. 35(3): 582–592. ArticlePubMed
- Hoetzel J, Suess B. 2022. Structural changes in aptamers are essential for synthetic riboswitch engineering. J Mol Biol. 434(18): 167631.ArticlePubMed
- Hong F, Ma D, Wu K, Mina LA, Luiten RC, et al. 2020. Precise and programmable detection of mutations using ultraspecific riboregulators. Cell. 183(3): 835–836. ArticlePubMedPMC
- Hook-Barnard IG, Brickman TJ, McIntosh MA. 2007. Identification of an AU-rich translational enhancer within the Escherichia coli fepB leader RNA. J Bacteriol. 189(11): 4028–4037. ArticlePubMedPMCLink
- Hsu LM, Cobb IM, Ozmore JR, Khoo M, Nahm G, et al. 2006. Initial transcribed sequence mutations specifically affect promoter escape properties. Biochemistry. 45(29): 8841–8854. ArticlePubMed
- Inui M, Kawaguchi H, Murakami S, Vertes AA, Yukawa H. 2004. Metabolic engineering of Corynebacterium glutamicum for fuel ethanol production under oxygen-deprivation conditions. J Mol Microbiol Biotechnol. 8(4): 243–254. ArticlePubMed
- Jiang Q, Geng F, Shen J, Zhu P, Lu Z, et al. 2024. Engineering a Lactobacillus lysine riboswitch to dynamically control metabolic pathways for lysine production in Corynebacterium glutamicum. Microorganisms. 12(3): 606.ArticlePubMedPMC
- Kallscheuer N, Marienhagen J. 2018. Corynebacterium glutamicum as platform for the production of hydroxybenzoic acids. Microb Cell Fact. 17: 1–13. ArticlePubMedPMC
- Kavita K, Breaker RR. 2023. Discovering riboswitches: the past and the future. Trends Biochem Sci. 48(2): 119–141. ArticlePubMed
- Kent R, Dixon N. 2019. Systematic evaluation of genetic and environmental factors affecting performance of translational riboswitches. ACS Synth Biol. 8(4): 884–901. ArticlePubMedPMCLink
- Kim DH, Hwang HG, Ye DY, Jung GY. 2024. Transcriptional and translational flux optimization at the key regulatory node for enhanced production of naringenin using acetate in engineered Escherichia coli. J Ind Microbiol Biotechnol. 51: kuae006.ArticlePubMedPMCPDF
- Kim J, Zhou Y, Carlson PD, Teichmann M, Chaudhary S, et al. 2019. De novo-designed translation-repressing riboregulators for multi-input cellular logic. Nat Chem Biol. 15(12): 1173–1182. ArticlePubMedPMCPDF
- Kondo T, Shimizu T. 2023. Translation enhancement by a short nucleotide insertion at 5′ UTR: application to an in vitro cell-free system and a photosynthetic bacterium. Appl Microbiol. 3(3): 687–697. Article
- Kosuri S, Goodman DB, Cambray G, Mutalik VK, Gao Y, et al. 2013. Composability of regulatory sequences controlling transcription and translation in Escherichia coli. Proc Natl Acad Sci USA. 110(34): 14024–14029. ArticlePubMedPMC
- Kozak M. 2005. Regulation of translation via mRNA structure in prokaryotes and eukaryotes. Gene. 361: 13–37. ArticlePubMed
- Kraus L, Duchardt-Ferner E, Brauchle E, Furbacher S, Kelvin D, et al. 2023. Development of a novel tobramycin dependent riboswitch. Nucleic Acids Res. 51(20): 11375–11385. ArticlePubMedPMCPDF
- Kudla G, Murray AW, Tollervey D, Plotkin JB. 2009. Coding-sequence determinants of gene expression in Escherichia coli. Science. 324(5924): 255–258. ArticlePubMedPMC
- Le SB, Onsager I, Lorentzen JA, Lale R. 2020. Dual UTR-A novel 5′ untranslated region design for synthetic biology applications. Synth Biol. 5(1): ysaa006.Article
- Lee JY, Cha S, Lee JH, Lim HG, Noh MH, et al. 2021a. Plug-in repressor library for precise regulation of metabolic flux in Escherichia coli. Metab Eng. 67: 365–372. ArticlePubMed
- Lee CY, Joshi M, Wang A, Myong S. 2024. 5'UTR G-quadruplex structure enhances translation in size dependent manner. Nat Commun. 15(1): 3963.ArticlePubMedPMCPDF
- Lee CY, McNerney C, Ma K, Zhao W, Wang A, et al. 2020. R-loop induced G-quadruplex in non-template promotes transcription by successive R-loop formation. Nat Commun. 11(1): 3392.ArticlePubMedPMCPDF
- Lee HM, Ren J, Kim WY, Vo PNL, Eyun SI, et al. 2021b. Introduction of an AU-rich element into the 5' UTR of mRNAs enhances protein expression in by S1 protein and Hfq protein. Biotechnol Bioprocess Eng. 26(5): 749–757. Article
- Lee HM, Ren J, Yu MS, Kim H, Kim WY, et al. 2021c. Construction of a tunable promoter library to optimize gene expression in Methylomonas sp. DH-1, a methanotroph, and its application to cadaverine production. Biotechnol Biofuels. 14: ArticlePDF
- Li M, Borodina I. 2015. Application of synthetic biology for production of chemicals in yeast Saccharomyces cerevisiae. FEMS Yeast Res. 15(1): 1–12. ArticlePMC
- Li N, Zeng W, Xu S, Zhou J. 2020. Obtaining a series of native gradient promoter-5'-UTR sequences in Corynebacterium glutamicum ATCC 13032. Microb Cell Fact. 19: 120.ArticlePubMedPMCPDF
- Lin J, Luo R, Pinello L. 2024. EPInformer: a scalable deep learning framework for gene expression prediction by integrating promoter-enhancer sequences with multimodal epigenomic data. bioRxiv. 2024.08.01.606099. Article
- Liu Y, Chen S, Chen J, Zhou J, Wang Y, et al. 2016. High production of fatty alcohols in Escherichia coli with fatty acid starvation. Microb Cell Fact. 15(1): 129.ArticlePubMedPMC
- Liu M, Jin Z, Xiang Q, He H, Huang Y, et al. 2024. Rational design of untranslated regions to enhance gene expression. J Mol Biol. 436(22): 168804.ArticlePubMed
- Liu YJ, Wang X, Sun Y, Feng Y. 2025. Bacterial 5' UTR: A treasure-trove for post-transcriptional regulation. Biotechnol Adv. 78: 108478.ArticlePubMed
- Lynch JP, Gonzalez-Prieto C, Reeves AZ, Bae S, Powale U, et al. 2023. Engineered Escherichia coli for the in situ secretion of therapeutic nanobodies in the gut. Cell Host Microbe. 31(4): 634–649. ArticlePubMedPMC
- Mandal M, Breaker RR. 2004. Gene regulation by riboswitches. Nat Rev Mol Cell Biol. 5(6): 451–463. ArticlePubMedPDF
- Matsuda Y, Itaya H, Kitahara Y, Theresia NM, Kutukova EA, et al. 2014. Double mutation of cell wall proteins CspB and PBP1a increases secretion of the antibody Fab fragment from Corynebacterium glutamicum. Microb Cell Fact. 13(1): 56.ArticlePubMedPMCPDF
- Mutalik VK, Guimaraes JC, Cambray G, Lam C, Christoffersen MJ, et al. 2013. Precise and reliable gene expression via standard transcription and translation initiation elements. Nat Methods. 10(4): 354–360. ArticlePubMedPDF
- Na D, Lee D. 2010. RBSDesigner: software for designing synthetic ribosome binding sites that yields a desired level of protein expression. Bioinformatics. 26(20): 2633–2634. ArticlePubMedPDF
- Na D, Lee S, Lee D. 2010. Mathematical modeling of translation initiation for the estimation of its efficiency to computationally design mRNA sequences with desired expression levels in prokaryotes. BMC Syst Biol. 4: 71.ArticlePubMedPMCPDF
- Nahvi A, Sudarsan N, Ebert MS, Zou X, Brown KL, et al. 2002. Genetic control by a metabolite binding mRNA. Chem Biol. 9(9): 1043.ArticlePubMed
- Noh MH, Lim HG, Woo SH, Song J, Jung GY. 2018. Production of itaconic acid from acetate by engineering acid-tolerant Escherichia coli W. Biotechnol Bioeng. 115(3): 729–738. ArticlePubMedLink
- Olenginski LT, Spradlin SF, Batey RT. 2024. Flipping the script: Understanding riboswitches from an alternative perspective. J Biol Chem. 300(3): 105730.ArticlePubMedPMC
- Pang Q, Han H, Liu X, Wang Z, Liang Q, et al. 2020. In vivo evolutionary engineering of riboswitch with high-threshold for N-acetylneuraminic acid production. Metab Eng. 59: 36–43. ArticlePubMed
- Pardee K, Green AA, Takahashi MK, Braff D, Lambert G, et al. 2016a. Rapid, low-cost detection of Zika virus using programmable biomolecular components. Cell. 165: 1255–1266. ArticlePubMed
- Pardee K, Slomovic S, Nguyen PQ, Lee JW, Donghia N, et al. 2016b. Portable, on-demand biomolecular manufacturing. Cell. 167(1): 248–259. ArticlePubMed
- Park S, Lee JW. 2021. Detection of coronaviruses using RNA toehold switch sensors. Int J Mol Sci. 22(4): 1772.ArticlePubMedPMC
- Pavlova N, Penchovsky R. 2022. Bioinformatics and genomic analyses of the suitability of eight riboswitches for antibacterial drug targets. Antibiotics (Basel). 11(9): 1177.ArticlePubMedPMC
- Pontrelli S, Chiu TY, Lan EI, Chen FYH, Chang P, et al. 2018. Escherichia coli as a host for metabolic engineering. Metab Eng. 50: 16–46. ArticlePubMed
- Proshkin S, Rahmouni AR, Mironov A, Nudler E. 2010. Cooperation between translating ribosomes and RNA polymerase in transcription elongation. Science. 328(5977): 504–508. ArticlePubMedPMC
- Rao X, Li D, Su Z, Nomura CT, Chen S, et al. 2024. A smart RBS library and its prediction model for robust and accurate fine-tuning of gene expression in Bacillus species. Metab Eng. 81: 1–9. ArticlePubMed
- Ren J, Shen J, Thai TD, Kim MG, Lee SH, et al. 2023. Evaluation of various Escherichia coli strains for enhanced lycopene production. J Microbiol Biotechnol. 33(7): 973–979. ArticlePubMedPMC
- Rocha EP, Danchin A, Viari A. 1999. Translation in Bacillus subtilis: roles and trends of initiation and termination, insights from a genome analysis. Nucleic Acids Res. 27(17): 3567–3576. ArticlePubMedPMC
- Rode AB, Endoh T, Sugimoto N. 2015. Tuning riboswitch-mediated gene regulation by rational control of aptamer ligand binding properties. Angew Chem Int Ed Engl. 54(3): 905–909. ArticlePubMed
- Saleski TE, Chung MT, Carruthers DN, Khasbaatar A, Kurabayashi K, et al. 2021. Optimized gene expression from bacterial chromosome by high-throughput integration and screening. Sci Adv. 7(7): eabe1767. ArticlePubMedPMC
- Salis HM. 2011. The ribosome binding site calculator. Methods Enzymol. 498: 19–42. ArticlePubMed
- Salis HM, Mirsky EA, Voigt CA. 2009. Automated design of synthetic ribosome binding sites to control protein expression. Nat Biotechnol. 27(10): 946–950. ArticlePubMedPMCPDF
- Salvail H, Breaker RR. 2023. Riboswitches. Curr Biol. 33(9): R343–R348. ArticlePubMedPMC
- Schneider J, Wendisch VF. 2010. Putrescine production by engineered Corynebacterium glutamicum. Appl Microbiol Biotechnol. 88(4): 859–868. ArticlePubMedPDF
- Seo SW, Yang JS, Cho HS, Yang J, Kim SC, et al. 2014. Predictive combinatorial design of mRNA translation initiation regions for systematic optimization of gene expression levels. Sci Rep. 4: ArticlePDF
- Seo SW, Yang JS, Kim I, Yang J, Min BE, et al. 2013. Predictive design of mRNA translation initiation region to control prokaryotic translation efficiency. Metab Eng. 15: 67–74. ArticlePubMed
- Sharma S, Pathania S, Bhagta S, Kaushal N, Bhardwaj S, et al. 2024. Microbial remediation of polluted environment by using recombinant E. coli: a review. Biotechnol Environ. 1(1): 8.ArticlePDF
- Sjekloća L, Ferré-D’Amaré AR. 2019. Binding between G quadruplexes at the homodimer interface of the Corn RNA aptamer strongly activates Thioflavin T fluorescence. Cell Chem Biol. 26: 1159–1168. ArticlePubMedPMC
- Song CW, Lee J, Ko YS, Lee SY. 2015. Metabolic engineering of Escherichia coli for the production of 3-aminopropionic acid. Metab Eng. 30: 121–129. ArticlePubMed
- Sung M, Yoo SM, Jun R, Lee JE, Lee SY, et al. 2016. Optimization of phage λ promoter strength for synthetic small regulatory RNA-based metabolic engineering. Biotechnol Bioprocess Eng. 21: 483–490. ArticlePDF
- Takahashi MK, Tan X, Dy AJ, Braff D, Akana RT, et al. 2018. A low-cost paper-based synthetic biology platform for analyzing gut microbiota and host biomarkers. Nat Commun. 9(1): 3347.ArticlePubMedPMCPDF
- Thai TD, Lim W, Na D. 2023. Synthetic bacteria for the detection and bioremediation of heavy metals. Front Bioeng Biotechnol. 11: 1178680.ArticlePubMedPMC
- Thavarajah W, Silverman AD, Verosloff MS, Kelley-Loughnane N, Jewett MC, et al. 2020. Point-of-use detection of environmental fluoride via a cell-free riboswitch-based biosensor. ACS Synth Biol. 9(1): 10–18. ArticlePubMedLink
- Tian T, Salis HM. 2015. A predictive biophysical model of translational coupling to coordinate and control protein expression in bacterial operons. Nucleic Acids Res. 43(14): 7137–7151. ArticlePubMedPMC
- Tietze L, Lale R. 2021. Importance of the 5′ regulatory region to bacterial synthetic biology applications. Microb Biotechnol. 14(6): 2291–2315. ArticlePubMedPMCLink
- Topp S, Gallivan JP. 2010. Emerging applications of riboswitches in chemical biology. ACS Chem Biol. 5(1): 139–148. ArticlePubMedPMC
- Tran KM, Lee HM, Thai TD, Shen J, Eyun SI, et al. 2021. Synthetically engineered microbial scavengers for enhanced bioremediation. J Hazard Mater. 419: 126516.ArticlePubMed
- Vikram, Mishra V, Rana A, Ahire JJ. 2022. Riboswitch-mediated regulation of riboflavin biosynthesis genes in prokaryotes. 3 Biotech. 12(10): 278.ArticlePubMedPMC
- Vind J, Sørensen MA, Rasmussen MD, Pedersen S. 1993. Synthesis of proteins in Escherichia coli is limited by the concentration of free ribosomes: expression from reporter genes does not always reflect functional mRNA levels. J Mol Biol. 231(3): 678–688. ArticlePubMed
- Vo P, Lee HM, Na D. 2019. Synthetic bacteria for therapeutics. J Microbiol Biotechnol. 29(6): 845–855. ArticlePubMed
- Walbrun A, Wang TH, Matthies M, Sulc P, Simmel FC, et al. 2024. Single-molecule force spectroscopy of toehold-mediated strand displacement. Nat Commun. 15(1): 7564.ArticlePubMedPMCPDF
- Wang T, Simmel FC. 2022. Riboswitch-inspired toehold riboregulators for gene regulation in Escherichia coli. Nucleic Acids Res. 50(8): 4784–4798. ArticlePubMedPMCPDF
- Wang X, Wei W, Zhao J. 2021. Using a riboswitch sensor to detect Co2+/Ni2+ transport in E. coli. Front Chem. 9: 631909.ArticlePubMedPMC
- Wanrooij PH, Uhler JP, Shi Y, Westerlund F, Falkenberg M, et al. 2012. A hybrid G-quadruplex structure formed between RNA and DNA explains the extraordinary stability of the mitochondrial R-loop. Nucleic Acids Res. 40(20): 10334–10344. ArticlePubMedPMC
- Watts A, Sankaranarayanan S, Watts A, Raipuria RK. 2021. Optimizing protein expression in heterologous systems: Strategies and tools. Meta Gene. 29: 100899.Article
- Wen JD, Kuo ST, Chou HD. 2021. The diversity of Shine-Dalgarno sequences sheds light on the evolution of translation initiation. RNA Biol. 18(11): 1489–1500. ArticlePubMed
- Winkler WC, Breaker RR. 2005. Regulation of bacterial gene expression by riboswitches. Annu Rev Microbiol. 59: 487–517. ArticlePubMed
- Winkler W, Nahvi A, Breaker RR. 2002. Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression. Nature. 419(6910): 952–956. ArticlePubMed
- Xiao J, Peng B, Su Z, Liu A, Hu Y, et al. 2020. Facilitating protein expression with portable 5'-UTR secondary structures in Bacillus licheniformis. ACS Synth Biol. 9(5): 1051–1058. ArticlePubMed
- Xu X, Qi LS. 2019. A CRISPR-dCas toolbox for genetic engineering and synthetic biology. J Mol Biol. 431(1): 34–47. ArticlePubMed
- Yang J, Han YH, Im J, Seo SW. 2021a. Synthetic protein quality control to enhance full-length translation in bacteria. Nat Chem Biol. 17(4): 421–427. ArticlePubMedPDF
- Yang D, Park SY, Lee SY. 2021b. Production of rainbow colorants by metabolically engineered Escherichia coli. Adv Sci. 8(13): e2100743. ArticleLink
- Yang ST, Zhang XF. 2023. ENGEP: advancing spatial transcriptomics with accurate unmeasured gene expression prediction. Genome Biol. 24(1): 293.ArticlePubMedPMCPDF
- Yim SS, An SJ, Choi JW, Ryu AJ, Jeong KJ. 2014. High-level secretory production of recombinant single-chain variable fragment (scFv) in Corynebacterium glutamicum. Appl Microbiol Biotechnol. 98(1): 273–284. ArticlePubMedPDF
- Yoo SM, Jung SW, Yeom J, Lee SY, Na D. 2020. Tunable gene expression system independent of downstream coding sequence. ACS Synth Biol. 9(11): 2998–3007. ArticlePubMed
- Yu S, Zheng B, Chen Z, Huo YX. 2021. Metabolic engineering of Corynebacterium glutamicum for producing branched-chain amino acids. Microb Cell Fact. 20(1): 230.ArticlePubMedPMCPDF
- Zhang Q, Ma D, Wu F, Standage-Beier K, Chen X, et al. 2021. Predictable control of RNA lifetime using engineered degradation-tuning RNAs. Nat Chem Biol. 17(7): 828–836. ArticlePubMedPMCPDF
- Zhao Y, Zhang JY, Zhang ZY, Tong TJ, Hao YH, et al. 2017. Real-time detection reveals responsive cotranscriptional formation of persistent intramolecular DNA and intermolecular DNA:RNA hybrid G-quadruplexes stabilized by R-loop. Anal Chem. 89(11): 6036–6042. ArticlePubMed
- Zhu P, Molina Resendiz M, von Ossowski I, Scheller S. 2024. A promoter-RBS library for fine-tuning gene expression in Methanosarcina acetivorans. Appl Environ Microbiol. 90(9): e0109224. ArticlePubMedLink
References
Supplementary Information
References
Citations

- Rhodo-Box: A Synthetic Biology Toolbox to Facilitate Metabolic Engineering of Rhodobacter sphaeroides
Matic Kostanjšek, Antoine Raynal, George Dimopoulos, Gerrich Behrendt, Vitor A. P. Martins dos Santos, Jules Beekwilder, Christos Batianis, Ruud A. Weusthuis, Enrique Asin-Garcia, Markus M. M. Bisschops
ACS Synthetic Biology.2026; 15(4): 1400. CrossRef - Production of the recombinant spider silk MaSp2 protein using the marine purple photosynthetic nonsulfur bacterium Rhodovulum sulfidophilum under autotrophic conditions
Miki Suzuki, Keiji Numata
NPG Asia Materials.2026;[Epub] CrossRef - Advancing microbial engineering through synthetic biology
Ki Jun Jeong
Journal of Microbiology.2025; 63(3): e2503100. CrossRef - Recombinase-Mediated Cassette Exchange-Based CRISPR Activation Screening Identifies Hyperosmotic Stress-Resistant Genes in Chinese Hamster Ovary Cells
Minhye Baek, Seokchan Kweon, Yujin Kim, Nathan E. Lewis, Jae Seong Lee, Gyun Min Lee
ACS Synthetic Biology.2025; 14(8): 3116. CrossRef

Fig. 1.
| Expression | UTR elements | Description and applications | Ref |
|---|---|---|---|
| Protein over-expression | AU-rich elements | A well-known AU-rich element is originated from sodB gene. S1 protein stabilizes AU-rich mRNAs by recruiting ribosomes and preventing degradation. Knockdown of S1 reduced GFP mRNA by 34% and RFP mRNA by 61%. In its absence, Hfq protein compensates to protect mRNAs from degradation. | |
| RG4 | The enhanced translation by an RG4 structure follows a physical barrier model, where bulky structures in the 5′ UTR guide ribosome movement toward the downstream start codon, increasing translation efficiency. | ||
| Key 5′ UTR features were optimized, including a 25–30 nt spacer between the stabilizer hairpin and RBS and 32% GC content in the spacer. Additional introduction of the RG4 RNA aptamer "Corn" into the 3′ UTR, significantly enhanced protein expression. Employing an optimal 5′ UTR increased recombinant luciferase protein expression by 1.8-fold. Similarly, modifying a gene with both the optimal 5′ UTR motif and the 3′ UTR motif (5×Corn) led to a 3.4-fold increase in recombinant Plasmodium falciparum lactate dehydrogenase (PfLDH) protein expression. | |||
| Synthetic dual UTRs | Two UTRs, enhancing transcription and translation, respectively, were identified from a library of <400,000 randomized UTRs in E. coli and concatenated. This synthetic dual UTR construct enhanced gene expression. The r31n47 dual UTR dramatically enhanced β-lactamase expression, compared with wild-type UTRs. For mCherry, transcript levels, fluorescence intensity, and half-life improved notably. Additionally, this dual UTR enhanced the solubility of both β-lactamase and mCherry proteins, further optimizing protein expression. | ||
| ProQC | The ProQC system was used to express vioB originated from C. violaceum, which converts L-tryptophan into violacein, a polyketide synthase product. This approach enhanced violacein production by 2.3-fold. Additionally, a synthetic fusion enzyme, CrtBI, was expressed using the ProQC system, while geranylgeranyl diphosphate synthase (crtE) was expressed, which resulted in a 2.2-fold increase in lycopene accumulation. | ||
| Expression fine-optimization | RBS | A total of 41 synthetic 5′ UTRs were designed using the RBSCalculator, with translation initiation rates rationally controlled over a 100,000-fold range, enabling precise gene expression regulation. The 5′ UTR_33k_30 exhibited the highest mRNA increasing lacZ by 5-fold, txAbF by 6-fold, and msfGFP by 2.5-fold. | |
| Optimal leader sequence was designed to minimize structure interference between SD and CDS. With the optimal leader sequence, RBS sequences were computationally designed using RBSDesigner to customize gene expression levels. Optimizing the expression levels of key metabolic enzymes encoded by cadA and PP3533 resulted in enhanced metabolite production. The expression-optimized strains produced 2.25 g/L of cadaverine (a 72% increase) and 2.59 g/L of L-proline (a 28% increase). | |||
| The shRBS library achieved a 10⁴-fold dynamic range in expression strength by modifying the spacer regions between the SD sequence and the start codon and in B. licheniformis. To assess the shRBS library for metabolic engineering, five shRBSs were selected to fine-tune leuS gene expression, promoting pulcherriminic acid production in B. licheniformis. Additionally, the rate-limiting enzyme YvmC was overexpressed. The production of pulcherriminic acid varied based on the strength of different shRBSs regulating leuS expression, achieving a two-fold range. The highest production of pulcherriminic acid was observed when U12-9 shRBS was utilized to leuS gene expression. | |||
| A high-throughput screening method using 30–300 bp DNA fragments identified a novel 5′ UTR sequence from the B. subtilis genome, with the highest expression rate. Subsequently, a 5′ UTR library was constructed from the 5′ UTR by introducing mutations to diversify gene expression level. An artificial rib operon in B. subtilisi, in which the genes’ 5′ UTR were replaced with the high-efficient UTR. This artificial rib operon resulted in a 2.09-fold increase in riboflavin production. This enhancement was 4.7-fold higher compared with the operon with its original UTR sequences. | |||
| A library of 90 promoter-UTRs was constructed based on RNA-Seq data analysis. The strongest promoter-UTR sequence, PNCgl1676-UTR, exhibited expression levels over five times higher than Psod-UTR, the most commonly used strong promoter-UTR in C. glutamicum. | |||
| A library of promoter–UTR sequences including RBS from various methanogens was constructed, which achieved a 140-fold dynamic range in expression strength in M. acetivorans. The strongest promoter-RBS was modified to generate six rationally designed high-expression 5′ -UTR variants. When evaluated using β-glucuronidase, they demonstrated a 140-fold range of expression strengths in M. acetivorans. | |||
| 5′ UTR libraries were designed using UTR Library Designer to diversify the expression levels of phlF and mcbR repressors. The expression levels of phlF and mcbR libraries were broadly diversified up to 18.57-fold and 15.14-fold, respectively. The most optimized strain using the libraries achieved a 2.82-fold increase in lycopene production compared with non-optimized strain. Additionally, a strain co-expressing phlF and mcbR with designed UTR sequences exhibited a 16.5-fold increase in 3-HP production compared with the parental strain. | |||
| 5′ UTR libraries of crtE, crtB, and crtI were generated using UTR Library Designer. For each gene, 16 different 5′ UTR sequences were designed. The highest lycopene titer was achieved by expressing crtE, crtB, and crtI, each using its optimally designed UTR sequence. | |||
| To fine-tune pckA expression, four 5′ UTR variants were generated using UTR Library Designer. The best UTR sequence, leading to fine-tuned pckA expression, resulted in a strain that exhibited a 49.8-fold increase in naringenin production compared with the non-optimized strain. | |||
| Dynamic expression regulation | Riboswitch | Riboswitch is a regulatory element in 5’ UTR, controlling the expression of downstream coding sequence due to its structural change upon interaction with its ligand. | |
| Two lysine riboswitches were utilized to activate and repress the expression of aspartate kinase III and homoserine dehydrogenase, respectively, in the lysine-producing strain C. glutamicum OW45. As a result, lysine production increased by 35% in strain QW48 (A263-lysC) and 43% in strain QW54 (R357-hom), compared with the parental strain QW45. | |||
| Toehold switches | Toehold switches are designed to detect specific RNA sequences, known as "triggers," with high sensitivity and specificity. Toehold switch was designed to detect coronavirus RNA, which acts like a trigger. The detection system implemented on a paper-cell free expression system could detect coronavirus. |
| Expression | UTR elements | Advantages | Limitations | Ref |
|---|---|---|---|---|
| Protein Over-expression | AU-rich | Incorporating AU-rich elements into the 5′ UTR region can stabilize mRNA through S1 and Hfq proteins, thereby enhancing protein production. | The relatively long AU-rich element derived from the sodB gene, for example, may enhance RNase E accessibility, potentially reducing mRNA stability. | |
| RG4 | RG4 structures act as internal ribosome entry sites, allowing translation initiation independent of a TSS. Furthermore, their incorporation into the 3′ UTR enhances mRNA stability and protects against RNase-mediated degradation. | The strong helical structure of RG4 complicates the incorporation of other elements like riboswitches into the 5' UTR, because ribonucleotides forming strong intramolecular interactions within the RG4 structure can also interact with other elements, disrupting their structures. | ||
| Synthetic dual UTRs | Dual UTRs elements enhance both transcription and translation with an optimal distance between two UTRs within the 5′ UTR region. | The concatenated UTRs may contain regulatory elements and thus the performance of optimized UTRs may vary under different environmental conditions or growth phases. | ||
| ProQC | The ProQC system ensures translation occurs only in the presence of full-length mRNAs by facilitating circularization through a toehold switch in the 5′ UTR and a cis-trigger sequence in the 3′ UTR. This mechanism enhances full-length translation, thereby improving protein quality in bacteria. | The incorporation of a toehold switch, forming a strong secondary structure, complicates its integration with other 5′ UTR elements. In addition, to further enhance full-length protein production, improving the circularization efficiency and stability of circular mRNA requires further investigation. | ||
| Expression fine-optimization | RBS | A UTR library offers a diverse set of UTR sequences, including RBSs, that can be incorporated into the 5′ UTR region to fine-tune target gene expression by varying ribosome binding affinity. | UTR libraries are designed for static fine-tuning rather than dynamic regulation. | |
| Dynamic expression regulation | Riboswitch | Riboswitches regulate gene expression by binding specific metabolites, switching expression "on" or "off," and controlling downstream gene activity. | Riboswitches may exhibit leaky expression in the OFF state or fail to achieve full activation in the ON state. | |
| Toehold switches | Toehold switches regulate gene expression by binding trigger RNAs, toggling expression "on" or "off," and controlling downstream gene activity. | The strong secondary structure of toehold switch complicates its integration with other 5′ UTR elements, and like riboswitches toehold switches often exhibit leaky expression in the OFF state or fail to achieve full activation in the ON state. Unintended interactions with endogenous RNAs can also lead to undesirable regulatory effects. |
| Software | Description | Ref | Exemplar applications |
|---|---|---|---|
| RBSDesinger | RBSDesigner is a mathematical model accounting for the thermodynamic RBS folding (30 nt from SD) and the interaction of RBS with ribosomes. The thermodynamic parameters in the model were calculated using UNAFold software package. The model’s prediction performance was R2=0.77 – 0.87. | The RBS of the ppc gene, optimized by RBSDesigner, enhanced both rapid cell growth and high 3-AP production in E. coli by facilitating the conversion of phosphoenolpyruvate to oxaloacetate, which is then converted to aspartate and ultimately to 3-AP ( |
|
| URL: |
|||
| RBSCalculator / Operon Calculator | RBSCalculator is a thermodynamic model composed of five energy parameters calculated from -35 to +35 nt of TIR by using NuPACK suite. Its prediction performance was R2=0.51 – 0.95 depending on the origin of UTR. | Optimized expression of fabH and fabZ using RBS sequences designed by RBSCalculator enhanced the synthesis of fatty acyl-ACP from acetyl-CoA. Fatty acyl-ACP was then converted to fatty alcohol by the far gene, resulting in a high production titer of fatty alcohol ( |
|
| Operon Calculator is a biophysical model based on the model of RBSCalculator to predict the translation of bi- and tri-cistronic genes. Its performance is Pearson R2=0.57 – 0.91. | |||
| URL: |
|||
| UTRDesigner / UTR Library Designer | UTRDesigner is a thermodynamic model that accounts for several parameters, calculated from -10 to +35 nt of the TIR using NuPACK. Its performance is R² = 0.81. UTR Library Designer is a tool used to generate UTR sequences with diverse expression levels, utilizing UTRDesigner. | Seo et al. ( |
For enhanced itaconic acid production, multiple genes (acs, gltA, and aceA) involved in acetate assimilation and the glyoxylate shunt pathway were overexpressed using optimized RBS sequences designed by UTRDesigner ( |
| URL: |
Table 1.
Table 2.
Table 3.
TOP
 MSK
MSK
 ePub Link
ePub Link Cite this Article
Cite this Article


