ABSTRACT
- Two novel bacterial species, designated as CJ85T and CJ88T, were isolated from the agricultural soil and the Han River, South Korea, respectively. Cells of both strains were Gram-staining-positive, short rod-shaped, non-motile, and yellow-pigmented. Strain CJ85T exhibited optimal growth in tryptic soy broth at 37°C and pH 7.0 in the absence of NaCl. Strain CJ88T showed optimal growth in lysogeny broth at 30°C and pH 7.0 in the absence of NaCl. Phylogenetic analysis based on 16S rRNA gene sequences revealed that strain CJ85T belonged to the genus Paramicrobacterium, showing the highest sequence similarity to Paramicrobacterium fandaimingii HY82T (97.6%). Strain CJ88T was assigned to the genus Microbacterium, with the highest sequence similarity to Microbacterium azadirachtae DSM 23848T (98.5%). The DNA G + C content was 64.8% for strain CJ85T and 70.5% for strain CJ88T. The genome-based analyses, including phylogenomic tree, digital DNA-DNA hybridization, and average nucleotide identity, clearly indicated that these strains represent novel species within their respective genera. The major fatty acids of both strains were anteiso-C15:0, anteiso-C17:0, and iso-C16:0. Based on the polyphasic taxonomy study, strains CJ85T and CJ88T represent novel species of the genera Paramicrobacterium and Microbacterium, respectively, for which names Paramicrobacterium salitolerans sp. nov. and Microbacterium fluminis sp. nov. are proposed. The type strains CJ85T (= KACC 23064T = JCM 36217T) and CJ88T (= KACC 24080T = JCM 38050T).
-
Keywords: Paramicrobacterium, Microbacterium, Paramicrobacterium salitolerans, Microbacterium fluminis, genome
Introduction
Plant-pathogenic bacteria belonging to the family Microbacteriaceae within the phylum Actinomycetota were first reported about a century ago (Smith, 1910, 1913) and the type species of the type genus of the family, Microbacterium lacticum isolated from dairy products was described afterwards (Orla-Jensen, 1919). Members of this family are Gram-positive, predominantly slender irregular rods, colored in colonies due to carotenoid pigments, chemo-organotrophic, aerobic, catalase-positive, mesophiles, and neutrophiles (Evtushenko, 2015). According to the List of Prokaryotic names with Standing in Nomenclature (LPSN), the family Microbacteriaceae currently encompasses 69 genera with validly published names (Parte et al., 2020).
The genus Microbacterium was proposed by Orla-Jensen (1919), of which the description was emended by Collins et al. (1983) and emended again by Takeuchi and Hatano (1998) to unite Microbacterium and Aureobacterium. The members of the genus Microbacterium have been isolated from diverse habitats such as seawater (Kim et al., 2008), soil (Lee et al., 2014), air (Zlamala et al., 2002), plants (Behrendt et al., 2001), and human blood (Clermont et al., 2009). They are characterized by the presence of N-glycolyl residues in the cell wall, menaquinone-11 (MK-11), MK-12, and MK-13 as primary respiratory quinones. The polar lipids consist of diphosphatidylglycerol, phosphatidylglycerol and one or more glycolipids; cellular fatty acids are predominantly anteiso-C15:0, anteiso-C17:0, and iso-C16:0 (Suzuki and Hamada, 2015). The genomic DNA G + C content ranged 61.6–71.8% (Lee et al., 2023). Currently, a total of 160 species with validly published names are reported under the genus Microbacterium, according to the LPSN (Parte et al., 2020).
The genus Paramicrobacterium was first proposed by Lee et al. (2023). The four members of genus Microbacterium–M. agarici, M. chengjingii, M. fandaimingii, and M. humi–were transferred to the genus Paramicrobacterium based on the phylogenomic analysis, including average nucleotide identity (ANI), average amino acid sequence identity (AAI), and digital DNA-DNA hybridization (dDDH) values (Lee et al., 2023). The members of the genus Paramicrobacterium are isolated from animals (Young et al., 2010) and plants (Zho et al., 2021). Chemotaxonomic characteristics of Paramicrobacterium were almost similar to those of Microbacterium (Lee et al., 2023). The genomic DNA G + C content values of Paramicrobacterium were 62.0–64.6% (Lee et al., 2023). During this study, we initially isolated two strains belonging to the genus Microbacterium but, strain CJ85T was transferred to the newly proposed genus Paramicrobacterium. Based on the polyphasic taxonomy and genomic approaches, strains CJ85T and CJ88T are proposed as novel species of the genus Paramicrobacterium and Microbacterium which isolated from the lettuce cultivation soil in Yongin-si and the Han River, South Korea.
Materials and Methods
Isolation of bacterial strains and culture conditions
Strains CJ85T and CJ88T were isolated from a lettuce cultivation soil in Yongin (37°9'30.67"N, 127°17'20.29"E) and from a freshwater sample of the Han River (37°24'5.45"N, 127°32'21.77"E), South Korea, respectively. Surface river water (0–1 m depth) and lettuce cultivation soil (5–10 cm depth) were collected. Each sample was inoculated into Reasoner’s 2A (R2A) broth (BD) at 30°C with shaking at 200 rpm. The resulting cultures were serially diluted and spread onto R2A agar. Single colonies were subsequently picked and purified through successive streaking on R2A agar at 30°C. For long-term storage, strains CJ85T and CJ88T were grown in R2A broth at 30°C for two days and subsequently maintained at –80°C in 30% (w/v) glycerol suspension.
Phylogeny analysis based on 16S rRNA genes
A single colony of each strain served as the template for PCR amplification of the 16S rRNA gene. The amplicons were sequenced at Biofact (Daejeon, Korea) using an automated DNA analyzer and sequencing primers 27F, 1492R, 785F, and 805R (Baker et al., 2003). The sequences were assembled to generate a consensus sequence for each strain. Alignment between the consensus sequence of each strain and those of related type strains selected from Ezbiocloud database (https://www.ezbiocloud.net/) (Chalita et al., 2024) was executed through the MUSCLE program (Edgar, 2004). Phylogenetic trees were constructed using the neighbor-joining (NJ), maximum-parsimony (MP), and maximum-likelihood (ML) methods. The NJ and MP trees were constructed using MEGA 12 software (Kumar et al., 2024). For NJ trees, evolutionary distances were calculated using the Jukes-Cantor model (Jukes and Cantor, 1969). The MP trees were inferred using the pairwise deletion option for gaps. The ML trees were reconstructed using IQ-TREE, and the best-fit substitution model was determined by ModelFinder Plus under the Bayesian Information Criterion (Minh et al., 2020). The selected model for strain CJ85T was the Transitional Model 3 with empirical base frequencies (+F) and a discrete Gamma distribution with three categories (+R3). For strain CJ88T, the model was the Transitional Model with empirical base frequencies (+F), a proportion of invariable sites (+I), and a discrete Gamma distribution with four rate categories (+G4). The topologies of all trees were evaluated based on bootstrap analysis of 1,000 datasets (Felsenstein, 1985).
Whole genome sequencing and de novo assembly
The extraction of genomic DNA from each strain was carried out using DNeasy Blood & Tissue Kits (QIAGEN, Germany) in accordance with the manufacturer’s protocols. Whole genome sequencing was performed at JS LINK (Korea) using a hybrid approach combining Oxford Nanopore and Illumina platforms. For long-read sequencing, a genomic DNA library was prepared using the Ligation Sequencing Kit (SQK-LSK109; Oxford Nanopore, UK) and sequenced on an Oxford Nanopore MinION instrument with R9.4.1 flow cells (FLO-MIN106; Oxford Nanopore). For short-read sequencing, a separate library preparation was performed using the TruSeq DNA PCR-Free Kit and sequenced on an Illumina NovaSeq 6000 platform. The resulting Nanopore and Illumina reads were subject to de novo hybrid genome assembly with Unicycler v0.5.0 (Wick et al., 2017). The completeness of the final genome assemblies was assessed using BUSCO v5.3.2 against the Microbacteriaceae single-copy ortholog dataset (Manni et al., 2021).
Phylogenomic analysis and genome-based characterization
To comply with the minimal standards for prokaryotic genomic taxonomy recommended by Chun et al. (2018), we estimated dDDH values through the Genome-to-Genome Distance Calculator (GGDC 3.0) (Meier-Kolthoff et al., 2022). ANI values were calculated using the OrthoANI pipeline (Lee et al., 2016). For genus-level classification, AAI values were calculated with the EzAAI software (Kim et al., 2021b), and the percentage of conserved proteins (POCP) values were determined as described by Qin et al. (2014). Reference genome sequences of closely related type strains were obtained from the national center for biotechnology information (NCBI) type material database (O'Leary et al., 2024). The up-to-date bacterial core genes 2 (UBCG2) pipeline was used to generate the concatenated alignment of 81 UBCG sequences (Kim et al., 2021a). Phylogenomic trees were constructed from the concatenated sequences using IQ-TREE with the general time reversible substitution model, empirical base frequency (+F), and the FreeRate model with ten categories (+R10) (Minh et al., 2020). Phylogenemic tree was visualized using iTOL v7 (Letunic and Bork, 2024). Protein-coding sequences (CDSs), rRNA genes, and tRNA genes were predicted and functionally annotated using Bakta (Schwengers et al., 2021). Functional metabolic pathways, carbohydrate-active enzymes, and proteolytic enzymes were predicted using the KEGG, CAZy, MEROPS databases, respectively, implemented in METABOLIC software (Zhou et al., 2022). Pan- and core-genome analyses were conducted using Panaroo pipeline (Tonkin-Hill et al., 2020). Antimicrobial resistance genes (ARGs) were identified based on the Comprehensive Antibiotic Resistance Database using the Resistance Gene Identifier with strict cutoff (Alcock et al., 2023).
Morphological and physiological characterization
Cell morphology was observed using transmission electron microscopy with cells grown on tryptic soy agar (TSA) at 30°C for 24 h. Growth was assessed in R2A broth, tryptic soy broth (TSB), Luria-Bertani (LB) broth, nutrient broth (NB), and marine broth (MB). The optimal growth temperature was determined on TSA (for strain CJ85T) and LB agar (for strain CJ88T) at temperatures ranging from 4 to 50°C (4, 10, 20, 25, 30, 37, 45, and 50°C). The pH range for growth was tested in TSB and LB broth media buffered to pH 4.0–5.0 (0.1 M citrate), pH 6.0–8.0 (0.1 M phosphate), and pH 9.0–10.0 (0.1 M bicarbonate-carbonate). NaCl tolerance was assessed in TSB and LB broth supplemented with 0.0–15% (w/v) NaCl in 1.0% increments. The pH and NaCl tolerance tests were carried out at 30°C for up to two days. Gram-staining was carried out using a commercial kit from Sigma-Aldrich, following the manufacturer’s protocols. Cell motility was examined by stab-inoculating a semi-solid medium (TSB or LB broth with 0.4% agar) and observing growth after seven days of incubation. Anaerobic growth was evaluated by incubating strains CJ85T and CJ88T on TSA and LB agar, respectively, at their optimal temperature for two weeks using the GasPak Anaerobe Pouch System (BD). Oxidase and catalase activities were determined using oxidase reagent (bioMérieux) and 3.0% (v/v) H2O2 solution, respectively. Hydrolysis of starch (1.0% w/v), cellulose (0.2% w/v, carboxymethyl cellulose), casein (3.0% w/v, skim milk), and DNA (DNase test agar, BD) was assessed after seven days of incubation. Carbon source utilization and additional enzymatic activities were determined using API 20NE kit according to the manufacturer’s instructions.
Chemotaxonomic analyses
For chemotaxonomic analyses, cells of strain CJ85T and CJ88T were cultured on TSA and LB agar, respectively, for 24 h at their optimal temperatures. The polar lipids were extracted from 100 mg of freeze-dried cells and analyzed by two-dimensional silica gel thin-layer chromatography (TLC) as described by Minnikin et al. (1984). For the first-dimension separation, a solvent mixture of chloroform:methanol:water (32.5:12.5:2, v/v/v) was employed, followed by a second dimension developed with chloroform:methanol:acetic acid:water (40:6:7.5:2, v/v/v/v). Separated polar lipids were visualized by spraying with specific reagents: 5% (w/v) phosphomolybdic acid for total lipids, molybdenum blue spray reagent (Sigma-Aldrich) for phospholipids, 0.25% (w/v) ninhydrin for aminolipids, and 15% (w/v) α-naphthol for glycolipids. For peptidoglycan analysis, whole cells were hydrolyzed in 6 N HCl at 121°C for 15 min, and the hydrolysates were analyzed by TLC on silica plates using the solvent system of Hasegawa et al. (1983). Respiratory quinones were extracted as described by Minnikin et al. (1984) and analyzed by high-performance liquid chromatography on a reverse phase column (Apollo C18, 250 × 4.6 mm, 5 μm) (Collins, 1985). Quinones were eluted with a methanol:iso-propanol mixture (4:1, v/v) at a flow rate of 1 ml/min and detected by UV absorbance at 270 nm. Cellular fatty acids were saponified, methylated, extracted, and analyzed by gas chromatography (Hewlett Packard 6890) using the standard protocol of the Microbial Identification System (Sherlock ver. 6.0B; RTSBA6 database) (Sasser, 1990).
Antimicrobial susceptibility test
The antimicrobial susceptibility was evaluated by the disk diffusion assay using antibiotic impregnated disks (Lioflchem, Italy). The following antibiotics were tested (disk content in μg): chloramphenicol (30), ciprofloxacin (5), clindamycin (2), erythromycin (15), fosfomycin (200), gentamicin (10), linezolid (10), rifampicin (5), streptomycin (10), tetracycline (30), trimethoprim (5), and vancomycin (30). Results were interpreted as susceptible (S), intermediate (I), or resistant (R) according to the clinical breakpoints for Staphylococcus spp. and Enterococcus spp. provided in the CLSI breakpoint table (Lewis II et al., 2025).
GenBank accession numbers
The GenBank accession numbers for the 16S rRNA gene sequences of strains CJ85T and CJ88T are PV837006 and PV837007, respectively, and for the genome sequences are GCA_051513965.1 and GCA_051549805.1, respectively.
Results and Discussion
16S rRNA phylogeny
Phylogenetic analysis based on the 16S rRNA gene sequences revealed that strain CJ85T formed a robust clade with members of the genus Paramicrobacterium, displaying the highest sequence similarity with P. fandaimingii HY82T (97.7%), followed by P. agarici DSM 21798T (97.5%) and P. chengjingii HY60T (97.4%; Fig. 1A); the topologies of the NJ, ML, and MP trees were congruent (Figs. 1A and S1). Strain CJ88T clustered with M. lemovicicum DSM 25044T with the highest 16S rRNA gene sequence similarity of 98.4% (Fig. 1B); however, the phylogenetic position of strain CJ88T was not consistent across the different analytical methods, showing a different topology in the MP tree compared to the NJ and ML trees (Figs. 1B and S2). Based on pairwise 16S rRNA gene sequence identities below the established threshold for species delineation (Kim et al., 2014; Rosselló-Móra and Amann, 2015), strains CJ85T and CJ88T represent novel species within the genus Paramicrobacterium and Microbacterium, respectively.
Genomic features
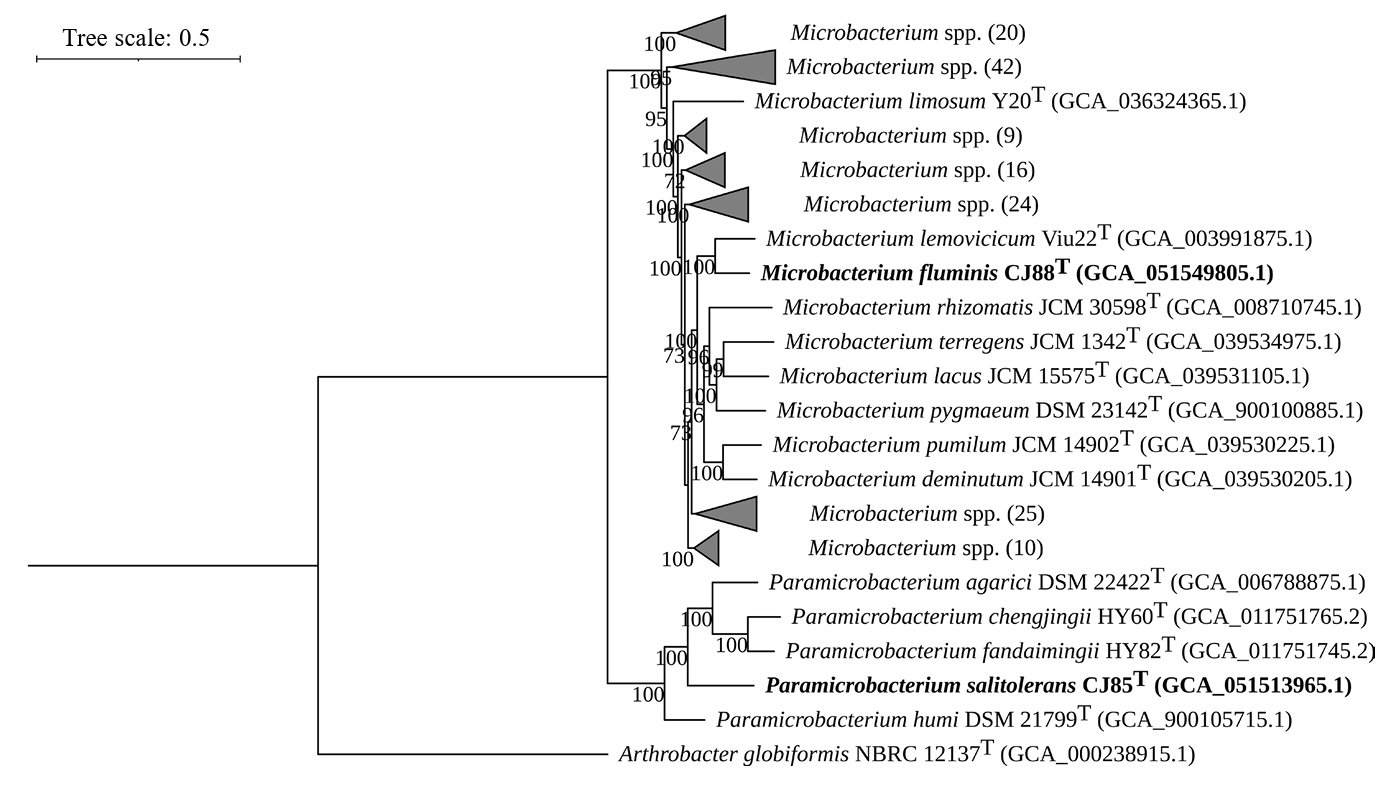
The genome assemblies of strains CJ85ᵀ and CJ88ᵀ exhibited 98.7% and 99.5% completeness (Fig. S3), with genome sizes of 3.17 Mbp and 3.24 Mbp, respectively. The genomic DNA G + C contents were 64.8% for strain CJ85T and 70.5% for strain CJ88T. The genome of strain CJ85T was predicted to contain 3,012 CDSs, 47 tRNA genes, and 6 rRNA genes, while that of strain CJ88T contained 3,065 CDSs, 48 tRNA genes, and 6 rRNA genes. Both strains harbored two identical copies of the 16S rRNA genes. The genomic characteristics of both strains and their related type strains are summarized in Table 1. A phylogenomic tree constructed using the concatenated sequences of 81 UBCGs from the two novel strains and 158 related type strains (Microbacterium and Paramicrobacterium) showed that strain CJ85T clustered within the genus Paramicrobacterium, forming a robust monophyletic clade with P. fandaimingii HY82T, P. agarici DSM 21798T, and P. chengjingii HY60T. Strain CJ88T was positioned within the genus Microbacterium, and indicated the closest phylogenetic relationship to M. lemovicicum DSM 25044T, complementing the incongruence of 16S rRNA gene-based phylogeny (Figs. 2 and S4).
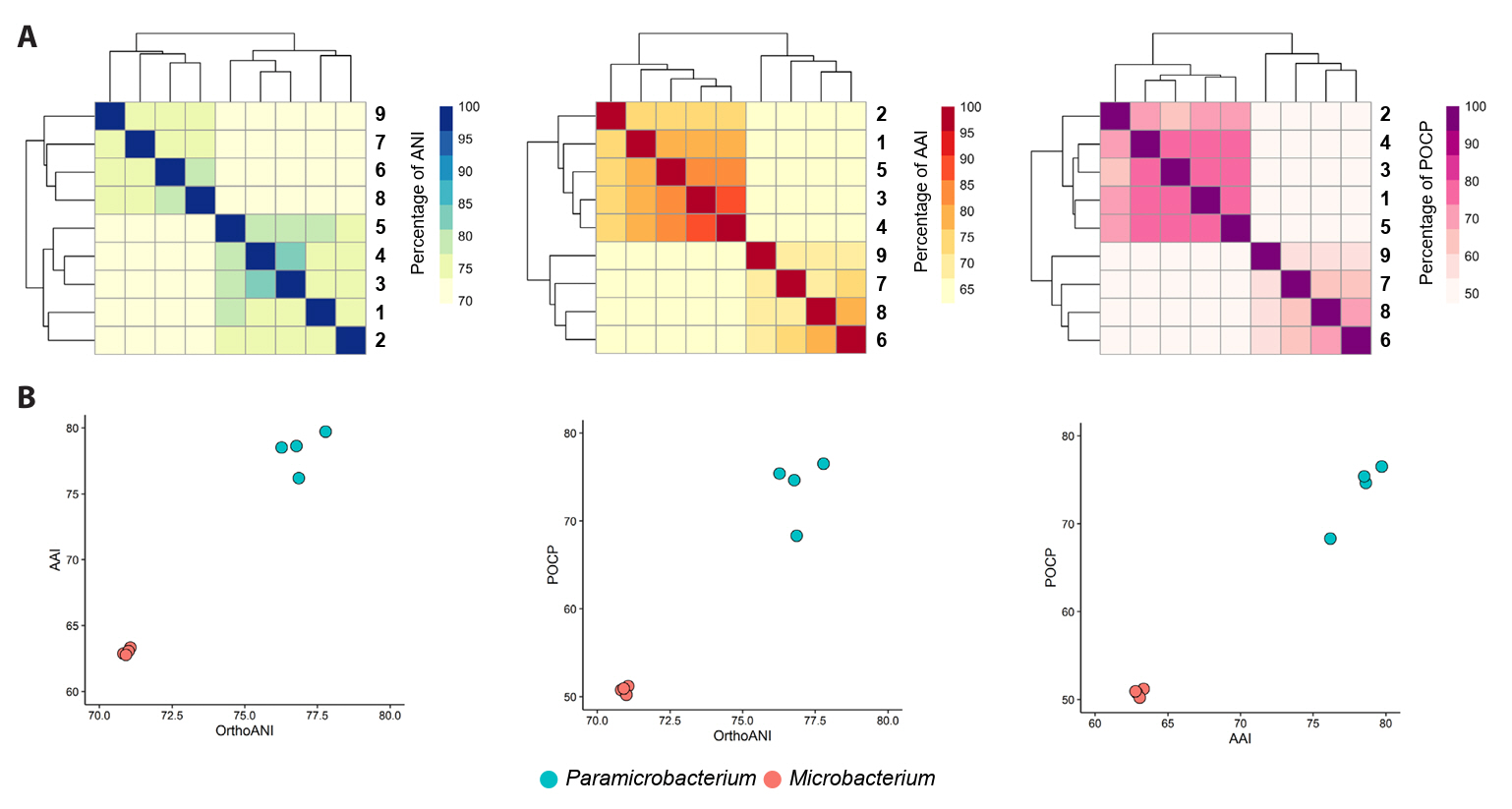
For taxonomic classification, a suite of genomic indices was calculated. The ANI values between strain CJ85T and its closest related type strains, P. fandaimingii HY82T, P. agarici DSM 21798T, P. chengjingii HY60T, and P. humi DSM 21799T, were 76.8%, 77.8%, 76.3%, and 76.9%, respectively (Table S1). The ANI values between strain CJ88T and M. azadirachtae DSM 23848T, M. lemovicicum ViU22T, and M. binotii JCM 16365T were 75.9%, 80.4%, and 76.9%, respectively (Table S1). All dDDH values between the novel strains and their related type strains were below 30% (Table S1). These results indicate that both strains represent a novel species within the genera Paramicrobacterium and Microbacterium, respectively, based on genomic ANI (< 95–96%) and dDDH values (< 70%) (Chun et al., 2018; Meier-Kolthoff et al., 2013; Richter and Rosselló-Móra, 2009). Furthermore, AAI and POCP values revealed a clear distinction between strains CJ85T and CJ88T and their related type strains at the genus level (Fig. 3, Table S2), supporting the separation of Paramicrobacterium from Microbacterium. Pangenome analysis of species of Paramicrobacterium and Microbacterium identified 1,711 and 963 core orthologous genes, respectively, whereas a combined analysis of both genera identified a shared core genome of 540 orthologous genes (Fig. S5).
Two glycopeptide ARGs (vanYB and vanYA) were predicted in the genome of strain CJ88T. However, the predicted resistance genotype was not consistent with the observed phenotype (Table S3). KEGG pathway analysis revealed that the genome of strains CJ85T and CJ88T encode 65 and 58 complete metabolic pathway modules, respectively (Fig. S6, Table S4). Both strains possess well-conserved central carbohydrate metabolism pathways, including glycolysis, glucogenesis, pentose phosphate pathway, phosphoribosyl diphosphate biosynthesis, and citrate cycle (Fig. S6). Notably, pathways for ectoine biosynthesis and β-lactam resistance (bla system) were found exclusively in genus Paramicrobacterium. Analysis of carbohydrate-active enzymes showed that the genome of strain CJ85T encoded two polysaccharide lyases (PLs) and 45 glycoside hydrolases (GHs), whereas the strain CJ88T genome encoded 22 GHs with no PL (Fig. S7A). Specifically, the genome of strain CJ85T harbored unique GH18 and GH19 families (Fig. S7B), which were absent from other related type strains. Strain CJ88T exhibited no distinct GH family profile compared to other related type strains (Fig. S7B). Both strains and the other related type strains all possessed GH enzymes required for cellulose degradation, suggesting that GH family profile is not consistent with the phenotypic characteristic of cellulose degradation (Fig. S7, Table 2); functional cellulolytic activity likely results from the synergistic action of multiple GH enzymes (Contato et al., 2025). The higher number of GH genes present in the CJ85T genome suggests that strain CJ85T possesses genomic content more optimized for growth in soil habitats compared to the freshwater-derived strain CJ88T. Analysis of proteolytic enzymes revealed that metallo, serine, and cysteine proteases were the predominant proteolytic enzymes in both strains (Fig. S8). Regarding flagellar biosynthesis genes, flhA gene was identified in the genome of strain CJ85T, whereas no flagellar-related gene was found in strain CJ88T. However, since the assembly of a functional flagellum requires a complex apparatus consisting of numerous proteins (Worrall et al., 2023), the presence of flhA gene alone is insufficient for flagellar formation. These genomic findings are consistent with the non-flagellated morphology of both strains, as observed in transmission electron microscopy images (Fig. S9).
Characterization of morphological, phenotypic, and biochemical features
Strains CJ85T and CJ88T were Gram-stain-positive, strictly aerobic, non-motile, and non-spore-forming rods. After 24 h of cultivation, colonies of strain CJ85T on TSA at 37°C were smooth, light-yellow, and circular, while colonies of CJ88T on LB agar at 30°C were smooth, yellow-pigmented, and circular. Transmission electron micrographs showed that both cells were rod-shaped (Fig. S9). Optimal growth for CJ85T was observed on TSA and for strain CJ88T on LB agar; no growth was observed for either strain on marine broth. Strain CJ85T grew at 10–45°C (optimum 37°C) and at pH 5.0–9.0 in the presence of 0.0–14.0% (w/v) NaCl. Strain CJ88T grew at 10–45°C (optimum 30°C) and at pH 6.0–9.0 in the presence of 0.0–3.0% (w/v) NaCl. For both strains, optimal growth was perceived at pH 7.0 in the absence of NaCl. Differential biochemical and physiological characteristics between the novel strains and their closely related type strains are provided in Table 2.
Chemotaxonomic characteristics
The predominant respiratory quinone was identified as MK-11 for strain CJ85T and MK-12 for strain CJ88T. The cell-wall peptidoglycan of both strains contained ornithine and meso-diaminopimelic acid. As shown in Table 3, the major cellular fatty acids of strain CJ85T were anteiso-C15:0 (42.3%), iso-C16:0 (22.7%), and anteiso-C17:0 (22.1%; Table 3). The polar lipids of strain CJ85T included diphospatidylglycerol, phosphatidylglycerol, two glycolipids, and two unidentified polar lipids (Fig. S10A). For strain CJ88T, the major cellular fatty acids were anteiso-C15:0 (40.2%), iso-C16:0 (21.5%), and anteiso-C17:0 (19.2%; Table 3). The polar lipids of strain CJ88T included diphospatidylglycerol, phosphatidylglycerol, glycolipid, and one unidentified polar lipid. Phosphatidylethanolamine was not detected in either strain (Fig. S10B). These chemotaxonomic features are consistent with those of their related genera within the family Microbacteriaceae.
Taxonomic conclusion
Genomic, phylogenetic, chemotaxonomic, and phenotypic analyses demonstrated that strain CJ85T belongs to the genus Paramicrobacterium and strain CJ88T belongs to the genus Microbacterium. The observed characteristics of both strains, including respiratory quinones, fatty acids, polar lipids, and carbon source utilization patterns, were consistent with the descriptions of their related species of each genus. For strain CJ85T, several characteristics such as positive oxidase and urease activity, hydrolysis of cellulose, and the absence of the dTDP-L-rhamnose biosynthesis module, distinguished the novel strain from its related type strains. Strain CJ88T was differentiated by its negative β-galactosidase activity and hydrolysis of DNA, whereas its related type strains were positive. In contrast, strain CJ88T exhibited positive hydrolysis of cellulose and casein, whereas its related type strains were negative. Based on genomic analyses, the low dDDH (< 70%) and ANI (< 95–96%) values clearly differentiated strains CJ85T and CJ88T from all known species of each genus. Consequently, based on the polyphasic taxonomy analyses, strain CJ85T represents a novel species of the genus Paramicrobacterium, for which the name Paramicrobacterium salitolerans sp. nov. is proposed and strain CJ88T represents a novel species of the genus Microbacterium, for which the name Microbacterium fluminis is proposed.
Description of Paramicrobacterium salitolerans sp. nov.
Paramicrobacterium salitolerans (sa.li.to'le.rans. L. masc. n. sal, salt; L. pres. part. tolerans, tolerating; N.L. part. adj. salitolerans, salt-tolerating).
Cells are aerobic, Gram-stain-positive, non-motile and rod-shaped. Colonies grown for 24 h on TSA at 37°C are light-yellow, convex, and smooth. Growth is observed at 10–45°C (optimum 37°C), at pH 5.0–9.0 (optimum pH 7.0) and in the presence of 0.0–14.0% NaCl (optimum 0.0%). Catalase- and oxidase-positive. Starch, carboxymethyl cellulose, DNA, and aesculin ferric citrate are hydrolyzed but casein and gelatin are not. Indole is not produced. Enzyme activities for urease and β-galactosidase are positive but negative for arginine dihydrolase. According to the API 20NE test, D-glucose, L-arabinose, D-mannose, D-mannitol, N-acetyl-glucosamine, D-maltose, and potassium gluconate are assimilated, but capric acid, adipic acid, malic acid, trisodium citrate, and phenylacetic acid are not. The predominant polar lipids are diphosphatidylglycerol, phosphatidylglycerol, two glycolipids, and two unidentified polar lipids. The major fatty acids are anteiso-C15:0, iso-C16:0, and anteiso-C17:0. The predominant respiratory quinone is MK-11. The type strain is CJ85T (= KACC 23064T = JCM 36217T), isolated from a lettuce cultivation soil in Yongin, Korea. The DNA G + C content of the type strain is 64.8%. The GenBank accession numbers for the 16S rRNA gene sequence and the whole genome sequence of strain CJ85T are PV837006 and GCA_051513965.1, respectively.
Description of Microbacterium fluminis sp. nov.
Microbacterium fluminis (flu'mi.nis. L. neut. n. flumen, a river; N.L. gen. n. fluminis, of a river).
Cells are aerobic, Gram-stain-positive, non-spore-forming, non-motile, and rod-shaped. Colonies grown for 24 h on LB agar at 30°C are yellow, convex, and smooth. Growth is observed at 10–45°C (optimum 30°C), at pH 6.0–9.0 (optimum pH 7.0) and in the presence of 0.0–3.0% NaCl (optimum 0.0%). Catalase- and oxidase-positive. Starch, carboxymethyl cellulose, casein, and aesculin ferric citrate are hydrolyzed but DNA and gelatin are not. Indole is not produced. Enzyme activity is negative for urease, β-galactosidase and arginine dihydrolase. According to the API 20NE test, D-glucose, D-mannose, D-mannitol, D-maltose, potassium gluconate, and malic acid are assimilated, but L-arabinose, N-acetyl-glucosamine, capric acid, adipic acid, trisodium citrate, and phenylacetic acid are not. The predominant polar lipids are diphospatidylglycerol, phosphatidylglycerol, glycolipid, and one unidentified polar lipid. The major fatty acids are anteiso-C15:0, iso-C16:0, and anteiso-C17:0. The predominant respiratory quinone is MK-12. The type strain is CJ88T (= KACC 24080T = JCM 38050T), isolated from the Han River, Korea. The DNA G + C content of the type strain is 70.5%. The GenBank accession numbers for the 16S rRNA gene sequence and the whole genome sequence of strain CJ88T are PV837007 and GCA_051549805.1, respectively.
Acknowledgments
This work was supported by a grant from the National Institute of Biological Resources (NIBR), funded by the Ministry of Environment (MOE) of the Republic of Korea (NIBR202502203) and the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (NRF-2023R1A2C1003654). This research was also supported by the Chung-Ang University Young Scientists Scholarship in 2024. The authors thank Professor Aharon Oren for his assistance with nomenclature.
Conflict of Interest
The authors declare no conflicts of interest.
Ethical Statements
Not applicable.
Supplementary Information
The online version contains supplementary material available at https://doi.org/10.71150/jm.2512014.
Table S1.
ANI and dDDH values between two novel strains and their related type strains. ANI values were shaded in blue and dDDG values in green color.
Strains: 1, P. salitolerans CJ85T; 2, P. fandaimingii HY82T; 3, P. agarici DSM 21798T; 4, P. chengjingii HY60T; 5, P. humi DSM 21799T; 6, M. fluminis CJ88T; 7, M. azadirachtae DSM 23848T; 8, M. lemovicicum ViU22T; 9, M. binotii JCM 16365T.
jm-2512014-Supplementary-Table-S1.pdf
Table S2.
AAI and POCP values between two novel strains and their related type strains. AAI values were shaded in red and POCP values in yellow color.
Strains: 1, P. salitolerans CJ85T; 2, P. fandaimingii HY82T; 3, P. agarici DSM 21798T; 4, P. chengjingii HY60T; 5, P. humi DSM 21799T; 6, M. fluminis CJ88T; 7, M. azadirachtae DSM 23848T; 8, M. lemovicicum ViU22T; 9, M. binotii JCM 16365T.
jm-2512014-Supplementary-Table-S2.pdf
Fig. S1.
Maximum-likelihood (A) and maximum-parsimony (B) phylogenetic trees of strain CJ85T and related type strains based on 16S rRNA gene sequences. Bootstrap values greater than 70% are shown at branch points based on 1,000 replicated datasets. Pseudoclavibacter soil DSM 23366T was used as an outgroup. Bar, 0.05 substitutions per nucleotide position for (A).
jm-2512014-Supplementary-Fig-S1.pdf
Fig. S2.
Maximum-likelihood (A) and maximum-parsimony (B) phylogenetic trees of strain CJ88T and related type strains based on 16S rRNA gene sequences. Bootstrap values greater than 70% are shown at branch points based on 1,000 replicated datasets. Agrococcus jenensis DSM 9580T was used as an outgroup. Bar, 0.05 substitutions per nucleotide position for (A).
jm-2512014-Supplementary-Fig-S2.pdf
Fig. S3.
Evaluation of the completeness of genome assembly results of strains CJ85T and CJ88T. The completeness of the assembled genomes was evaluated with the datasets of Microbacteriaceae lineage.
jm-2512014-Supplementary-Fig-S3.pdf
Fig. S4.
Phylogenomic tree of two novel strains and their related type strains based on 81 UBCGs. The phylogenomic tree was reconstructed using 158 type strains within the genera Paramicrobacterium and Microbacterium. Bootstrap value greater than 70% are shown at branch points based on maximum-likelihood analysis of 1,000 replicated datasets. The Arthrobacter globiformis NBRC 12137T was used as an outgroup. Bar, 0.5 substitutions per nucleotide position
jm-2512014-Supplementary-Fig-S4.pdf
Fig. S5.
Pangenome analysis of orthologous gene families within genera Paramicrobacterium and Microbacterium. The analysis depicts the distribution of orthologous gene families calculated separately for members of the genus Paramicrobacterium, members of the genus Microbacterium, and all strains from both genera combined. The numbers represent the counts of shared (core) or unique orthologous gene families among the strains.
Strains: 1, P. salitolerans CJ85T; 2, P. humi DSM 21799T; 3, P. chengjingii HY60T; 4, P. fandaimingii HY82T; 5, P. agarici DSM 21798T; 6, M. fluminis CJ88T; 7, M. binotii JCM 16365T; 8, M. lemovicicum ViU22T; 9, M. azadirachtae DSM 23848T.
jm-2512014-Supplementary-Fig-S5.pdf
Fig. S7.
Comparative analysis of carbohydrate-active enzymes among two novel strains and their related type strains. (A) The bar plot showed the number of carbohydrate-active enzymes classified by CAZy enzyme classes. (B) The heatmap indicated the number of carbohydrate-active enzymes according to CAZy enzyme families.
jm-2512014-Supplementary-Fig-S7.pdf
Fig. S9.
Transmission electron micrograph of strains CJ85T (A) and CJ88T (B). Cells were grown on TSA (A) and LB agar (B) for four days at 37°C. Bar, 1.0 μm
jm-2512014-Supplementary-Fig-S9.pdf
Fig. S10.
Two-dimensional TLC of the total polar lipids from strain CJ85T (A) and CJ88T (B). Abbreviations: PG, phosphatidylglycerol; DPG, diphosphatidylglycerol; GL, glycolipid; L, polar lipid.
jm-2512014-Supplementary-Fig-S10.pdf
Fig. 1.Neighbor-joining phylogenetic trees of strains CJ85T (A) and CJ88T (B) and their related type strains based on 16S rRNA gene sequences. Closed circles indicate nodes recovered in the three trees generated by neighbor-joining, maximum-likelihood, and maximum-parsimony methods. Bootstrap values greater than 70% are shown at branch points based on 1,000 replicated datasets. Pseudoclavibacter soil DSM 23366T (A) and Agrococcus jenensis DSM 9580T (B) were used as an outgroup, respectively. Bars, 0.01 substitutions per nucleotide position.
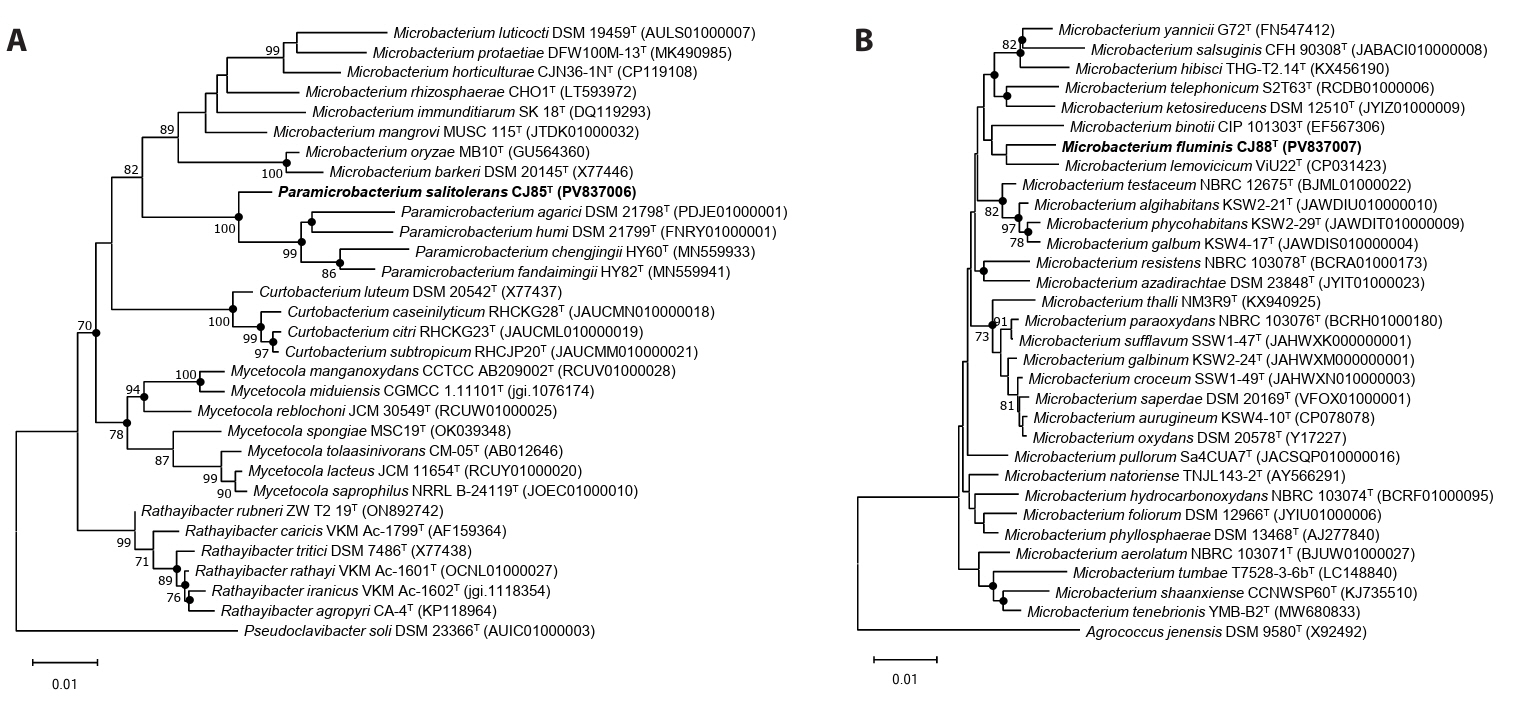
Fig. 2.Phylogenomic tree of strains CJ85T and CJ88T and their related type strains based on the concatenated sequences of 81 UBCGs. This is a summarized phylogenomic tree of the whole phylogenomic tree reconstructed using 158 type strains belonging to the genera Paramicrobacterium and Microbacterium. The whole phylogenomic tree is provided in Fig. S4. Numbers in parentheses indicate the number of species included in each collapsed clade. Bootstrap values greater than 70% are shown at branch points based on maximum-likelihood analysis of 1,000 replicated datasets. The Arthrobacter globiformis NBRC 12137T was used as an outgroup. Bar, 0.5 substitutions per nucleotide position.
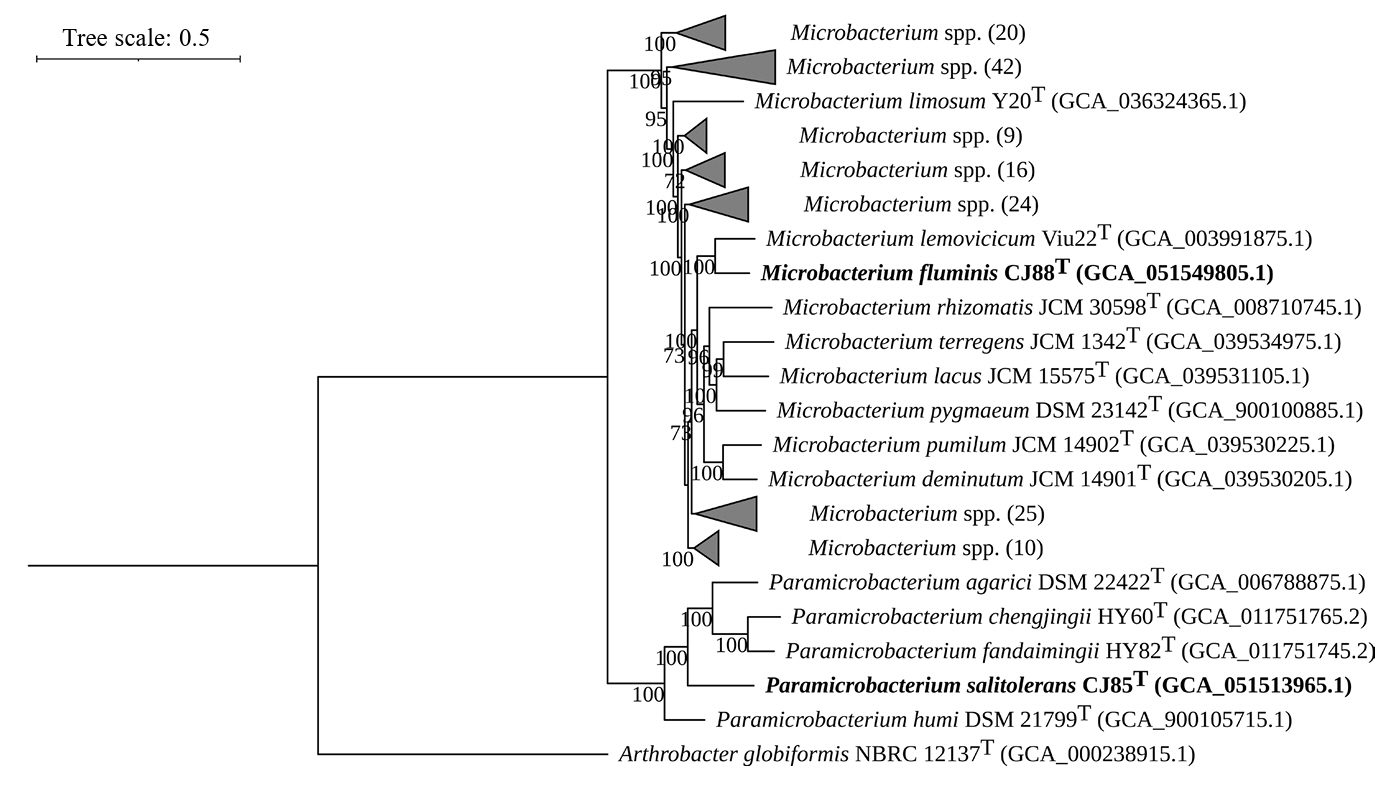
Fig. 3.
Overall genomic relatedness indices (OGRIs) of strains CJ85T and CJ88T and related type strains. (A) Heatmaps showing pairwise values for OrthoANI (blue), AAI (red), and POCP (purple) values among genomes of the genera Paramicrobacterium and Microbacterium. The dendrograms indicate hierarchical clustering of the genomes based on each OGRI. (B) Correlation plots showing the OGRI values calculated between Paramicrobacterium and Microbacterium species. Each value represents the set of values for a pairwise comparison against CJ85T, with the 100% self-comparison value excluded.
Strains: 1, P. salitolerans CJ85T; 2, P. humi DSM 21799T; 3, P. chengjingii HY60T; 4, P. fandaimingii HY82T; 5, P. agarici DSM 21798T; 6, M. fluminis CJ88T; 7, M. binotii JCM 16365T; 8, M. lemovicicum ViU22T; 9, M. azadirachtae DSM 23848T.
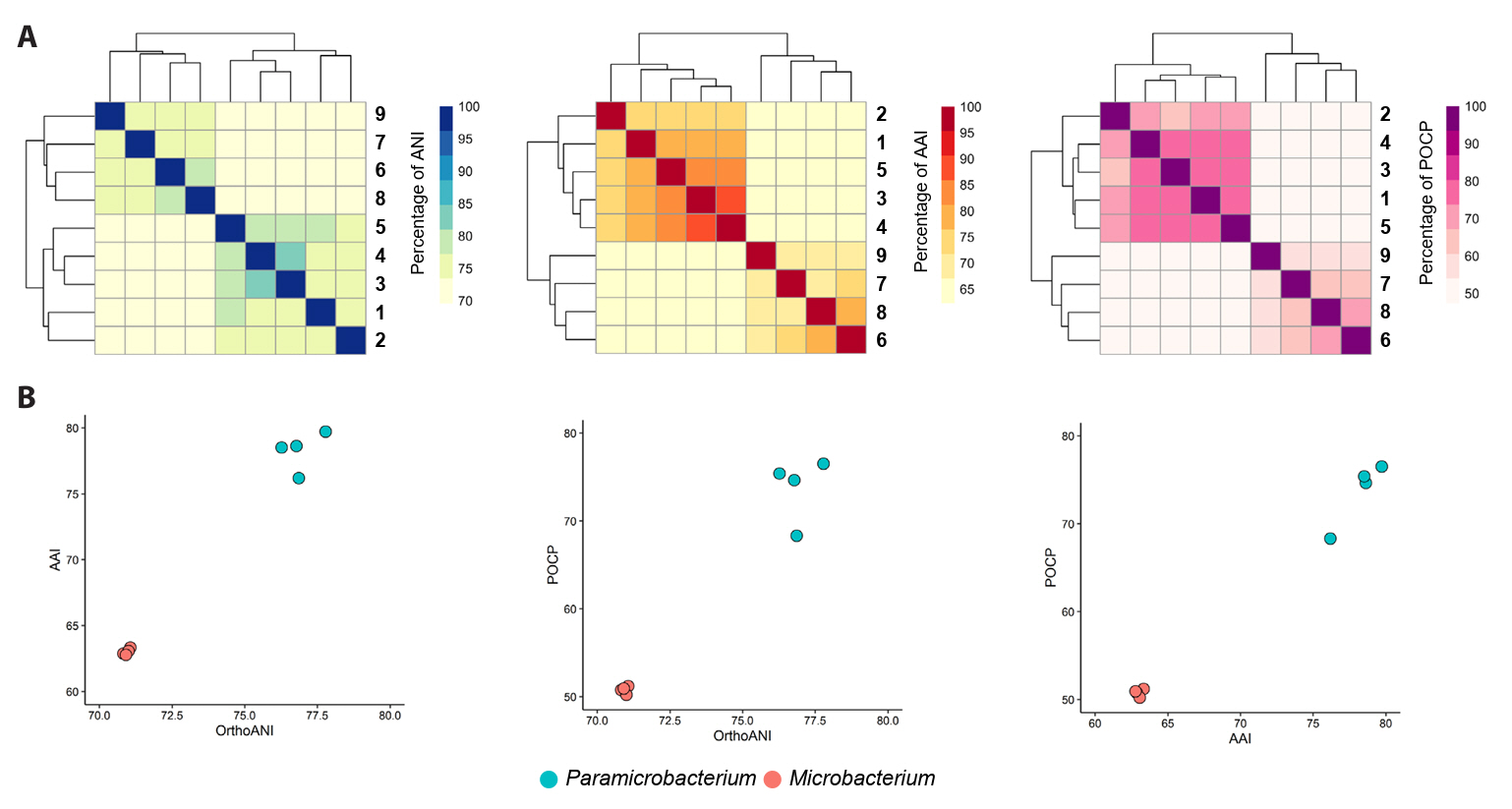
Table 1.Genomic features of strains CJ85T and CJ88T with their related type strains of the genera Paramicrobacterium and Microbacterium. All genomic features were predicted and annotated using Bakta.
|
Strain |
Accession number |
Size (Mbp) |
G + C contents (%) |
Number of |
|
contigs |
protein coding genes |
rRNAs |
tRNAs |
|
P. salitolerans CJ85T
|
GCA_051513965.1†
|
3.17 |
64.8 |
1 |
3,012 |
6 |
47 |
|
P. fandaimingii HY82T
|
GCA_011751745.2‡
|
3.62 |
63.3 |
1 |
3,464 |
6 |
47 |
|
P. agarici DSM 21798T
|
GCA_006788875.1‡
|
3.35 |
64.3 |
1 |
3,252 |
6 |
48 |
|
P. chengjingii HY60T
|
GCA_011751765.2‡
|
3.62 |
61.9 |
1 |
3,466 |
6 |
46 |
|
P. humi DSM 21799T
|
GCA_900105715.1‡
|
3.36 |
67.3 |
2 |
3,278 |
6 |
46 |
|
M. fluminis CJ88T
|
GCA_051549805.1†
|
3.24 |
70.5 |
1 |
3,065 |
6 |
48 |
|
M. azadirachtae DSM 23848T
|
GCA_000956545.1‡
|
4.04 |
70.5 |
86 |
3,755 |
5 |
45 |
|
M. lemovicicum ViU22T
|
GCA_003991875.1‡
|
3.56 |
70.9 |
1 |
3,297 |
6 |
48 |
|
M. binotii JCM 16365T
|
GCA_039532115.1‡
|
3.22 |
69.3 |
38 |
3,134 |
5 |
52 |
Table 2.Differential characteristics of Paramicrobacterium strain CJ85T and Microbacterium strain CJ88T and their related type strains
Strains: 1, P. salitolerans CJ85T; 2, P. fandaimingii HY82T; 3, P. agarici DSM 21798T; 4, P. chengjingii HY60T; 5, P. humi DSM 21799T; 6, M. fluminis CJ88T; 7, M. azadirachtae DSM 23848T; 8, M. lemovicicum ViU22T; 9, M. binotii JCM 16365T. All data are from this study unless indicated. All strains are positive for activity of catalase and aesculin hydrolysis. All strains are negative for indole production, activity of arginine dihydrolase, and assimilation of capric acid.
|
Characteristic |
1 |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
|
Isolation source
|
Freshwater |
Bat feces |
Mushroom |
Bat feces |
Mushroom |
Soil |
Rhizoplane |
Soil |
Human blood |
|
Colony color
|
Light-yellow |
White |
Yellow |
White |
Yellow |
Yellow |
Light-yellow |
Orange |
Yellow |
|
Growth*
|
|
|
|
|
|
|
|
|
|
|
Temperature (˚C) |
10–45 |
10–30 |
20–37 |
10–30 |
20–37 |
10–45 |
10–37 |
15–37 |
15–37 |
|
pH |
5–9 |
6–10 |
6–11 |
6–10 |
5–11 |
6–9 |
5–10 |
ND |
ND |
|
NaCl (%, w/v) |
0.0–14.0 |
0.5–10.5 |
0.5–10.5 |
0.5–10.5 |
0.5–10.5 |
0.0–3.0 |
0.0–5.5 |
0.0–4.5 |
0.0–6.0 |
|
Nitrate reduction
|
+ |
+ |
+ |
+ |
+ |
− |
+ |
− |
− |
|
Fermentation of
|
|
|
|
|
|
|
|
|
|
|
Glucose |
− |
− |
− |
− |
− |
− |
− |
− |
+ |
|
Enzyme activity
|
|
|
|
|
|
|
|
|
|
|
Oxidase |
+ |
− |
− |
− |
− |
+ |
− |
+ |
− |
|
Urease |
+ |
− |
− |
− |
− |
− |
− |
− |
− |
|
β-Galactosidase |
+ |
+ |
+ |
+ |
+ |
− |
+ |
+ |
+ |
|
Hydrolysis of
|
|
|
|
|
|
|
|
|
|
|
Starch |
+ |
+ |
+ |
− |
+ |
+ |
− |
+ |
− |
|
Cellulose |
+ |
− |
− |
− |
− |
+ |
− |
− |
− |
|
Casein |
− |
− |
− |
− |
− |
+ |
− |
− |
− |
|
DNA |
+ |
+ |
+ |
+ |
+ |
− |
+ |
+ |
+ |
|
Gelatin |
− |
− |
− |
− |
− |
− |
− |
+ |
− |
|
Assimilation of
|
|
|
|
|
|
|
|
|
|
|
D-Glucose |
+ |
+ |
+ |
− |
+ |
+ |
+ |
+ |
+ |
|
L-Arabinose |
+ |
− |
+ |
− |
+ |
− |
+ |
− |
+ |
|
D-Mannose |
+ |
− |
+ |
− |
+ |
+ |
+ |
− |
+ |
|
D-Mannitol |
+ |
+ |
+ |
− |
+ |
+ |
+ |
− |
+ |
|
N-Acetyl-glucosamine |
+ |
− |
+ |
− |
+ |
− |
+ |
− |
− |
|
D-Maltose |
+ |
− |
+ |
− |
+ |
+ |
+ |
+ |
+ |
|
Potassium gluconate |
+ |
− |
+ |
− |
+ |
+ |
+ |
− |
+ |
|
Adipic acid |
− |
− |
+ |
− |
+ |
− |
− |
− |
+ |
|
Malic acid |
− |
− |
+ |
− |
+ |
+ |
+ |
− |
+ |
|
Trisodium citrate |
− |
− |
+ |
− |
+ |
− |
+ |
− |
+ |
|
Phenylacetic acid |
− |
− |
− |
− |
− |
− |
− |
− |
+ |
Table 3.Fatty acid composition (%) of Paramicrobacterium strain CJ85T and Microbacterium strain CJ88T and the related type strains
Strains: 1, P. salitolerans CJ85T; 2, P. fandaimingii HY82T; 3, P. agarici DSM 21798T; 4, P. chengjingii HY60T; 5, P. humi DSM 21799T; 6, M. fluminis CJ88T; 7, M. azadirachtae DSM 23848T; 8, M. lemovicicum ViU22T; 9, M. binotii JCM 16365T. All data are obtained from this study.
|
Fatty acid |
1 |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
|
Straight
|
|
|
|
|
|
|
|
|
|
|
C14:0
|
TR |
- |
TR |
- |
TR |
- |
- |
- |
- |
|
C16:0
|
2.5 |
- |
3.9 |
- |
1.6 |
TR |
- |
- |
2.5 |
|
Unsaturated
|
|
|
|
|
|
|
|
|
|
|
C15:1 ω5c
|
3.4 |
- |
6.7 |
- |
TR |
- |
- |
- |
- |
|
Branched
|
|
|
|
|
|
|
|
|
|
|
iso-C10:0
|
- |
9.0 |
- |
10.0 |
- |
- |
7.5 |
10.9 |
- |
|
iso-C14:0
|
1.6 |
- |
1.8 |
- |
TR |
TR |
- |
- |
- |
|
iso-C15:0
|
4.4 |
10.8 |
4.7 |
9.5 |
7.1 |
11.2 |
10.1 |
- |
- |
|
iso-C16:0
|
22.7 |
15.0 |
18.3 |
14.1 |
19.8 |
21.5 |
16.2 |
17.5 |
19.5 |
|
iso-C17:0
|
TR |
- |
1.3 |
- |
2.6 |
5.0 |
3.6 |
- |
- |
|
anteiso-C15:0
|
42.3 |
44.3 |
43.5 |
39.7 |
41.0 |
40.2 |
34.9 |
50.5 |
39.7 |
|
anteiso-C17:0
|
22.1 |
17.7 |
17.7 |
17.5 |
23.4 |
19.2 |
23.8 |
13.8 |
35.8 |
|
anteiso-C15:1 A |
- |
- |
1.3 |
- |
- |
- |
- |
- |
- |
References
- Alcock BP, Huynh W, Chalil R, Smith KW, Raphenya AR, et al. 2023. CARD 2023: expanded curation, support for machine learning, and resistome prediction at the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res. 51: D690–D699. ArticlePubMedPMCPDF
- Baker GC, Smith JJ, Cowan DA. 2003. Review and re-analysis of domain-specific 16S primers. J Microbiol Methods. 55: 541–555. Article
- Behrendt U, Ulrich A, Schumann P. 2001. Description of Microbacterium foliorum sp. nov. and Microbacterium phyllosphaerae sp. nov., isolated from the phyllosphere of grasses and the surface litter after mulching the sward, and reclassification of Aureobacterium resistens (Funke et al. 1998) as Microbacterium resistens comb. nov. Int J Syst Evol Microbiol. 51: 1267–1276. ArticlePubMed
- Chalita M, Kim YO, Park S, Oh HS, Cho JH, et al. 2024. EzBioCloud: a genome-driven database and platform for microbiome identification and discovery. Int J Syst Evol Microbiol. 74: 006421.Article
- Chun J, Oren A, Ventosa A, Christensen H, Arahal DR, et al. 2018. Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol. 68: 461–466. Article
- Clermont D, Diard S, Bouchier C, Vivier C, Bimet F, et al. 2009. Microbacterium binotii sp. nov., isolated from human blood. Int J Syst Evol Microbiol. 59: 1016–1022. ArticlePubMed
- Collins MD. 1985. 11 Analysis of isoprenoid quinones. In Bergan T. (ed.), Methods in microbiology, vol. 18th, pp. 329–366. Academic Press. Link
- Collins MD, Jones D, Kroppenstedt RM. 1983. Reclassification of Brevibacterium imperiale (Steinhaus) and “Corynebacterium laevaniformans” (Dias and Bhat) in a redefined genus Microbacterium (Orla-Jensen), as Microbacterium imperiale comb. nov. and Microbacterium laevaniformans nom. rev., comb. nov. Syst Appl Microbiol. 4: 65–78. Article
- Contato AG, Pasin TM, Polizeli M. 2025. Microbial enzymes for biomass conversion. Annu Rev Microbiol. 79: 663–683. ArticlePubMed
- Edgar RC. 2004. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32: 1792–1797. Article
- Evtushenko LI. 2015. Microbacteriaceae. In Trujillo ME, Dedysh S, DeVos P, Hedlund B, Kämpfer P, et al. (eds.), Bergey’s manual of systematics of archaea and bacteria, 1st edn, pp. 1–14. John Wiley & Sons, Inc. Article
- Felsenstein J. 1985. Confidence limits on phylogenies: an approach using the bootstrap. Evolution. 39: 783–791. ArticlePubMedLink
- Hasegawa T, Takizawa M, Tanida S. 1983. A rapid analysis for chemical grouping of aerobic actinomycetes. J Gen Appl Microbiol. 29: 319–322. Article
- Jukes TH, Cantor CR. 1969. Mammalian protein metabolism. In Munro HN. (ed.), Evolution of protein molecules, pp. 21–132. Academic PressPDF
- Kim KK, Lee KC, Oh HM, Lee JS. 2008. Microbacterium aquimaris sp. nov., isolated from seawater. Int J Syst Evol Microbiol. 58: 1616–1620. ArticlePubMed
- Kim J, Na SI, Kim D, Chun J. 2021a. UBCG2: up-to-date bacterial core genes and pipeline for phylogenomic analysis. J Microbiol. 59: 609–615. ArticlePDF
- Kim M, Oh HS, Park SC, Chun J. 2014. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol. 64: 346–351. Article
- Kim D, Park S, Chun J. 2021b. Introducing EzAAI: a pipeline for high throughput calculations of prokaryotic average amino acid identity. J Microbiol. 59: 476–480. ArticlePDF
- Kumar S, Stecher G, Suleski M, Sanderford M, Sharma S, et al. 2024. MEGA12: molecular evolutionary genetic analysis version 12 for adaptive and green computing. Mol Biol Evol. 41: msae263.ArticlePubMedPMC
- Lee LH, Azman AS, Zainal N, Eng SK, Mutalib NA, et al. 2014. Microbacterium mangrovi sp. nov., an amylolytic actinobacterium isolated from mangrove forest soil. Int J Syst Evol Microbiol. 64: 3513–3519. Article
- Lee I, Kim YO, Park SC, Chun J. 2016. OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol. 66: 1100–1103. ArticlePubMed
- Lee SD, Yang HL, Kim IS. 2023. Four new Microbacterium species isolated from seaweeds and reclassification of five Microbacterium species with a proposal of Paramicrobacterium gen. nov. under a genome-based framework of the genus Microbacterium. Front Microbiol. 14: 1299950.ArticlePubMedPMC
- Letunic I, Bork P. 2024. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52: W78–W82. ArticlePubMedPDF
- Lewis II JS, Mathers AJ, Bobenchik AM, Bryson AL, Campeau S, et al. 2025. CLSI M100-ED35:2025 performance standards for antimicrobial susceptibility testing, 35th edn. Available from https://em100.edaptivedocs.net/GetDoc.aspx. Link
- Madhaiyan M, Poonguzhali S, Lee JS, Lee KC, Saravanan VS, et al. 2010. Microbacterium azadirachtae sp. nov., a plant-growth-promoting actinobacterium isolated from the rhizoplane of neem seedlings. Int J Syst Evol Microbiol. 60: 1687–1692. Article
- Manni M, Berkeley MR, Seppey M, Zdobnov EM. 2021. BUSCO: assessing genomic data quality and beyond. Curr Protoc. 1: e323. ArticlePubMedLink
- Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M. 2013. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinformatics. 14: 60.ArticlePubMedPDF
- Meier-Kolthoff JP, Sarda Carbasse J, Peinado-Olarte RL, Göker M. 2022. TYGS and LPSN: a database tandem for fast and reliable genome-based classification and nomenclature of prokaryotes. Nucleic Acids Res. 50: D801–D807. ArticlePubMedPMCPDF
- Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, et al. 2020. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol. 37: 1530–1534. ArticlePubMedPMCPDF
- Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, et al. 1984. An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods. 2: 233–241. Article
- Mondani L, Piette L, Christen R, Bachar D, Berthomieu C, et al. 2013. Microbacterium lemovicicum sp. nov., a bacterium isolated from a natural uranium-rich soil. Int J Syst Evol Microbiol. 63: 2600–2606. Article
- O’Leary NA, Cox E, Holmes JB, Anderson WR, Falk R, et al. 2024. Exploring and retrieving sequence and metadata for species across the tree of life with NCBI datasets. Sci Data. 11: 732.ArticlePubMedPMC
- Orla-Jensen S. 1919. The lactic acid bacteria. A. F. Høst.Link
- Parte AC, Sarda Carbasse J, Meier-Kolthoff JP, Reimer LC, Göker M. 2020. List of prokaryotic names with standing in nomenclature (LPSN) moves to the DSMZ. Int J Syst Evol Microbiol. 70: 5607–5612. ArticlePubMedPMC
- Qin QL, Xie BB, Zhang XY, Chen XL, Zhou BC, et al. 2014. A proposed genus boundary for the prokaryotes based on genomic insights. J Bacteriol. 196: 2210–2215. ArticlePubMedPMCLink
- Richter M, Rosselló-Móra R. 2009. Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA. 106: 19126–19131. ArticlePubMedPMC
- Rosselló-Móra R, Amann R. 2015. Past and future species definitions for bacteria and archaea. Syst Appl Microbiol. 38: 209–216. Article
- Sasser M. 1990. Identification of bacteria by gas chromatography of cellular fatty acids. MIDI technical note 101. MIDI Inc.. PDF
- Schwengers O, Jelonek L, Dieckmann MA, Beyvers S, Blom J, et al. 2021. Bakta: rapid and standardized annotation of bacterial genomes via alignment-free sequence identification. Microb Genom. 7: 000685.ArticlePubMed
- Smith EF. 1910. A new tomato disease of economic importance. Science. 31: 794–796.Link
- Smith EF. 1913. A new type of bacterial disease. Science. 38: 926.Article
- Suzuki K, Hamada M. 2015. Microbacterium. In Trujillo ME, Dedysh S, DeVos P, Hedlund B, Kämpfer P, et al. (eds.), Bergey’s manual of systematics of archaea and bacteria, 1st edn, pp. 1–52. John Wiley & Sons, Inc.Link
- Takeuchi M, Hatano K. 1998. Union of the genera Microbacterium Orla-Jensen and Aureobacterium Collins et al. in a redefined genus Microbacterium. Int J Syst Bacteriol. 48: 739–747. ArticlePubMed
- Tonkin-Hill G, MacAlasdair N, Ruis C, Weimann A, Horesh G, et al. 2020. Producing polished prokaryotic pangenomes with the Panaroo pipeline. Genome Biol. 21: 180.ArticlePubMedPMCPDF
- Wick RR, Judd LM, Gorrie CL, Holt KE. 2017. Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput Biol. 13: e1005595. ArticlePubMedPMC
- Worrall LJ, Majewski DD, Strynadka NCJ. 2023. Structural insights into type III secretion systems of the bacterial flagellum and injectisome. Annu Rev Microbiol. 77: 669–698. Article
- Young CC, Busse HJ, Langer S, Chu JN, Schumann P, et al. 2010. Microbacterium agarici sp. nov., Microbacterium humi sp. nov. and Microbacterium pseudoresistens sp. nov., isolated from the base of the mushroom Agaricus blazei. Int J Syst Evol Microbiol. 60: 854–860. ArticlePubMed
- Zho J, Huang Y, Yang J, Lai XH, Jin D, et al. 2021. Microbacterium chengjingii sp. nov. and Microbacterium fandaimingii sp. nov., isolated from bat faeces of Hipposideros and Rousettus species. Int J Syst Evol Microbiol. 71: 004858.Article
- Zhou Z, Tran PQ, Breister AM, Liu Y, Kieft K, et al. 2022. METABOLIC: high-throughput profiling of microbial genomes for functional traits, metabolism, biogeochemistry, and community-scale functional networks. Microbiome. 10: 33.ArticlePubMedPDF
- Zlamala C, Schumann P, Kämpfer P, Valens M, Rosselló-Móra R, et al. 2002. Microbacterium aerolatum sp. nov., isolated from the air in the 'Virgilkapelle' in Vienna. Int J Syst Evol Microbiol. 52: 1229–1234. ArticlePubMed
Citations
Citations to this article as recorded by






 ePub Link
ePub Link Cite this Article
Cite this Article
 MSK
MSK





